Abstract
Familial tumor suppressor genes comprise two subgroups: caretaker genes (CTs) that repair DNA, and gatekeeper genes (GKs) that trigger cell death. Since GKs may also induce cell cycle delay and thus enhance cell survival by facilitating DNA repair, we hypothesized that the prosurvival phenotype of GKs could be selected during cancer progression, and we used a multivariable systems biology approach to test this. We performed multidimensional data analysis, non-negative matrix factorization and logistic regression to compare the features of GKs with those of their putative antagonists, the proto-oncogenes (POs), as well as with control groups of CTs and functionally unrelated congenital heart disease genes (HDs). GKs and POs closely resemble each other, but not CTs or HDs, in terms of gene structure (P<0.001), expression level and breadth (P<0.01), DNA methylation signature (P<0.001) and evolutionary rate (P<0.001). The similar selection pressures and epigenetic trajectories of GKs and POs so implied suggest a common functional attribute that is strongly negatively selected—that is, a shared phenotype that enhances cell survival. The counterintuitive finding of similar evolutionary pressures affecting GKs and POs raises an intriguing possibility: namely, that cancer microevolution is accelerated by an epistatic cascade in which upstream suppressor gene defects subvert the normal bifunctionality of wild-type GKs by constitutively shifting the phenotype away from apoptosis towards survival. If correct, this interpretation would explain the hitherto unexplained phenomenon of frequent wild-type GK (for example, p53) overexpression in tumors.
Similar content being viewed by others
Introduction
Somatic tumorigenesis involves loss-of-function events affecting a heterogeneous class of regulators designated tumor suppressor genes (TSGs), which are sometimes also termed as recessive oncogenes to distinguish them from their dominant transforming (gain-of-function, proto-oncogene; PO) counterparts.1 Two functional TSG subgroups are hence defined: gatekeeper genes (GKs) such as RB1 and TP53, which control cell cycling and cell death, and caretaker genes (CTs) such as BRCA1 and MLH1, which maintain DNA repair and genome stability.2 Loss-of-function mutations often affect GKs such as TP53 and others in sporadic tumors,3, 4 consistent with an important role for GKs in regulating clonal tumor proliferation. In contrast, we have shown in earlier work that CT dysfunction, which could be subtle, polymorphic and/or multiallelic5—is better tolerated in the germline than GK dysfunction is, implicating environmental selection for germline CT gene methylation as a mechanism that may contribute both to maintenance of genome plasticity for the species, and also to heritable but non-familial cancer predisposition.6
Frequent occurrence in sporadic tumors of TSG protein overexpression without gene mutation—as has been reported for TP53,7 CDKN2A,8 RB1,9 SMAD410 and CDH1,11 among others—with tumor-promoting effects is less understood.12, 13 As confirmed by literature data mining,14 such overexpression is more strongly associated with GKs rather than with CTs (P<0.001; Supplementary Table S1); this suggests that GKs express a selectable phenotype, which is lacking in CTs. Relevant to this hypothesis, we and others have shown that certain POs (such as MYC and HER2) can promote anti-oncogenic cell phenotypes such as differentiation,15 growth arrest16 or apoptosis in cells with relatively normal cell cycle control machinery;17 in contrast, when selected for ‘cooperative’ GK knockout or preexisting constitutive PO expression (for example, of the RAS family18), overexpression of heterologous POs causes growth and/or transformation.19, 20, 21, 22 These observations suggest that a dynamic balance exists between expression thresholds of opposing genes within the same network,23 and that abnormal perturbation of one part of the network could possibly select for normal GK hyperfunction.24
Such apparent bifunctionality may be explained by epistasis, a concept, which teaches that the function of a given gene is not invariant but rather depends upon the genetic background.25 In the above example, a possible explanation is that GK protein overexpression may be not only tolerated but also selected in the presence of a block affecting heterologous TSGs.26, 27 Consistent with this, CT inactivation is a common tumorigenetic event predisposing to GK mutation;28 for example, repair deficiency (for example, microsatellite instability caused by mismatch repair gene inactivation) may lead to secondary mutations (for example, frameshifts) affecting GKs or POs in preinvasive tissues.29, 30 Moreover, as repair genes are involved in afferent sensing of DNA damage,31, 32 CT inactivation in tumors could also bias GK function towards a pro-mitogenic phenotype by raising the apoptotic threshold.33
Systems biology is an emerging field of interdisciplinary research that seeks to elucidate complex biological interactions by integrating otherwise disparate data sources; it is not a single well-defined field, but is a dynamic set of experimental approaches that seek to clarify central functional patterns within systems by integrating multiple information sources.34 Such an approach is increasingly used in which the description of a complex set of discrete observations fails to explain the behavior of the whole system, such as in the example above of suppressor gene overexpression in sporadic cancer. The types of techniques often used in systems biology include transcriptomics, genomics, methylomics, metabolomics and similar structure–function analytical techniques.
