Abstract
Among the people living in the five deep Southern Thai provinces, Thai-Malay Muslims (MUS) constitute the majority, while the remaining are Thai Buddhists (BUD). Cultural, linguistic and religious differences between these two populations have been previously reported. However, their biological relationship has never been investigated. In this study, we aimed to reveal the genetic structure and genetic affinity between MUS and BUD by analyzing 15 autosomal short tandem repeats. Both distance and model-based clustering methods showed significant genetic homogeneity between these two populations, suggesting a common biological ancestry. After Islamization in this region during the fourteenth century AD, gradual albeit nonstatistically significant genetic changes occurred within these two populations. Cultural barriers possibly influenced these genetic changes. MUS have closer admixture to Malaysian-Malay Muslims than BUD countrywide. Admixture proportions also support certain degree of genetic dissimilarity between the two studied populations, as shown by the unequal genetic contribution from Malaysian-Malay Muslims. Cultural transformation and recent minor genetic admixture are the likely processes that shaped the genetic structure of both MUS and BUD.
Similar content being viewed by others
Introduction


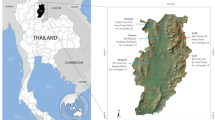
Thailand, one of the biggest Buddhist countries, is in the geographical heart of Southeast Asia. The country is divided into six regions: the North, the Northeast, the East, the West, the Central and the South. Stretching down the Malay Peninsula, the South of Thailand borders the Gulf of Thailand to the East, the Andaman Sea to the West, and Malaysia to the South. On the basis of the most recent census of 65.9 million Thais,1 the vast majority practices Buddhism (93.6% or 61.7 million) and the largest minority practices Islam (4.9% or 3.2 million).2 The percentages flip in Southern Thailand.3 Over 80% of the 3.2 million Muslims in Thailand live in the five deep Southern Provinces of Narathiwat, Pattani, Yala, Satun and Songkhla (Figure 1).
Southeast Asian map showing the locations of studied populations (represented by ) and compared populations (represented by ) in Thailand, Malaysia and Philippines. The sample size for each population varies from 46 to 304 unrelated individuals. The references for the compared populations are given in Table 1.
On the basis of historical and archeological evidence, prehistoric people inhabited the territory of deep Southern Thailand, and the area was nurtured by contacts with other cultures since the first century. Hindus from India migrated to occupy this area during the Indianization period (third to fifth centuries). The Buddhist Mon people then inhabited this area between the fifth and eighth centuries. Subsequently, the Malays populated the area, found the Hindu–Buddhist kingdom of Srivijaya, and dominated this peninsular from the eighth to eleventh centuries. The Hindu–Buddhist Malay kingdom of Patani Sultunate was established as early as the ninth century. The Khmer then governed the area from the eleventh to thirteenth centuries, followed by the southward movements of the Thai people from Siam.4 Islamization of the Patani Sultunate occurred in the fourteenth century. This territory became a vassal state under the Ayutthaya Kingdom in the early sixteenth century.4 Throughout history, Siam (Ayutthaya Kingdom, Thonburi Kingdom and then Thailand) attacked the Patani Sultunate many times over many hundred years.5 Finally, it was annexed as part of Siam under the Anglo-Siamese Treaty of 1909.6
Muslims in Thailand can be categorized into two groups: the Sunni-Malay Muslims (henceforth known as Thai-Malay Muslims (MUS)) and Shiite-Muslims (non-Malay Muslims). The non-Malay Muslims descended from Arabs and Persians, and they are fewer in numbers compared with the MUS who are of Malay ancestry. The non-Malay Muslims speak Thai as their main language and assimilate well in the Thai society. In contrast, the MUS speak ‘Yawee’, a local dialect of Bahasa Melayu (an Austronesian linguistic family). They follow the Malay traditions, use Malay names and listen to Malay music. All of these hinder population admixture among Thai Buddhists (BUD) and MUS.3,7 The modern history of Southern Thailand is marred by various separatist movements carried out by locals who resist the assimilation of their culture with the Bangkok/Buddhist-centric government. In other words, the ethnocentric policies of Thailand alienate the MUS.8 Therefore, MUS and BUD have remained culturally, linguistically, religiously and politically distant from each other. However, comparison of genetic structure between these two populations has never been carried out until this study.
