Abstract
Although, there have been rigorous research on the Indian caste system by several disciplines, it is still one of the most controversial socioscientific topic. Previous genetic studies on the subcontinent have supported a classical hierarchal sharing of genetic component by various castes of India. In the present study, we have used high-resolution mtDNA and Y chromosomal markers to characterize the genetic structuring of the Uttarakhand populations in the context of neighboring regions. Furthermore, we have tested whether the genetic structuring of caste populations at different social levels of this region, follow the classical chaturvarna system. Interestingly, we found that this region showed a high level of variation for East Eurasian ancestry in both maternal and paternal lines of descent. Moreover, the intrapopulation comparison showed a high level of heterogeneity, likely because of different caste hierarchy, interpolated on asymmetric admixture of populations inhabiting on both sides of the Himalayas.
Similar content being viewed by others
Introduction
South Asia is one of the major hotspots for the modern human migration, which has been attested by the molecular data provided by different genetic systems.1, 2, 3, 4, 5 However, the inferences drawn from these studies are contrasting.6, 7, 8, 9 As far as the mitochondrial DNA (mtDNA) is concerned, it is clear that most of the maternal ancestry of the subcontinent is autochthonous.10, 11, 12, 13, 14 However, the Y chromosome studies still needs a thorough examination.15, 16 The Autosomal studies have suggested the presence of two major components,6, 17 nevertheless require validation from ascertainment bias free data that is, complete genome sequence data.18 Apart from population dispersal and admixture at intercontinental level, local migration and admixture can also have significant role in shaping the genetic diversity of the subcontinent.19, 20, 21 Therefore, this study is an attempt to decipher the variations in this context at a higher resolution thereby conferring the fine-scale genetic structure of the populations living in the Uttarakhand state of North India.
The state of Uttarakhand, formerly known as Uttaranchal is one of the Himalayan states of India. It is bordered in the North by Tibet, South by Uttar Pradesh, East by Nepal and West by Himachal Pradesh. The size of its present-day population is about 10 million with major proportion of Brahmin and Kshatriya populations.22 The population density of this region is 2.5 times lower than average India due to cold climate and steep hilly regions. The state comprises 13 districts divided into two zones, known as Gharwal and Kumaon (Figure 1). This state also comprises of several Hindu pilgrimages (Devbhumi—literal meaning the Land of Gods), and is considered as a land of religious significance among Hindus. Considering the prehistoric human occupation in the light of ancient rock paintings and Palaeolithic stone tools, this region can be fundamental for the information on South Asian relic lineages.23, 24, 25
The caste structure of this region is similar to that classified in the Indian classical chaturvarna system.26 Majority of the caste and tribal populations of this region speak various branches of Indo-Aryan languages, collectively known as Pahari branch.27 To cover a detailed understanding about the genetic structure of this state, we have analyzed the maternal and paternal lines of ancestries of various ethnic groups of this region. Moreover, keeping in mind the geographical position of this state, we first evaluated the sex-specific genetic admixture with the Eastern and Western Eurasians, and further evaluated genetics of the classical caste system.
Materials and methods
Sampling
About 5.0 ml blood samples were collected from each individual belonging to six caste populations from three major districts of the Gharwal division of the Himalayan state of Uttarakhand (Figure 1). Altogether 323 samples were analyzed for mtDNA and Y chromosomal markers (Table 1; Supplementary Tables 1 and 2). The sampling involved a detailed interview procedure with queries pertaining to name of the caste, which they belonged, and any oral history about their ancestry and so on. The related individuals with similar family background were avoided. This project was approved by the Institutional Ethical Committee of the Council of Scientific and Industrial Research (CSIR)-Centre for Cellular and Molecular Biology, Hyderabad, India. The informed written consent was obtained from all the volunteers.
