Abstract
With increasing public concern about infertility and the frequent involvement of chromosomal anomalies in miscarriage, analyses of copy number variations (CNVs) have been used to identify the genomic regions responsible for each process of childbearing. Although associations between CNVs and diseases have been reported, many CNVs have also been identified in healthy individuals. Like other types of mutations, phenotypically indefinite CNVs may have been retained and accumulated during anthropogenesis. Therefore to distinguish causative variants from other variants is a formidable task. Furthermore, because previous studies have predominantly focused on European and African populations, comprehensive detection of common Asian CNVs is eagerly awaited. Here, using a high-resolution genotyping array and samples from 411 Japanese women with normal parity without significant complications, we have compiled 1043 copy number variable regions. In total, the collected regions cover 164 Mb, or up to 0.5% of the genome. The copy number differences in these regions may be irrelevant not only to infertility but also to a wide range of diseases. The utility of this resource in reducing the candidate pathogenetic variants, especially in Japanese subjects, is also demonstrated.
Similar content being viewed by others
Introduction
The advent of new technologies has allowed the identification of structural variants that have a more significant impact on human diversity than does the entire set of single-nucleotide polymorphisms (SNPs). Copy number variations (CNVs) are one such type of structural variant and constitute the largest proportion of genomic variations.1, 2, 3 CNVs result from the duplication or deletion of a DNA segment and are commonly observed in human genomes.4, 5, 6, 7 When a genomic event results in a CNV, not only the copy number of a gene can be altered but also its genic sequences. Therefore, CNVs can cause disease or contribute to disease susceptibility,8, 9, 10 and they have been compiled in several databases for public use.9, 10, 11
Although a number of deleterious changes may have been negatively selected during human evolution, it is likely that phenotypically neutral changes have been retained, transmitted and accumulated over generations. Increasing numbers of CNVs are found in phenotypically normal human individuals.1 Accordingly, each ethnic group tends to have distinct features in terms of the positions, copy numbers and frequencies of their CNVs, and it is possible that fixed CNVs have contributed to ethnic differences in phenotypic variations and disease susceptibility.12, 13, 14, 15 Therefore, it is important to have a list of CNVs for each ethnic group, especially for medical purposes. However, the number of reported CNVs from Asian populations is small compared with those of Europeans and Africans. Extensive examination of Asian CNVs is eagerly awaited by Asian researchers.
The compilation of nonpathogenic variations, in addition to disease-related variations, is also important for a better understanding of the genetic landscape of the human genome. Data sets including both these sorts of variations should be helpful in pinpointing causative mutations. Even when we search for variations using patient samples, most of the variations identified would be normal polymorphisms, together with a few pathogenic mutations. Although we can consider most of the available variation data nonpathogenic, it is difficult to know which variations are pathogenic. Therefore, the collection of data from normal controls is essential. To investigate phenotypically ‘normal’ samples in this study, we considered reproduction and child development, and chose parous Japanese women, who had experienced normal pregnancies and deliveries.
Although the origin of the Japanese population remains controversial, the last major migration to the Japanese Archipelago is thought to have occurred approximately 2000 years ago.16, 17 The population has been mixed well with various Asian ethnic groups during previous migrations, but has remained relatively isolated for 2000 years. However, although the current population of Japan is 127 million, far fewer CNVs have been documented in Japanese samples than in Europeans. Compiling a list of Japanese CNVs is also important from the perspective of medical science in Japan.
Materials and methods
Subject recruitment and SNP genotyping with a high-resolution microarray
We examined 411 unrelated Japanese women who had had one or more normal parities, with no significant abnormalities in any of their pregnancies, deliveries or neonates. Ethical approval was also obtained from each review board of the hospitals that participated in the study. The informed consent of all the subjects was obtained. To avoid cell-culture-induced chromosomal rearrangements, genomic DNAs were extracted directly from blood using the QIAsymphony DNA Midi Kit (Qiagen, Venlo, The Netherlands) with the QIAsymphony SP instrument and analyzed with a high-resolution SNP-based genotyping microarray, HumanOmni2.5-8 BeadChip (Illumina, San Diego, CA, USA). Only data that met the quality control guidelines of the manufacturer were used for further analyses.