Microsatellites or short tandem repeats (STRs) have been proven as a powerful tool for investigating phylogenetic relationships and for forensic identification, as these loci are distributed throughout the human genome, have a high mutation rate, and are highly polymorphic.9,10 Thousands of microsatellites have been mapped, but <20 (the CODIS and the ENFSI loci) are currently in use for individualization by forensic and population geneticists worldwide.11 In Thailand, studies on forensic STRs and their utilization for population genetics are quite limited.12, 13, 14, 15, 16
The present study used the 15 forensic autosomal microsatellites to investigate the genetic variation and structure of two Southern Thai populations from the three deep Southern Provinces of Thailand. The genetic relationships of the two studied populations and other populations residing in Thailand, Malaysia and the Philippines from earlier studies13,17,18 were also evaluated.
Materials and methods
Samples
Buccal swabs were collected, under informed consent, from 150 unrelated individuals from the deep South of Thailand, in the area of Narathiwat, Pattani and Yala Provinces (Thai-Malay Muslim, n=104, of which 17 were from Narathiwat, 44 from Pattani and 43 from Yala; Thai Buddhist, n=46, of which 6 were from Narathiwat, 15 from Pattani and 25 from Yala). Ethical approval was obtained from the Ethics Board of Prince of Songkla University. Since the statistical analyses in this study were on the basis of Bayesian-clustering algorithm, the compared populations used to determine population affinity were selected from previous studies in which raw genotypic data of 15 STRs are available. Nine neighboring populations from Malaysia (ML1, ML2, CH-ML and IN-ML), Thailand (TH-North, TH-Isan, TH-Central and TH-Southern) and Philippines (FI) were utilized (Figure 1 and Table 1).
DNA extraction and STR typing
Genomic DNAs were extracted from the buccal samples using the Promega IQ kit (Promega Corp., Madison, WI, USA). Extracted DNAs were quantified using the Quantifiler Human DNA Quantification Kit (Applied Biosystem, Foster City, CA, USA). Multiplex PCR amplification from 0.5 ng of template DNA was performed using a commercial AmpFℓSTR Identifiler PCR Amplification kit (Applied Biosystem), which amplifies 15 autosomal STR loci and one gender determination locus: D8S1179, D21S11, D7S820, CSF1PO, D3S1358, TH01, D13S317, D16S539, vWA, TPOX, D18S51, D5S818, FGA, D19S433, D2S1338 and amelogenin. Amplicons were genotyped by multicapillary electrophoresis in an ABI Prism 3130xl Genetic Analyzer (Applied Biosystem) following the manufacturer’s instruction. The results were further analyzed using GeneMapper v.3.2.1 (Applied Biosystem).
Statistical analyses
ARLEQUIN 3.5 software19 was used to determine STR allele frequencies, Hardy–Weinberg P-values, observed heterozygosity (Ho), expected heterozygosity, number of allele and gene diversity. Significance levels for Hardy–Weinberg P-values were adjusted using sequential Bonferroni correction (α=0.05/15 or 0.0033).20 Several parameters of forensic and population genetic importance, including power of discrimination, match probability, polymorphic information content, power of exclusion and typical paternity index, were obtained with the Excel PowerStats v.12 spread sheet (www.promega.com/geneticidtools/powerstats).
To elucidate population affinity, both distance-based and model-based clustering approaches were employed. Pairwise genetic distances on basis of allele frequency variance (Fst), as well as the statistical significance using 1000 permutations were calculated using ARLEQUIN 3.5. The Fst distance matrix was then plotted in three dimensions by means of multidimensional scaling (3D MDS) using the STATISTICA 10.0 software (StatSoft, Tulsa, OK, USA). Principal coordinate analysis (PCoA) was constructed by a covariance matrix with data standardization using GENALEX 6.3 software.21
Population structure was evaluated according to religion (Islam, Buddhism and Christianity) through hierarchical analysis of molecular variance,22 as implemented in Arlequin. Further investigation of population structure was performed using Bayesian-clustering method implemented in STRUCTURE 2.323, 24, 25 under assumptions of admixture, correlated allele frequencies and LOCPRIOR model.25 The number of clusters (K) was from 1 to 8 with five replicates performed using an MCMC chain burn-in length of 50 000 iterations followed by 100 000 iterations run length. STRUCTURE Harvester26 was used to calculate the greatest estimated mean posterior probability (Ln Prob) averaged over five replicates,23 as well as a second order rate of change logarithmic probability between the subsequent K values (delta K)27 to identify the optimal K in the data.