Genotyping
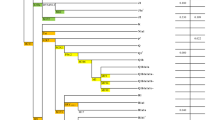
We sequenced the hypervariable segment I of mtDNA and the variations were scored against the revised Cambridge Reference Sequence28 and Reconstructed Sapiens Reference Sequence29 (Supplementary Tables 1 and 2). Haplogroups were assigned collectively based on hypervariable segment I variations and by genotyping the coding regions mutations published till date in PhyloTree (build 16).30 For the comparative analysis, we have merged our data with the published sources.14, 16, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45 A total of 30 Y chromosome biallelic markers from previously published data set46 were used in this study for assigning the haplogroup to each individual. For haplogroup assignment we have used Karmin et al.47 and recently updated nomenclature of The International Society of Genetics Genealogy (ISOGG).48 We have also merged data of Tharu population from the same region published elsewhere.14 The most parsimonious tree for mtDNA (Supplementary Figures 1–3) and Y chromosome (Supplementary Figure 4), were built following the median-joining algorithm.49
Statistical analysis
Principal component analysis for mtDNA and Y chromosomal haplogroups was performed with merged data sets by using software POPSTR (http://harpending.humanevo.utah.edu/popstr/), kindly provided by H. Harpending. We have used hypervariable segment I sequence data for mtDNA and binary coded (0 as ancestral and 1 as derived) for Y chromosome. Subsequently, the genetic distances among populations and populations groups were generated from Arlequin 3.5.50
Results and discussion
The genetic structure of the Uttarakhand populations was first studied in the context of Eurasian populations. The analysis of maternal and paternal lineages showed a contrasting sharing of ancestry with East and West Eurasians (Table 1 and Supplementary Table 3). Analyses of the haplogroups and sub-haplogroups revealed a prevalent East Eurasian component for both male and female gene pools, more evident in maternal ancestry of the Uttarakhand. We observed that the maternal lineages of the Uttarakhand comprised of a high proportion of east Eurasian haplogroups, consistent with the other Himalayan states.14, 18, 36, 51 Conversely to the maternal ancestry, the paternal heritage was mainly comprised of the autochthonous Indian and West Eurasian-specific lineages (Supplementary Figure 4 and Supplementary Table 3). When we compared maternal lineage of Uttarakhand at the haplotype level, we found that majority of East Eurasian-specific haplotypes were prevalent in Tibet (Supplementary Figures 1–3), supporting a similar scenario observed in Nepal.18 The massive haplotype sharing with the neighboring Uttar Pradesh and Nepal (rather than Northeast India) was evident in the Network analyses (Supplementary Figures 1–3).
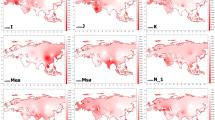
The frequency based principle component analysis for both mtDNA and Y chromosome haplogroups placed the Uttarakhand population over the North Indian-Nepali cluster (Figure 2). The East Eurasian vs South Asian lineages were major precursor for the PC1 in both the genetic systems, whilst, PC2 was derived by Tibetan vs Southeast Asian lineages. The high heterogeneity of maternal lineages observed in frequency distribution (Table 1) and Network analysis (Supplementary Figures 1–3), is also attested by the principal component analysis. Four Y chromosome haplogroups were the major predictors of the principal component analysis structure. East Eurasian-specific haplogroups (for example, O2a1-M95 and O3a2c1-M134) were responsible for their affinity with East/Southeast Asia, whereas haplogroups R1a1a-M17 and R2a-M124 were playing antagonistic roles. The substantial presence of Y chromosomal haplogroup Q-P36.2 also links them with the Central Asian populations (Supplementary Table 3). In turn, the West Eurasian-specific lineages show substantial elevation than observed in the state of Uttar Pradesh.
Principal component analysis (PCA) plots constructed on the basis of mtDNA (upper panel) and Y chromosome (lower panel) haplogroup frequencies. Data from neighboring regions are compiled from published sources (given in Materials and methods section). A full color version of this figure is available at the Journal of Human Genetics journal online.
To gain a comprehensive overview of inter and intra-caste relations of this region, we compared the maternal and paternal heritage of various caste and tribal populations. Previous genetic studies on different caste populations have failed to make a consensus on their origins and affinities.19, 52, 53, 54 It was also suggested that the restricted marriage practice was the major contributing factor for a fine tapestry of the social system.43, 54 However, contradictory conclusion were drawn for the roles of males and females in shaping the caste stratification.19, 43, 54, 55 When comparing with the previous analyses,43, 52 our data showed significant (unpaired t-test P<0.0001) higher proportion of West Eurasian-specific lineages among traditionally higher caste (Brahmin and Kshatriya) populations for maternal lineages, although it was non-significant for the paternal ancestry (unpaired t-test P=0.5468).