Identification of CNVs and CNVRs
Two distinct algorithms were used to maximize the specificity of our CNV calling: a likelihood-based method with CNVPartition version 3.2.0 (http://www.illumina.com/software/illumina_connect.ilmn) and a hidden Malkov method with PennCNV version (27 August 2009).18 The parameters applied with these tools were referred to those typically used by many research groups (at least three consecutive probes to define a CNV, using the GC wave adjustment option, etc.). These programs computed confidence scores that can be used to filter out CNV regions that are likely to be false positives. However, we should note that the two programs calculated the scores in different ways, with different scales. To minimize false positives, we first chose only CNVs with high confidence scores; that is, more than 100 with CNVPartition, and selected copy number variable regions (CNVRs) that overlapped those called by PennCNV for at least 80% of their lengths. For PennCNV, we generated a list of B allele frequencies using a collection of signal intensities for 47 samples from HapMap Japanese in Tokyo with the compile_pfb script (Figure 1a).
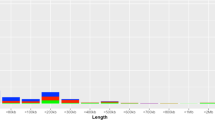
(a) Data processing flow. The initial 26 150 regions identified with CNVPartition were validated with PennCNV. (b) Chromosomal distribution of the CNVRs. Each CNVR is shown by a horizontal bar. Gain- and loss-type CNVs are distinguished by bars on the left and right, respectively. The numbers of each type of CNV are also shown, and are drawn with Idiographica (http://www.ncrna.org/idiographica/). CNV, copy number variation; CNVR, copy number variable regions.
Multiplex PCR assay
Multiplex polymerase chain reaction (PCR) assay was used to confirm regions that had been called homozygously deleted. The reactions were performed with both a control primer pair that generated a 296-bp fragment and a test primer pair that amplified a target region. The thermal cycling conditions were initial denaturation at 95 °C for 2 min, followed by 35 cycles of denaturation at 95 °C for 30 s, annealing at 60 °C for 30 s and extension at 72 °C for 30 s, and a final extension at 72 °C for 3 min. Detailed information on these primers is given in Supplementary Table S3.
Results
Genetic ancestry of the subjects
First, the population structure was inferred with the Structure software (http://pritchardlab.stanford.edu/structure.html) to confirm the Japanese ancestry of the subjects.19 A cluster analysis of our samples together with the sequences of 499 HapMap individuals from three ancestral populations (European, African and Asian) was performed using 1959 unlinked tag SNPs on chromosome 21. The expected ancestry of all the subjects was confirmed with a minimum coefficient of 0.85. We also performed a principal components analysis with the pca.jar program (Biobank Japan project; http://genome-analysis.src.riken.jp/PCP/). The results indicated that all but one subject were derived from the main islands of Japan and that the remaining singleton was Ryukyuan.20
Characterization of CNVs and CNVRs
The CNVPartition software (Illumina) identified 26 150 candidate regions as CNVs. We then used another program, PennCNV,18 which is based on an integrated hidden Markov algorithm, to maximize the specificity of the analysis. If a candidate CNV was also supported by PennCNV for at least 80% of its length, it was retained. In this way, we identified 6871 CNVs and 1043 regions with variable copy numbers from 411 Japanese individuals, with an average of 16.7 CNVs per diploid genome (Supplementary Table S1). Detailed information on all the SNP probes used for the CNV calls is tabulated (Supplementary Table S2). The mean length of the CNVs was 79.9 kb, ranging from 169 bases to 2.27 Mb. These 6871 CNVs corresponded to 1043 CNVRs (588 losses and 455 gains). Figure 1 shows the chromosomal distribution of the observed CNVRs. The total length of all of these CNVRs was 163 720 kb, which is equivalent to 0.5% of the whole human genome. The CNVRs can be divided into gain regions and loss regions, depending on whether their copy numbers have increased or decreased. Of the 1043 regions identified, 1033 overlap the latest database of genomic variants (DGVs) (released on 23 July 2013) reported at the DGV. More than half the CNVRs, including 72% of the gain CNVRs and 36% of the loss CNVRs, intersect RefSeq gene loci.