The estimator of admixture coefficient was calculated using ADMIX 2.0.28 As the estimated coefficients are less affected by the stochasticity of the mutation process, and as the short time scale of human admixture process is influenced by genetic drift rather than accumulated mutation,29 both mutation and molecular divergence between the alleles were not taken into account.28,30
Results
Genetic variability and genetic structure
The average Ho and gene diversity in MUS were 0.787 and 0.786, respectively, whereas 0.801 (Ho) and 0.784 (gene diversity) were observed in BUD. The Ho and gene diversity values in both populations were in the same range as the other compared populations (Table 1). Deviation from Hardy–Weinberg equilibrium was detected in only one locus (D18S51) in MUS after Bonferroni correction. In MUS, the combined match probability was 1 in 4.4917 × 1017 and the combined power of exclusion was 0.9999986, whereas BUD had 1 in 2.4579 × 1016 combined match probability and 0.9999996 combined power of exclusion. The combined power of discrimination for these loci was robust (>0.999999999999999 in both populations). In MUS, the most polymorphic locus was FGA, reflected by the highest Ho and number of alleles, whereas in BUD this was found in D21S11 and D8S1179, respectively. As expected, the most polymorphic loci in both populations were highly discriminating, as shown by the relatively high PD values. This demonstrates that this set of loci will be useful for forensic identification. Two private alleles in MUS (32.3 at D21S11 and 15 at D7S820, respectively) and only one allele in BUD (31 at FGA) were found. The allele frequency distributions, population genetic parameters and forensic parameters are listed in Supplementary Tables 1 and 2 for MUS and BUD, respectively.
The proportion of genetic variation within and between the different religious groups was assessed by analysis of molecular variance (Table 2). Interpopulation differences were higher when the populations constituted a single group (0.59%) compared with when populations were grouped according to religions (0.41%). The nonsignificant Fct statistics (P>0.01) observed when populations were grouped according to religion indicates that genetic structure was only minimally influenced by religion.
Population affinity
We performed pairwise Fst comparison with test for genetic differences between MUS and BUD, as well as compared them with previously studied populations. Forty out of fifty-five pairwise differences were statistically significant (P<0.01) (Table 3), but MUS and BUD exhibited nonsignificant genetic difference. MUS also showed nonsignificant genetic difference from ML1, ML2 and TH-South. BUD did not differ from the following five populations: ML1, TH-North, TH-Isan, TH-Central and TH-South. Among the 11 compared populations, IN-ML, FI and CH-ML statistically differed from other populations, indicating high genetic divergence.
We also conducted distance-based clustering (MDS and PCoA) to determine the genetic relationships among these populations. The 3D MDS plot shows genetic differentiation of IN-ML, FI and CH-ML in dimensions 1, 2 and 3, respectively (Figure 2). This agrees with the pairwise Fst analysis. The remaining eight populations clustered together around the center of the plot, which indicates close genetic affinity. Within the cluster itself, two population subgroups were weakly separated in dimension 3. MUS was grouped with two Malaysian populations (ML1 and ML2), whereas BUD was clustered with other Thai populations nationwide (TH-North, TH-Isan, TH-Central and TH-South). The PCoA result strongly agrees with the MDS result. IN-ML and FI were segregated from all populations on axis 1 and axis 2, which explained 49.34 and 23.54% of the distance matrix (Figure 3). CH-ML was dispersed further in an upward position on axis 3, which explained 16.11% of the variation (Figure 3). Again, a cluster of populations was seen in the middle of the plot. MUS lay near ML1 and ML2 but BUD formed a continuum running to axis 2 with TH-North, TH-Isan, TH-Central and TH-South. Interestingly, TH-South occupied the intermediate position among groups of populations from Thailand and Malaysia. This probably represents gene flow within TH-South.