While comparing the genetic distances, we found that the Brahmins and Kshatriya consistently remained as outliers. Therefore, to evaluate the roles of males and females in the social stratification of this region, we calculated the distances of studied populations, with respect to the Brahmins, who were the extreme outlier in the tree (Figure 3). For this, we have grouped populations according to their classical social status—Brahmins, Kshatriya, Vaishya (Goswami and Shah) and Shudra (Arya and Tamta). Interestingly, the maternal structure of this region was in congruent with the classical social system, where distance from Brahmins to other groups was following a social ladder type structure (Figure 3a). However, the male line of descent did not reveal any kind of such local structure (Figure 3b). Notably, the distance from Brahmins to Vaishya was significantly higher than Brahmins to Shudra (t-test P<0.001). Moreover, Arya (Shudra) was closest to Brahmins and Kshatriya than to any other non-Brahmin and Kshatriya populations. The affinity of Arya towards Brahmins and Kshatriya is due to the analogous high frequency of haplogroups R1a1a-M17 and R2a-M124 (Supplementary Figure 4 and Supplementary Table 3). A similar observation was reported elsewhere,19 however, the most parsimonious reason given for such discrepancy is highly unlikely in our case giving out the closer paternal ancestry of Shudra (Arya) with the Brahmin and Kshatriya (Supplementary Figure 4).
In conclusions, our extensive analysis on uniparentally inherited markers led to the most precise identification of East/Southeast Asian, South Asian and West Eurasian haplogroups among Uttarakhand populations and a better understanding of the extent of admixture from East and West. We have observed that the admixture of East Eurasian lineages to Uttarakhand population was inclined towards higher female geneflow. The unexpected link of Shudra (Arya) with the higher traditional caste (Brahmin and Kshatriya) was major factor likely to be responsible for disordering the traditional caste barrier in the Uttarakhand state. In addition, generating well resolved mtDNA and Y chromosomal lineages allowed us to detect the directional geneflow and their accretion from other side of the Himalayas.
References
Kivisild, T., Papiha, S. S., Rootsi, S., Parik, J., Kaldma, K., Reidla, M. et al in An Indian Ancestry: A Key for Understanding Human Diversity in Europe and Beyond (eds Renfrew, C. & Boyle, K .) 267–279 (McDonald Institute for Archaeological Research University of Cambridge, Cambridge, 2000).
Chaubey, G., Metspalu, M., Kivisild, T. & Villems, R. Peopling of South Asia: investigating the caste-tribe continuum in India. Bioessays 29, 91–100 (2007).
Majumder, P. P. The human genetic history of South Asia. Curr. Biol. 20, R184–R187 (2010).
Majumder, P. P. & Basu, A. A genomic view of the peopling and population structure of India. Cold Spring Harb. Perspect. Biol. 7, a008540 (2014).
Pala, M., Soares, P., Chaubey, G., Richards, M. in Archaeogenetics (eds Barker, Graeme & Goucher, Candice ) 26–54 (Cambridge University Press, Cambridge, UK, 2015).
Metspalu, M., Romero, I. G., Yunusbayev, B., Chaubey, G., Mallick, C. B., Hudjashov, G. et al. Shared and unique components of human population structure and genome-wide signals of positive selection in South Asia. Am. J. Hum. Genet. 89, 731–744 (2011).
Moorjani, P., Thangaraj, K., Patterson, N., Lipson, M., Loh, P. R., Govindaraj, P. et al. Genetic evidence for recent population mixture in India. Am. J. Hum. Genet. 93, 422–438 (2013).
Chaubey, G. Language isolates and their genetic identity: a commentary on mitochondrial DNA history of Sri Lankan ethnic people: their relations within the island and with the Indian subcontinental populations. J. Hum. Genet. 59, 61–63 (2014).