As far as we know, three studies have examined the Japanese population with array-based methods: two of them used samples from HapMap and the other used healthy individuals.21, 22, 23 These results are summarized with our data set (Table 1). Although those three studies had together already reported 82 regions, more than half the regions reported in the present study were not detected by them. It is probable that the higher resolution of our analysis and our larger sample size allowed us to detect additional CNVRs. Depending on the types of platform used, array-based CNV studies occasionally show discrepancies in the regions of CNVs.21, 22 Differences in the array architectures, scanning machines and calling algorithms could affect the final data sets. Using reported CNV data from SNP arrays, we counted the overlapping regions among studies that focused on other populations or HapMap data13, 14, 24, 25, 26 (Table 2). The similarities among these studies are comparable, but our results suggest a greater similarity between the Japanese and Chinese populations.
Homozygous deletions found in parous Japanese women
In our study, 1628 homozygous deletions that could affect 112 RefSeq gene loci were called in a total of 822 chromosomes. Although the CNV analysis was unable to determine the precise breakpoints, our data indicate that some exonic sequences are disrupted by homozygous deletions (Table 3). Using multiplex PCR with both control and test primer pairs, we confirmed the null genotypes caused by deletions (Supplementary Figure S1 and Supplementary Table S3). Five genes, FCGR3B, FCGR2B, UGT2B17, HLA-DRB1 and CYP2A6, are described as disease related in the OMIM database. The FCGR3, FCGR2B and HLA-DRB1 genes have roles in the immune system. FCGR3B and FCGR2B encode the crystallizable region of immunoglobulin G. Several studies have shown that a low copy number at the FCGR3B–FCGR2B locus is associated with a susceptibility to systemic lupus erythematosus in the Caucasian population,27, 28, 29 but not in the Chinese population.28 UGT2B17 encodes a protein that belongs to the family of UDP-glucuronosyltransferases enzymes, which catalyzes the glucuronidation of steroid hormones. A case–control study of osteoporosis-related fracture suggested that a CNV at the UGT2B17 locus contributes to osteoporosis.30 Jakobsson et al.31 found that its null genotype was more common in Koreans (67%) than in Swedish (9%). Our array results also showed a high frequency (74%) of the null genotype. The CYP2A6 protein metabolizes nicotine and coumarin in the liver. The lack of a CYP2A6 gene may affect nicotine levels in individuals and probably has a protective effect against tobacco dependence.32 Another study reported that the frequency of homozygotes for the CYP2A6 gene deletion was lower in Japanese lung cancer patients than in control samples.33 Except for HLA-DRB1, these disease-related genes have been reported to be frequently deleted in Asian populations.25, 34, 35, 36 Because we limited our samples to parous women only, it is unlikely that the CNVRs identified in the present study are related to human reproduction.
Discussion
In the present study, we compiled a catalog of copy number variable regions identified in phenotypically normal Japanese samples, especially those with a history of full-term pregnancy and deliveries without major complications. The data set will be useful in the search for novel or rare CNVs that increase the individual’s susceptibility to congenital diseases and complications during pregnancy. It is unlikely that the newly identified CNVs are related to infertility or miscarriage. CNVs in parous women without complications have never before been investigated. Although the copy numbers of these regions were not thoroughly validated with other methods; such as, quantitative PCR, according to DGV, most of the CNVRs identified here have been reported in previous studies, indicating that they should be observed by other methods or techniques. Because our identification strategy was based on a microarray technique, it is inevitable that errors would have occurred. Besides routine data processing, we also carefully curated the data by examining the B allele frequencies and signal intensities (log R ratio) for each CNVR using the GenomeStudio software (Illumina) (Figure 2). We found that many implausible calls were situated in regions with high G+C contents; for example, in subtelomeric regions. All of them were copy number gain-type CNVs rather than copy number loss-type CNVs. Although further research is required, it is important to note that CNVRs tend to be detected in those regions by SNP microarrays. Even if such CNVRs are false positives, our data set is still useful for screening large numbers of candidate CNVs.