Multidimensional scaling plot depicting population affinity based on Fst distances. The stress value is 0.021. Genetic differentiation of IN-ML, FI and CH-ML are observed in dimensions 1, 2 and 3, respectively. Other populations clustered at the center of the plot. Within the cluster itself, two population subgroups are separated in dimension 3: MUS is grouped with Malaysians (ML1 and ML2), whereas BUD is grouped with other Thais (TH-North, TH-Isan, TH-Central and TH-South). See the meaning of population abbreviations in Table 1.
PCoA was performed using a covariance matrix with data standardization. Axis 1, 2 and 3 explained 49.34%, 23.54% and 16.11% of the distance matrix, respectively. IN-ML and FI are separated from all the other populations. Similarly, CH-ML is above the other populations on axis 3 (lower panel). MUS lies near ML1 and ML2 but BUD forms a continuum running to axis 2 with TH-North, TH-Isan, TH-Central and TH-South. See the meaning of population abbreviations in Table 1.
Model-based clustering method was performed using STRUCTURE 2.3 to clarify population structures and their relationships. Because of the low level of genetic divergence among populations (Fst of 0–0.0189), we executed the analysis using sampling information, which provides more information.31 Cluster (K) values were varied from 1 to 8. According to the highest LnP (K), the most suitable K23 was observed at K=3. The ad hoc statistic delta K27 showed a strong modal peak at K=2. Table 4 shows the proportion of membership of each pre-defined population in each of the two or three clusters with mean LnP (K) and δK. At K=2, most populations belonged to cluster 1, with IN-ML as the only exception (membership proportion of 0.648 in cluster 2). When K was increased to 3, a new delineated cluster belonging to FI emerged (0.725). Clusters 1 and 2 still displayed similar proportion of membership. The consistency between MDS, PCoA and STRUCTURE results indicate genetic distinction of IN-ML, FI and to a lesser extent the CH-ML. STRUCTURE result also reveals no clear population differentiation in cluster 1, which was comprised of the two studied populations (MUS and BUD) and other populations from all parts of Thailand (TH-North, TH-Isan, TH-Central and TH-South) and Malaysia (ML1, ML2 and CH-ML). In other words, our analysis clearly suggests genetic homogeneity among these populations.
Admixture estimation
MDS and PCoA positioned BUD, MUS and TH-South midway between ML1 and ML2, TH-North, TH-Isan, TH-Central and FI. This suggests an admixed origin for BUD and MUS; thus we further evaluated population admixture. In the admixture analysis, two groups of populations (the admixed ones and the parental ones) were defined. The BUD, MUS and TH-South were considered as admixed populations who received genetic contributions from their putative parental sources: the Malay group (ML1 and ML2) and the Thai group (TH-North, TH-Isan and TH-Central). We discounted FI as a parental population based on historical evidence, even though FI was located periphery to the admixed groups. Figure 4 depicts the estimators of admixture coefficient. The parental Malay group contributed a lower proportion to BUD, MUS and TH-South (2.83, 33.49 and 36.11%) when compared with the parental Thai group (97.17, 66.51 and 63.89%).
Discussion
Several studies on molecular genetic loci have been reported with current Thai populations. However, those studies focused on linguistic and geographic factors in determining a population's genetic structure.15,16,32 In this study, we focused on the religious background of populations to determine the genetic relationship of MUS and BUD in the deep South of Thailand using 15 autosomal STR loci. On the basis of historical and anthropological facts, the people in Patani Sultunate originally practiced Hindu–Buddhism. Islamization took place around the 12th to 15th centuries, and the Patani Sultanate continued to prosper until it fell under Siamese control in the early 16th century.33 Local legend told of a learned Muslim who cured Raja Indra (the ruler of Patani Sultanate at that time) on the condition that he would be converted to Islam. From then on, two distinct religious populations have inhabited this area: the Muslims and the Buddhists.