Chaubey, G., Kadian, A., Bala, S. & Rao, V. R. Genetic affinity of the Bhil, Kol and Gond mentioned in epic ramayana. PloS ONE 10, e0127655 (2015).
Sun, C., Kong, Q. P., Palanichamy, M. G., Agrawal, S., Bandelt, H. J., Yao, Y. G. et al. The dazzling array of basal branches in the mtDNA macrohaplogroup M from India as inferred from complete genomes. Mol. Biol. Evol. 23, 683–690 (2006).
Palanichamy, M., Sun, C., Agrawal, S., Bandelt, H. J., Kong, Q. P., Khan, F. et al. Phylogeny of mtDNA macrohaplogroup N in India based on complete sequencing: implications for the peopling of South Asia. Am. J. Hum. Genet. 75, 966–978 (2004).
Thangaraj, K., Chaubey, G., Singh, V. K., Vanniarajan, A., Thanseem, I., Reddy, A. G. et al. In situ origin of deep rooting lineages of mitochondrial Macrohaplogroup 'M' in India. BMC Genom. 7, 151 (2006).
Chandrasekar, A., Kumar, S., Sreenath, J., Sarkar, B. N., Urade, B. P., Mallick, S. et al. Updating phylogeny of mitochondrial DNA macrohaplogroup m in India: dispersal of modern human in South Asian corridor. PloS ONE 4, e7447 (2009).
Chaubey, G., Singh, M., Crivellaro, F., Tamang, R., Nandan, A., Singh, K. et al. Unravelling the distinct strains of Tharu ancestry. Eur. J. Hum. Genet. 22, 1404–1412 (2014).
Sahoo, S., Singh, A., Himabindu, G., Banerjee, J., Sitalaximi, T., Gaikwad, S. et al. A prehistory of Indian Y chromosomes: evaluating demic diffusion scenarios. Proc. Natl Acad. Sci. USA 103, 843–848 (2006).
Sengupta, S., Zhivotovsky, L. A., King, R., Mehdi, S. Q., Edmonds, C. A., Chow, C. E. et al. Polarity and temporality of high-resolution y-chromosome distributions in India identify both indigenous and exogenous expansions and reveal minor genetic influence of Central Asian pastoralists. Am. J. Hum. Genet. 78, 202–221 (2006).
Reich, D., Thangaraj, K., Patterson, N., Price, A. L. & Singh, L. Reconstructing Indian population history. Nature 461, 489–494 (2009).
Wang, H. W., Li, Y. C., Sun, F., Zhao, M., Mitra, B., Chaudhuri, T. K. et al. Revisiting the role of the Himalayas in peopling Nepal: insights from mitochondrial genomes. J. Hum. Genet. 57, 228–234 (2012).
Bamshad, M. J., Watkins, W. S., Dixon, M. E., Jorde, L. B., Rao, B. B., Naidu, J. M. et al. Female gene flow stratifies Hindu castes. Nature 395, 651–652 (1998).
Sharma, G., Tamang, R., Chaudhary, R., Singh, V. K., Shah, A. M., Anugula, S. et al. Genetic affinities of the central Indian tribal populations. PloS ONE 7, e32546 (2012).
Thangaraj, K., Naidu, B. P., Crivellaro, F., Tamang, R., Upadhyay, S., Sharma, V. K. et al. The influence of natural barriers in shaping the genetic structure of maharashtra populations. PloS ONE 5, e15283 (2010).
Uttarakhand Census [Internet]. Available at http://www.census2011.co.in/census/state/uttarakhand.html.
Tewari, R. Some Important Stone Images from Uttaranchal Himalaya. South Asian Stud. 23, 95–106 (2007).
Khanduri, B. Archaeology of central Himalaya: a review In: Puraratna Emerging Trends in Archaeology, Art, Anthropology, Conservation, and History: in Honour of Shri Jagat Pati Joshi 1, 213 (2002).
Lahiri, N. & Bacus, E. Exploring the archaeology of Hinduism In: World Archaeology 36, 313–325 (2004).