(a) A copy number variation (CNV) located on chromosome 7. The panel shows the region at nucleotides 108 541 155–108 698 960 in hg19. This CNV was called with high-intensity probes. The B allele frequencies (BAFs) were separated into four levels, which corresponded to AAA, AAB, ABB and BBB, respectively. (b) Another CNV located in the subtelomeric region on chromosome 4. The panel shows the region at nucleotides 863 513–1 113 194. Despite high-intensity probes used, as in the example shown above, the four levels of BAFs were not observed, suggesting that the call might be implausible. Such CNVs tended to be called in G+C-rich regions; for example, 58% G+C content in this case. The snapshot was made with the IGV program (http://www.broadinstitute.org/igv/). A full color version of this figure is available at Journal of Human Genetics online.
It is unclear whether CNVs are selectively neutral on the basis of genetic drift, but they are certainly distributed throughout all human populations. Using the genotypes of mitochondrial DNA and Y chromosome, geneticists and anthropologists have surmised various intriguing scenarios about the history of humans.37, 38, 39, 40 However, these genetic materials have been transmitted exclusively through maternal and paternal lineages, respectively. In contrast, the CNVs reported here occur in the more extensive remaining genome regions; that is, on autosomes or the X chromosome. Therefore, they have acted some times as maternal alleles at and at other times as paternal alleles. They might also have been subjected to crossingover. CNV data from various parts of the world are essential to substantiate these hypothetical scenarios.
Chromosomal anomalies are found with conventional cytogenetic techniques in approximately half of all early sporadic miscarriages.41 It is possible that miscarriages and pregnancy losses are also caused by submicroscopic chromosomal changes, including CNVs. Twenty-eight CNVs have been reported as candidate miscarriage-related variations when instances of recurrent pregnancy loss were examined by Rajcan-Separovic et al.42 When 17 Caucasian and three African-American couples with recurrent pregnancy losses and their miscarriage samples were examined, CNVs that may have been related to miscarriages were reported.42 They reported 11 novel CNVs in miscarriage samples and three in the parent samples and suggested that these CNVs were probably mutations causing susceptibility to miscarriage. Of the 11 CNVs in the miscarriage samples, one on chromosome 12 (130 060 706–130 430 847 in hg18) and another one on chromosome X (6 498 521–8 091 951) overlapped with our data set. Whereas the first one on chromosome 12 was up to 370 kb in length and encompassed the GPR133 gene, the corresponding variable region in our data set is much shorter and includes no known genes. The GPR133 gene encodes one of the orphan G-protein-coupled receptors, but its function is unknown.43 It is possible that this receptor protein has a role in several signal-transduction pathways via classical receptor/G-protein interactions. Therefore, the CNV mentioned above may be a variant that causes miscarriage. However, one of the CNVs on chromosome X is consistent with our data set, suggesting that it is a commonly observed variant. In fact, Rajcan-Separovic et al.42 tried to define the common CNVs using a collective repository in the DGV, but insufficient phenotypic information was available to refine the data. Taking these observations together, it seems that to define a set of common CNVs, it will be necessary to collect a large number of control data that focus on a specific phenotype; such as, normal parity in this case.
The Japanese are an admixture of ancient Asian populations that inhabited regions outside the Japanese Archipelago. We investigated the similarities among the CNVRs detected in various populations and noted that around 15% of Japanese CNVRs overlap those of other populations (Table 2). It has been suggested that the number of overlapping CNVs is influenced by the number of subjects. For instance, Japanese and Tibetan data showed dissimilarity because of the limited number of Tibetan subjects. Although the sample sizes of the Korean and Chinese populations are smaller than those of the European and African populations, similarities between the Japanese and other East Asian populations were similar to those of the European and African populations. This probably suggests strong similarities between the Japanese and other East Asian populations.