The present study applies the systems biology approach by using a variety of such analyses for each of the four gene groups, including gene expression (transcription) analysis, gene structure (genomic) analysis, methylation-dependent ‘signature’ (methylation and mutation) analysis and evolutionary rate (evolvability) analysis. To test the notion that GKs and POs may share functional attributes depending upon the genetic environment—that is, with wild-type alleles of both the gene families maintaining differentiation clonal outgrowth in cells with heterologous suppressor gene defects—we compare and distinguish the structural and functional features of GKs, CTs, POs and a control group of genes implicated in the pathogenesis of congenital heart disease (HD). As detailed here, this analysis provides the counter-intuitive conclusion that GKs share the genomic and evolutionary properties of POs, raising the possibility of inapparent phenotypic similarities relevant to carcinogenesis.
Materials and methods
Gene identification, classification and ontology analysis
We chose 157 genes (39 CTs, 36 GKs, 41 POs, 41 HDs; Supplementary Tables S1 and S5) from published works (http://www.sanger.ac.uk/genetics/CGP/Census).35, 36 Familial cancer genes with repair functions37, 38 were categorized as CTs, whereas other familial suppressors39—most of which were confirmed to mediate apoptosis40—were classified as GKs. In addition, only putative POs with transforming viral homologs were classed as POs. Given the multigenic interdependence of DNA repair and cellular apoptosis,33 unambiguous identification of genes that exclusively mediate one of these two processes is not straightforward. We sought to minimize uncertainties over this functional overlap in two ways: first, we used a familial tumor suppressor gene database35, 41 to restrict the choice of genes to those with major neoplastic effects (that is, heritable cancer syndromes) when deleted in the germline; this provided a total of 75 TSGs; and second, by cross-correlating the former data set with a database of DNA repair genes38 we subclassified this familial cancer susceptibility gene subset as CTs (n=39), then designated the remainder—the majority of which were confirmed to mediate apoptosis40—as GKs (n=36). An additional control group of heart development genes implicated in congenital HDs was also defined using the NCBI Entrez Gene (27 September 2009 release) and keyword search on ‘congenital heart disease’ and ‘Homo sapiens’. Gene ontology analysis42 was performed using the Panther database (http://www.pantherdb.org/).43
Analyses of gene sequences and mutations, and non-negative matrix factorization
We used sequence analysis to infer functional properties of gene sequences as in our previously cited publications. Briefly, enzymatic methylation of cytosines in CpG dinucleotides clustered within gene promoters leads to transcriptional repression and chromatin condensation, whereas methylcytosine residues in coding regions may undergo oxidative deamination to form thymine residues; if the mismatch repair system fails to rectify these mutations, an excess of CG → TA transitional mutations becomes a quantifiable hallmark of the foregoing methylation events.44 This interaction between methylation-dependent trans repression and mutation is in turn a factor of positive selection,45 perhaps contributing to the nonrandom correlations between the codon structure and function that we have previously reported.46, 47 Inter-species changes in genomic GC content48 could also derive, in part, from such a mutational mechanism. Nucleotide sequences may similarly be examined for the presence or otherwise of strand-specific (sense vs antisense) dinucleotide asymmetries,49 whereas transcription-coupled repair creates asymmetric patterns of base composition that can be mined to support retrospective inferences of differential transcription.50 Human and mouse reference sequences, and species gene numbers were downloaded from NCBI Gene (http://www.ncbi.nlm.nih.gov/Gene), whereas mutation data were downloaded from the Human Gene Mutation Database. A variety of packages from R 2.81 (http://www.r-project.org) were used for statistical analysis, including coin, biomaRt, GeneR, nlmc and others. To analyze coding sequences, we used reference data from the NCBI Entrez Gene or ENSEMBL and updated using R scripts. For multiple splicing forms, the longest coding sequence was used for analysis; mono- and dinucleotide composition was assessed using in-house Perl scripts and/or GeneR package in R2.81. Comparison of mutation rates in germline (that is, familial cancers) and somatic (sporadic) tumors was based on Cancer Genome Anatomy Project Gene Census list (http://www.sanger.ac.uk/genetics/CGP/Census).35, 36 Frame-dependent dinucleotide composition and asymmetries were analyzed using the GeneR package. For comparative analyses of 5′-and 3′-untranslated regions (UTR), reference sequences were downloaded from ENSEMBL (Release 52) using R package biomaRt. Non-negative matrix factorization (NNMF) was performed using MATLAB (7.6) Statistical Toolbox (5.1) (http://www.mathworks.com) for principal component analyses, and non-parametric tests.