According to the historical evidence, present day MUS could be the descendants of either local Hindu–Buddhist converts or Muslims from India who arrived during the spread of Islamic faith into the Malay Peninsula. Our prominent genetic finding is the genetic homogeneity between MUS and BUD (Figures 2 and 3). This supports the theory that MUS descends from the Hindu–Buddhist converts, who underwent cultural transformation during the reign of Raja Indra. Cultural transformation has shaped several Muslim communities, for example, Indian Muslims and Vietnamese Cham.34, 35, 36
Despite the genetic resemblance between MUS and BUD, some degree of differentiation was observed. MUS seemed to be genetically closer to the Malaysian Malays (ML1 and ML2), whereas BUD were more closely related to the remaining Thai populations nationwide. This suggests demographic movement through politics and trades within this micro-geographic region after the Islamization event.37 Therefore, we performed admixture analysis to infer recent genetic history between MUS and BUD. Parental Malay population contributed 30% and 3% to the gene pool of MUS and BUD, respectively (Figure 4). This finding is consistent with MDS and PCoA results (Figures 2 and 3). Admixture analysis indicates weak genetic divergence between these two studied populations, possibly due to a recent cultural barrier. Admixture is common in human evolution and exists in various populations worldwide.15,38,39 The local history indicates that the two populations differ in culture, language and religion. Previous studies have shown that these factors can be barriers that hinder genetic admixing.7 As such, the weak divergence observed could be because MUS amalgamate with the Malays, whereas BUD tend to mix with other Thai populations. Interestingly, the TH-South population from Shotivaranon et al.13 also displayed admixed origin. Nearly one-third of their genetic content was contributed by the Malay populations. TH-South samples were collected randomly from 211 Southern Thai individuals regardless of their ethnic affiliation. Therefore, genetic heterogeneity among individuals composing of BUD and MUS could be seen in current admixture analysis.
The studied populations displayed high heterozygosity for the 15 STR loci used. Only one locus deviated from HWE (D18S51 in MUS), indicating non-consanguineous marriages in both populations. The forensic parameters of this STR panel support previous findings that the loci are appropriate for forensic testing of Thai samples.12,13,15 Moreover, the loci reveal genetic relations of the two deep South populations. With the recent expansion of the CODIS loci from 13 to 20 loci40 and the availability of new 24-loci multiplexes (for example, GlobalFiler PCR Amplification kit), it is possible that we will be able to gain more insight into the genetic history of BUD and MUS. These newer loci have been selected due to their high heterozygosity.41 Thus, it might be possible to better differentiate the two populations. Additional information from more advanced techniques such as high throughput SNP genotyping platform, as well as the uniparentally-inherited loci (Y-STRs and mitochondrial DNA) should help elucidate the genetic relationships of the Thai populations.15,42,43
Our analyses using forensic loci on the genetic structure and affinity among BUD and MUS residing in the deep South of Thailand showed that (1) BUD and MUS exhibited genetic homogeneity due to common biological ancestry; (2) despite genetic similarity, some genetic divergence was noticeable. MUS are genetically closer to the Malaysian Malays, whereas BUD are more closely related to the other Buddhist populations in Thailand. Cultural barriers might have obstructed recent admixture between the studied populations. In contrast, admixture in short time-scale between MUS and Malaysian Malays could have been promoted by religious affiliation; and (3) cultural transformation and religious difference apparently contributed to the genetic structure of BUD and MUS.
References
National Statistical Office. The 2010 Population and Housing Census (Thai), Ministry of Information and Communication Technology: Bangkok, Thailand (2012).
National Statistical Office in Advance Report-The 2010 Population and Housing Census (Thai), National Statistical Office: Bangkok, Thailand, (2011).
Jampaklay, A., Sakulsri, T. & Prasith-hima, A.-A. in Proc. 8th National IPSR Conference: Marginalised Populations, Institute for Population and Social Research, Mahidol University (2012).
Schliesinger, J. Ethnic groups of Thailand: Non-Tai-speaking peoples, White Lotus Press: Chonburi, Thailand, (2000).
Syukri, I. History of the Malay Kingdom of Patani, Ohio University Press: USA, (1985).
Office of the Geographer. International Boundary Study: Malaysia - Thailand Boundary, Bureau of Intelligence and Research: USA, (1965).
Perez-Miranda, A.M., Alfonso-Sanchez, M.A., Kalantar, A., Garcia-Obregon, S., de Pancorbo, M.M., Pena, J.A. et al. Microsatellite data support subpopulation structuring among Basques. J. Hum. Genet. 50, 403–414 (2005).
Feigenblatt, O.F.V. The Muslim Malay Community in Southern Thailand: a ‘Small People’ Facing Existential Uncertainty. RJAPS 27, 53–63 (2010).