Previous studies have predominantly targeted European and African populations, but CNVs have been observed at different frequencies or copy numbers in different populations; for example, variations in the salivary amylase gene.44 Many CNVs; such as, those at the AMY1 locus, may be associated with diabetes, asthma, hypertension, allergy and other diseases of affluence in each ethnic group. Although CNVRs may result from the accumulation of tolerable structural mutations in the course of an ethnic history, they could start to influence the population’s susceptibility to disease once its lifestyle is altered. The allelic frequencies of SNPs and short indels in each population have recently been documented.45 The complete documentation of the CNVRs in each ethnic group is similarly important. The development of an innovative method to achieve this; such as, one involving next-generation sequencing and informatics, is another challenge.
References
Redon, R., Ishikawa, S., Fitch, K. R., Feuk, L., Perry, G. H., Andrews, T. D. et al. Global variation in copy number in the human genome. Nature 444, 444–454 (2006).
Stranger, B. E., Forrest, M. S., Dunning, M., Ingle, C. E., Beazley, C., Thorne, N. et al. Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 315, 848–853 (2007).
Pang, A. W., Migita, O., Macdonald, J. R., Feuk, L. & Scherer, S. W. Mechanisms of formation of structural variation in a fully sequenced human genome. Hum. Mut. 34, 345–354 (2013).
Frazer, K. A., Chen, X., Hinds, D. A., Pant, P. V., Patil, N. & Cox, D. R. Genomic DNA insertions and deletions occur frequently between humans and nonhuman primates. Genome Res. 13, 341–346 (2003).
Locke, D. P., Segraves, R., Carbone, L., Archidiacono, N., Albertson, D. G., Pinkel, D. et al. Large-scale variation among human and great ape genomes determined by array comparative genomic hybridization. Genome Res. 13, 347–357 (2003).
Sebat, J., Lakshmi, B., Troge, J., Alexander, J., Young, J., Lundin, P. et al. Large-scale copy number polymorphism in the human genome. Science 305, 525–528 (2004).
Sharp, A. J., Locke, D. P., McGrath, S. D., Cheng, Z., Bailey, J. A., Vallente, R. U. et al. Segmental duplications and copy-number variation in the human genome. Am. J. Hum. Genet. 77, 78–88 (2005).
Marshall, C. R. & Scherer, S. W. Detection and characterization of copy number variation in autism spectrum disorder. Methods Mol. Biol. 838, 115–135 (2012).
Swaminathan, G. J., Bragin, E., Chatzimichali, E. A., Corpas, M., Bevan, A. P., Wright, C. F. et al. DECIPHER: web-based, community resource for clinical interpretation of rare variants in developmental disorders. Hum. Mol. Genet. 21, R37–R44 (2012).
Firth, H. V., Richards, S. M., Bevan, A. P., Clayton, S., Corpas, M., Rajan, D. et al. DECIPHER: Database of Chromosomal Imbalance and Phenotype in Humans Using Ensembl Resources. Am. J. Hum. Genet. 84, 524–533 (2009).
Macdonald, J. R., Ziman, R., Yuen, R. K., Feuk, L. & Scherer, S. W. The Database of Genomic Variants: a curated collection of structural variation in the human genome. Nucleic Acids Res. 42, D986–D992 (2014).
Li, J., Yang, T., Wang, L., Yan, H., Zhang, Y., Guo, Y. et al. Whole genome distribution and ethnic differentiation of copy number variation in Caucasian and Asian populations. PLoS ONE 4, e7958 (2009).
Lou, H., Li, S., Yang, Y., Kang, L., Zhang, X., Jin, W. et al. A map of copy number variations in Chinese populations. PLoS ONE 6, e27341 (2011).
Zhang, Y. B., Li, X., Zhang, F., Wang, D. M. & Yu, J. A preliminary study of copy number variation in Tibetans. PLoS ONE 7, e41768 (2012).
Kanduri, C., Ukkola-Vuoti, L., Oikkonen, J., Buck, G., Blancher, C., Raijas, P. et al. The genome-wide landscape of copy number variations in the MUSGEN study provides evidence for a founder effect in the isolated Finnish population. Eur. J. Hum. Genet. 21, 1411–1516 (2013).