Chaubey, G. The demographic history of india: a perspective based on genetic evidence. http://hdl.handle.net/10062/15240 PhD thesis. Universitatis Tartuensis, Estonia, (2010).
Grierson, G. A. The Pahari Language, British India Press, (1915).
Andrews, R. M., Kubacka, I., Chinnery, P. F., Lightowlers, R. N., Turnbull, D. M. & Howell, N. Reanalysis and revision of the Cambridge reference sequence for human mitochondrial DNA. Nat. Genet. 23, 147 (1999).
Behar, D. M., van Oven, M., Rosset, S., Metspalu, M., Loogväli, E. L., Silva, N. M. et al. A "copernican" reassessment of the human mitochondrial DNA tree from its root. Am. J. Hum. Genet. 90, 675–684 (2012).
van Oven, M. & Kayser, M. Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation. Hum. Mutat. 30, E386–E394 (2009).
Basu, A., Mukherjee, N., Roy, S., Sengupta, S., Banerjee, S., Chakraborty, M. et al. Ethnic India: a genomic view, with special reference to peopling and structure. Genome Res. 13, 2277–2290 (2003).
Metspalu, M., Kivisild, T., Metspalu, E., Parik, J., Hudjashov, G., Kaldma, K. et al. Most of the extant mtDNA boundaries in south and southwest Asia were likely shaped during the initial settlement of Eurasia by anatomically modern humans. BMC Genet. 5, 26 (2004).
Cordaux, R., Weiss, G., Saha, N. & Stoneking, M. The northeast Indian passageway: a barrier or corridor for human migrations? Mol. Biol. Evol. 21, 1525–1533 (2004).
Kivisild, T., Rootsi, S., Metspalu, M., Mastana, S., Kaldma, K., Parik, J. et al. The genetic heritage of the earliest settlers persists both in Indian tribal and caste populations. Am. J. Hum. Genet. 72, 313–332 (2003).
Sharma, S., Saha, A., Rai, E., Bhat, A. & Bamezai, R. Human mtDNA hypervariable regions, HVR I and II, hint at deep common maternal founder and subsequent maternal gene flow in Indian population groups. J. Hum. Genet. 54, 47–55 (2005).
Fornarino, S., Pala, M., Battaglia, V., Maranta, R., Achilli, A., Modiano, G. et al. Mitochondrial and Y-chromosome diversity of the Tharus (Nepal): a reservoir of genetic variation. BMC Evol. Biol. 9, 154 (2009).
Hill, C., Soares, P., Mormina, M., Macaulay, V., Meehan, W., Blackburn, J. et al. Phylogeography and ethnogenesis of aboriginal Southeast Asians. Mol. Biol. Evol. 23, 2480–2491 (2006).
Qin, Z., Yang, Y., Kang, L., Yan, S., Cho, K., Cai, X. et al. A mitochondrial revelation of early human migrations to the Tibetan Plateau before and after the last glacial maximum. Am. J. Phys. Anthropol. 143, 555–569 (2010).
Chaubey, G., Metspalu, M., Choi, Y., Mägi, R., Romero, I. G., Soares, P. et al. Population genetic structure in indian austroasiatic speakers: the role of landscape barriers and sex-specific admixture. Mol. Biol. Evol. 28, 1013–1024 (2011).
Debnath, M., Palanichamy, M. G., Mitra, B., Jin, J. Q., Chaudhuri, T. K. & Zhang, Y. P. Y-chromosome haplogroup diversity in the sub-Himalayan Terai and Duars populations of East India. J. Hum. Genet. 56, 765–771 (2011).
Kumar, V., Reddy, A. N. S., Babu, J. P., Rao, T. N., Langstieh, B. T., Thangaraj, K. et al. Y-chromosome evidence suggests a common paternal heritage of Austro-Asiatic populations. BMC Evol. Biol. 7, 47 (2007).
Trivedi, R., Sahoo, S., Singh, A., Bindu, G., Banerjee, J., Tandon, M. et al. Genetic imprints of pleistocene origin of indian populations: a comprehensive phylogeographic sketch of indian Y-Chromosomes. Int. J. Hum. Genet. 8, 97–118 (2008).