Ellegren, H. Microsatellite mutations in the germline: implications for evolutionary inference. Trends Genet. 16, 551–558 (2000).
Rosenberg, N.A., Pritchard, J.K., Weber, J.L., Cann, H.M., Kidd, K.K., Zhivotovsky, L.A. et al. Genetic structure of human populations. Science 298, 2381–2385 (2002).
Butler, J.M. Genetics and genomics of core short tandem repeat loci used in human identity testing. J. Forensic Sci. 51, 253–265 (2006).
Rerkamnuaychoke, B., Rinthachai, T., Shotivaranon, J., Jomsawat, U., Siriboonpiputtana, T., Chaiatchanarat, K. et al. Thai population data on 15 tetrameric STR loci-D8S1179, D21S11, D7S820, CSF1PO, D3S1358, TH01, D13S317, D16S539, D2S1338, D19S433, vWA, TPOX, D18S51, D5S818 and FGA. Forensic Sci. Int. 158, 234–237 (2006).
Shotivaranon, J., Chirachariyavej, T., Leetrakool, N. & Rerkamnuaychoke, B. DNA database of populations from different parts in the Kingdom of Thailand. Forensic Sci. Int. Genet. 4, e37–e38 (2009).
Listman, J.B., Malison, R.T., Sughondhabirom, A., Yang, B.Z., Raaum, R.L., Thavichachart, N. et al. Demographic changes and marker properties affect detection of human population differentiation. BMC Genet. 8, 21 (2007).
Kutanan, W., Kampuansai, J., Colonna, V., Nakbunlung, S., Lertvicha, P., Seielstad, M. et al. Genetic affinity and admixture of northern Thai people along their migration route in northern Thailand: evidence from autosomal STR loci. J. Hum. Genet. 56, 130–137 (2011).
Kutanan, W. & Kampuansai, J. Genetic variation of the Yuan in Saraburi province of central Thailand revealed by autosomal forensic STRs. Chiang Mai J. Sci. 41, 39–47 (2014).
Maruyama, S., Minaguchi, K., Takezaki, N. & Nambiar, P. Population data on 15 STR loci using AmpF/STR Identifiler kit in a Malay population living in and around Kuala Lumpur, Malaysia. Leg. Med. 10, 160–162 (2008).
Seah, L.H., Jeevan, N.H., Othman, M.I., Jaya, P., Ooi, Y.S., Wong, P.C. et al. STR Data for the AmpFlSTR Identifiler loci in three ethnic groups (Malay, Chinese, Indian) of the Malaysian population. Forensic Sci. Int. 138, 134–137 (2003).
Excoffier, L. & Lischer, H.E. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 10, 564–567 (2010).
Rice, W.R. Analyzing tables of statistical tests. Evolution 43, 223–225 (1989).
Peakall, R. & Smouse, P.E. GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes. 6, 288–295 (2006).
Excoffier, L., Smouse, P.E. & Quattro, J.M. Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131, 479–491 (1992).
Pritchard, J., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Falush, D., Stephens, M. & Pritchard, J.K. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164, 1567–1587 (2003).
Hubisz, M.J., Falush, D., Stephens, M. & Pritchard, J.K. Inferring weak population structure with the assistance of sample group information. Mol. Ecol. Resour. 9, 1322–1332 (2009).
Earl, D.A. & vonHoldt, B.M STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 4, 359–361 (2011).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol. Ecol. 14, 2611–2620 (2005).
Dupanloup, I. & Bertorelle, G. Inferring admixture proportions from molecular data: Extension to any number of parental populations. Mol. Biol. Evol. 18, 672–675 (2001).
Belle, E.M., Landry, P.A. & Barbujani, G. Origins and evolution of the Europeans' genome: evidence from multiple microsatellite loci. Proc. Biol. Sci. 273, 1595–1602 (2006).
Bertorelle, G. & Excoffier, L. Inferring admixture proportions from molecular data. Mol. Biol. Evol. 15, 1298–1311 (1998).
Latch, E.K., Dharmarajan, G., Glaubitz, J.C. & Rhodes, O.E. Relative performance of Bayesian clustering software for inferring population substructure and individual assignment at low levels of population differentiation. Conserv. Genet. 7, 295–302 (2006).