Hanihara, K. Dual structure model for the population history of the Japanese. Jpn Rev. 2, 1–33 (1991).
Japanese Archipelago Human Population Genetics C., Jinam, T., Nishida, N., Hirai, M., Kawamura, S., Oota, H. et al. The history of human populations in the Japanese Archipelago inferred from genome-wide SNP data with a special reference to the Ainu and the Ryukyuan populations. J. Hum. Genet. 57, 787–795 (2012).
Wang, K., Li, M., Hadley, D., Liu, R., Glessner, J., Grant, S. F. et al. PennCNV: an integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res. 17, 1665–1674 (2007).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Kumasaka, N., Yamaguchi-Kabata, Y., Takahashi, A., Kubo, M., Nakamura, Y. & Kamatani, N. Establishment of a standardized system to perform population structure analyses with limited sample size or with different sets of SNP genotypes. J. Hum. Genet. 55, 525–533 (2010).
McCarroll, S. A., Kuruvilla, F. G., Korn, J. M., Cawley, S., Nemesh, J., Wysoker, A. et al. Integrated detection and population-genetic analysis of SNPs and copy number variation. Nat. Genet. 40, 1166–1174 (2008).
Conrad, D. F., Pinto, D., Redon, R., Feuk, L., Gokcumen, O., Zhang, Y. et al. Origins and functional impact of copy number variation in the human genome. Nature 464, 704–712 (2010).
Koike, A., Nishida, N., Yamashita, D. & Tokunaga, K. Comparative analysis of copy number variation detection methods and database construction. BMC Genet. 12, 29 (2011).
Moon, S., Kim, Y. J., Hong, C. B., Kim, D. J., Lee, J. Y. & Kim, B. J. Data-driven approach to detect common copy-number variations and frequency profiles in a population-based Korean cohort. Eur. J. Hum. Genet. 19, 1167–1172 (2011).
Vogler, C., Gschwind, L., Rothlisberger, B., Huber, A., Filges, I., Miny, P. et al. Microarray-based maps of copy-number variant regions in European and sub-Saharan populations. PLoS ONE 5, e15246 (2010).
Shaikh, T. H., Gai, X., Perin, J. C., Glessner, J. T., Xie, H., Murphy, K. et al. High-resolution mapping and analysis of copy number variations in the human genome: a data resource for clinical and research applications. Genome Res. 19, 1682–1690 (2009).
Fanciulli, M., Norsworthy, P. J., Petretto, E., Dong, R., Harper, L., Kamesh, L. et al. FCGR3B copy number variation is associated with susceptibility to systemic, but not organ-specific, autoimmunity. Nat. Genet. 39, 721–723 (2007).
Willcocks, L. C., Lyons, P. A., Clatworthy, M. R., Robinson, J. I., Yang, W., Newland, S. A. et al. Copy number of FCGR3B, which is associated with systemic lupus erythematosus, correlates with protein expression and immune complex uptake. J. Exp. Med. 205, 1573–1582 (2008).
McKinney, C. & Merriman, T. R. Meta-analysis confirms a role for deletion in FCGR3B in autoimmune phenotypes. Hum. Mol. Genet. 21, 2370–2376 (2012).
Yang, T. L., Chen, X. D., Guo, Y., Lei, S. F., Wang, J. T., Zhou, Q. et al. Genome-wide copy-number-variation study identified a susceptibility gene, UGT2B17, for osteoporosis. Am. J. Hum. Genet. 83, 663–674 (2008).
Jakobsson, J., Ekström, L., Inotsume, N., Garle, M., Lorentzon, M., Ohlsson, C. et al. Large differences in testosterone excretion in Korean and Swedish men are strongly associated with a UDP-glucuronosyl transferase 2B17 polymorphism. J. Clin. Endocrinol. Metab. 91, 687–693 (2006).
Pianezza, M. L., Sellers, E. M. & Tyndale, R. F. Nicotine metabolism defect reduces smoking. Nature 393, 750 (1998).
Miyamoto, M., Umetsu, Y., Dosaka-Akita, H., Sawamura, Y., Yokota, J., Kunitoh, H. et al. CYP2A6 gene deletion reduces susceptibility to lung cancer. Biochem. Biophys. Res. Commun. 261, 658–660 (1999).