Thanseem, I., Thangaraj, K., Chaubey, G., Singh, V. K., Bhaskar, L. V. K. S., Reddy, B. M. et al. Genetic affinities among the lower castes and tribal groups of India: inference from Y chromosome and mitochondrial DNA. BMC Genet. 7, 42 (2006).
Zerjal, T., Pandya, A., Thangaraj, K., Ling, E. Y. S., Kearley, J., Bertoneri, S. et al. Y-chromosomal insights into the genetic impact of the caste system in India. Hum. Genet. 121, 137–144 (2007).
Kayser, M., Brauer, S., Cordaux, R., Casto, A., Lao, O., Zhivotovsky, L. A. et al. Melanesian and Asian origins of Polynesians: mtDNA and Y chromosome gradients across the Pacific. Mol. Biol. Evol. 23, 2234–2244 (2006).
Karafet, T. M., Mendez, F. L., Meilerman, M. B., Underhill, P. A., Zegura, S. L. & Hammer, M. F. New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree. Genome Res. 18, 830–838 (2008).
Karmin, M., Saag, L., Vicente, M., Sayres, M. A. W., Järve, M., Talas, U. G. et al. A recent bottleneck of Y chromosome diversity coincides with a global change in culture. Genome Res. 25, 459–466 (2015).
Y-DNA Haplogroup Tree 2015, Version: 10.38, Date: 2 August 2015 (Internet). 2015 (cited on 4.08.2015). Available at http://www.isogg.org/tree/ISOGG_YDNATreeTrunk.html.
Bandelt, H. J., Forster, P. & Rohl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 16, 37–48 (1999).
Excoffier, L. & Lischer, H. E. L. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 10, 564–567 (2010).
Thangaraj, K., Chaubey, G., Kivisild, T., Selvi Rani, D., Singh, V. K., Ismail, T. et al. Maternal footprints of Southeast Asians in North India. Hum. Hered. 66, 1–9 (2008).
Bamshad, M., Kivisild, T., Watkins, W. S., Dixon, M. E., Ricker, C. E., Rao, B. B. et al. Genetic evidence on the origins of Indian caste populations. Genome Res. 11, 994–1004 (2001).
Wooding, S., Ostler, C., Prasad, B. V. R., Watkins, W.S., Sung, S., Bamshad, M. et al. Directional migration in the Hindu castes: inferences from mitochondrial, autosomal and Y-chromosomal data. Hum. Genet. 115, 221–229 (2004).
Cordaux, R., Aunger, R., Bentley, G., Nasidze, I., Sirajuddin, S. M. & Stoneking, M. Independent origins of Indian caste and tribal paternal lineages. Curr. Biol. 14, 231–235 (2004).
Cordaux, R., Deepa, E., Vishwanathan, H. & Stoneking, M. Genetic evidence for the demic diffusion of agriculture to India. Science 304, 1125 (2004).
Acknowledgements
We thank all the volunteer donors for their participation in this study. RT was supported by the European Social Funds Doctoral Studies and Internationalisation Programme DoRa. GC acknowledges the financial support from European Union European Regional Development Fund through the Centre of Excellence in Genomics to Estonian Biocentre and University of Tartu by Tartu University grant (PBGMR06901), by Estonian Personal grants PUT-766 and Estonian Institutional Research grants IUT24-1. LS was supported by Bhatnagar (CSIR) and JC Bose (DST) Fellowships, Government of India. KT was supported by the Council of Scientific and Industrial Research, Government of India (GENESIS: BSC0121) and (BSC 0208). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. All the sequence files have been deposited to GenBank (id KT362381-KT362703).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website .
Rights and permissions
About this article
Cite this article
Negi, N., Tamang, R., Pande, V. et al. The paternal ancestry of Uttarakhand does not imitate the classical caste system of India. J Hum Genet 61, 167–172 (2016). https://doi.org/10.1038/jhg.2015.121
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2015.121
This article is cited by
-
The matrilineal ancestry of Nepali populations
Human Genetics (2023)