Kutanan, W., Kampuansai, J., Fuselli, S., Nakbunlung, S., Seielstad, M., Bertorelle, G. et al. Genetic structure of the Mon-Khmer speaking groups and their affinity to the neighbouring Tai populations in Northern Thailand. BMC Genet. 12, 56 (2011).
Azra, A. The Origins of Islamic Reformism in Southeast Asia: Networks of Malay-Indonesian and Middle Eastern 'Ulamā' in the Seventeenth and Eighteenth Centuries, Allen and Unwin: NSW, Australia (2004).
Eaaswarkhanth, M., Dubey, B., Ramakodi Meganathan, P., Noor, S. & Haque, I. Microsatellite diversity delineates genetic relationships of Shia and Sunni Muslim populations of Uttar Pradesh, India. Hum. Biol. 81, 427–445 (2009).
Eaaswarkhanth, M., Dubey, B., Meganathan, P.R., Ravesh, Z., Khan, F.A., Singh, L. et al. Diverse genetic origin of Indian Muslims: evidence from autosomal STR loci. J. Hum. Genet. 54, 340–348 (2009).
Peng, M.S., Quang, H.H., Dang, K.P., Trieu, A.V., Wang, H.W., Yao, Y.G. et al. Tracing the Austronesian footprint in Mainland Southeast Asia: a perspective from mitochondrial DNA. Mol. Biol. Evol. 27, 2417–2430 (2010).
Mishra, P.P. The History of Thailand, Greenwood (2010).
Pfaff, C.L., Parra, E.J., Bonilla, C., Hiester, K., McKeigue, P.M., Kamboh, M.I. et al. Population structure in admixed populations: effect of admixture dynamics on the pattern of linkage disequilibrium. Am. J. Hum. Genet. 68, 198–207 (2001).
Wen, B., Xie, X., Gao, S., Li, H., Shi, H., Song, X. et al. Analyses of genetic structure of Tibeto-Burman populations reveals sex-biased admixture in southern Tibeto-Burmans. Am. J. Hum. Genet. 74, 856–865 (2004).
Hares, D.R. Expanding the CODIS core loci in the United States. Forensic Sci. Int. Genet. 6, e52–e54 (2012).
Butler, J.M. & Hill, C.R. Biology and genetics of new autosomal STR loci useful for forensic DNA analysis. Forensic. Sci. Rev. 24, 15–26 (2012).
Bodner, M., Zimmermann, B., Rock, A., Kloss-Brandstatter, A., Horst, D., Horst, B. et al. Southeast Asian diversity: first insights into the complex mtDNA structure of Laos. BMC Evol. Biol. 11, 49 (2011).
Hatin, W.I., Nur-Shafawati, A.R., Zahri, M.K., Xu, S., Jin, L., Tan, S.G. et al. Population genetic structure of peninsular Malaysia Malay sub-ethnic groups. PLoS ONE 6 e18312 (2011).
Acknowledgements
This work is funded by the Faculty of Science Research Grant, Prince of Songkla University (no. 157001) and Khon Kaen University, Thailand, under the Incubation Researcher Project. We acknowledge the help of the DNA Analysis Center, Scientific Crime Detection Division 10, Royal Thai Police. We also thank Dr Daoroong Kangwanpong for valuable comments and suggestions on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Kutanan, W., Kitpipit, T., Phetpeng, S. et al. Forensic STR loci reveal common genetic ancestry of the Thai-Malay Muslims and Thai Buddhists in the deep Southern region of Thailand. J Hum Genet 59, 675–681 (2014). https://doi.org/10.1038/jhg.2014.93
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2014.93
This article is cited by
-
Reanalyzing the genetic history of Kra-Dai speakers from Thailand and new insights into their genetic interactions beyond Mainland Southeast Asia
Scientific Reports (2023)
-
Forensic and genetic characterizations of diverse southern Thai populations based on 15 autosomal STRs
Scientific Reports (2022)
-
Characterizing the diversity of MHC conserved extended haplotypes using families from the United Arab Emirates
Scientific Reports (2022)
-
Allelic frequencies of fifteen autosomal STRs in the northeastern Thai people
International Journal of Legal Medicine (2020)
-
Genetic structure of Tibetan populations in Gansu revealed by forensic STR loci
Scientific Reports (2017)