Lv, J., Yang, Y., Zhou, X., Yu, L., Li, R., Hou, P. et al. FCGR3B copy number variation is not associated with lupus nephritis in a Chinese population. Lupus 19, 158–161 (2010).
Xue, Y., Sun, D., Daly, A., Yang, F., Zhou, X., Zhao, M. et al. Adaptive evolution of UGT2B17 copy-number variation. Am. J. Hum. Genet. 83, 337–346 (2008).
Oscarsona, M., McLellana, R. A., Gullsteènb, H., Yuec, Q. Y., Langd, M. A., Bernale, M. L. et al. Characterisation and PCR-based detection of a CYP2A6 gene deletion found at a high frequency in a Chinese population. FEBS Lett. 448, 105–110 (1999).
Hammer, M. F. & Horai, S. Y chromosomal DNA variation and the peopling of Japan. Am. J. Hum. Genet. 56, 951–962 (1995).
Shinka, T., Tomita, K., Toda, T., Kotliarova, S. E., Lee, J., Kuroki, Y. et al. Genetic variations on the Y chromosome in the Japanese population and implications for modern human Y chromosome lineage. J. Hum. Genet. 44, 240–245 (1999).
Dulik, M. C., Zhadanov, S. I., Osipova, L. P., Askapuli, A., Gau, L., Gokcumen, O. et al. Mitochondrial DNA and Y chromosome variation provides evidence for a recent common ancestry between Native Americans and Indigenous Altaians. Am. J. Hum. Genet. 90, 229–246 (2012).
Oppenheimer, S. Out-of-Africa, the peopling of continents and islands: tracing uniparental gene trees across the map. Philos. Trans. R. Soc. Lond. Ser. B 367, 770–784 (2012).
van den Berg, M., van Maarle, M., van Wely, M. & Goddijn, M. Genetics of early miscarriage. Biochim. Biophys. Acta 1822, 1951–1959 (2012).
Rajcan-Separovic, E., Diego-Alvarez, D., Robinson, W. P., Tyson, C., Qiao, Y., Harvard, C. et al. Identification of copy number variants in miscarriages from couples with idiopathic recurrent pregnancy loss. Hum. Reprod. 25, 2913–2922 (2010).
Bohnekamp, J. & Schoneberg, T. Cell adhesion receptor GPR133 couples to Gs protein. J. Biol. Chem. 286, 41912–41916 (2011).
Perry, G. H., Dominy, N. J., Claw, K. G., Lee, A. S., Fiegler, H., Redon, R. et al. Diet and the evolution of human amylase gene copy number variation. Nat. Genet. 39, 1256–1260 (2007).
1000 Genomes Project Consortium, Abecasis, G. R., Auton, A., Brooks, L. D., DePristo, M. A., Durbin, R. M. et al. An integrated map of genetic variation from 1092 human genomes. Nature 491, 56–65 (2012).
Acknowledgements
We are grateful to all the participants in the present study, including the 411 women. Computation time was partly provided by the supercomputer system, Shirokane, at the Human Genome Centre, Institute of Medical Science, University of Tokyo. This work was supported by CREST Program ‘Epigenomic analysis of the human placenta and endometrium constituting the fetal-maternal interface’ of Japan Science and Technology Agency (JST) and Health and Labor Sciences Research Grants for Research into Rare and Intractable Diseases (H23 Jitsuyoka (Nanbyo)-Ippan-003 and H25 Jisedai-Ippan-001), and was also partly supported by KAKENHI 23770273, 24657151, 24390251, 24592494, 24659742, 25293345 and 25860258.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors received no financial support from Illumina KK and the company had no role in the study design. The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Rights and permissions
About this article
Cite this article
Migita, O., Maehara, K., Kamura, H. et al. Compilation of copy number variants identified in phenotypically normal and parous Japanese women. J Hum Genet 59, 326–331 (2014). https://doi.org/10.1038/jhg.2014.27
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2014.27