Analysis of evolutionary rate
The latest gene evolutionary rate data was downloaded from ENSEMBL (release 52) using biomaRt packages running in R platform. Evolutionary rates were computed with maximum likelihood method using the PAML packages. Gene expression intensity and breadth (specificity) were related to molecular evolutionary rates as approximated by the ratio of non-synonymous to synonymous mutation rate (Ka/Ks or dN/dS, in which the former is treated as non-neutral whereas the latter is treated as neutral).51
Gene expression analysis
We mined reference data from the University of California Santa Cruz genome database, then analyzed these data using a non-supervised Euclid distance hierarchical cluster method.52 The aim of such cluster analysis is to measure similarities between different data points, which is otherwise difficult to achieve visually for a relatively small data set of 157 genes; we therefore visualized the four gene groups in n-dimensional space where n is the number of tissue classes, in which context Euclidean distance is simple to compute. The Clustergrams so derived represent heatmaps with dendrograms for hierarchical clustering of matrix data with column dendrograms; the rows represent genes and the columns represent samples. Default clusters are created by average linkage with Euclidean distance metric, whereas hierarchical cluster trees are created using a single-linkage algorithm in which the input Y is a distance matrix. Similar results were generated using other methods. In addition, to quantify the transcriptomic similarity of GKs and POs, and their distinction from CTs and HDs, we conducted a multinomial logistic regression analysis. As our interest focuses on the four modules (CTs, GKs, POs, HDs), we used the median value of the given categories for analysis. Fisher's exact test was used to compare the modules.
To address the problem of computing the P-value for a cluster, we also used an R package, ‘pvclust’ (www.is.titech.ac.jp/%7Eshimo/prog/pvclust/), which is suitable for computing unbiased P-values. We downloaded the relevant gene reference expression data, combined from both human and mouse genomes, using the University of California Santa Cruz gene sorter (GNF Atlas 2—GNF Expression Atlas 2 Data from U133A and GNF1H Chips), analyzed the median value for each group of genes, then conducted hierarchical cluster analysis with multiscale bootstrap (number of bootstrap=10 000 simulations) using an average method and correlation-based dissimilarity matrix.
Results
Gene subgroup identification
Three classes of ‘cancer genes’—CTs, GKs and POs—were defined by hereditary cancer syndromes (CTs and GKs) and/or homologous viral oncogenesis (POs). The genes so identified are listed in Supplementary Table S2. An additional control group of 41 functionally unrelated cardiac development genes implicated in congenital HDs, was also defined (see Materials and methods).
Functional gene classification and mutation frequency
We first determined the biological processes associated with CTs, GKs, POs and HDs using Panther, and established that the functions of POs and GKs are overlapping, but that this is not true for CTs or HDs: the top four processes implicated for GKs and POs (such as, oncogenesis, cell cycle control, cell proliferation and differentiation and protein phosphorylation) are common to both groups, whereas the top five processes for CTs (DNA repair, DNA metabolism, nucleoside metabolism, meiosis and DNA recombination) are exclusive to that group (Supplementary Table S3). A similar molecular function comparison confirms that the top five functional strings (related to kinases, receptors and transcription factors) associated with GKs are shared by POs, whereas CTs and HDs share only one function with GKs and POs (Supplementary Table S4).
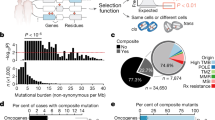
A further comparison of ‘cancer gene’ mutation rates in the germline (that is, predisposing to familial cancers) and in somatic (sporadic) tumors was then performed on CTs, GKs and POs, showing that there are 34 genes with both germline and somatic mutations, 38 genes with germline mutations only and 310 genes with somatic mutations only; germline-only mutations often involve DNA repair (‘mutation modifier’) genes, somatic-only mutations more typically affect POs, whereas GKs may be affected in either context (Supplementary Figure S1). These data confirm a clear soma-germline difference in gene mutation profile: tumor-permissive repair gene (including CT) dysfunction is selectively tolerated in the germline, whereas other tumorigenic mutations (that is, including GK and PO) are more common in somatic tissues (χ2-test, P<2.2e−16). The conclusion that germline GK and PO losses are more embryonic lethal than CTs again supports a shared pivotal role for these genes in enhancing survival.
Gene length and GC content
Like the height and weight of animals, the length and GC content of coding sequences are fundamental features of genes, with implications for transcription and methylation frequency53, 54, 55 that in turn reflect interactions between stochastic genetic determinants and fluctuating environments.56, 57 Figure 1a shows that the coding sequence lengths of GKs and POs are similar to each other but significantly shorter than that of CTs, whereas GC and GC3 contents (GC content within the third site of codons) are significantly higher in GKs and POs than in CTs (Figures 1b and c) as revealed by a multiple logistic model (P=0.0301, P=0.0003 and P=0.0001, for length, GC and CG3s, respectively). Gene length may thus be decreasing, whereas GC and GC3 are increasing from CTs to GKs/POs (Kendall trend analysis, τ=−0.14, two-sided P=0.0540 for length, τ=0.287; two-sided P=7.605 × 10−5 for GC, τ=0.327, two-sided P=6.681 × 10−6). Simple length vs GC3 plot illustrates the structural similarity of GKs and POs, and their distinction from CTs (Figure 1d). We also conducted post hoc non-parametric Nemenyi–Damico–Wolfe–Dunn testing (Supplementary Table S5), including pairwise comparisons (Supplementary Table S6), confirming that GKs resemble POs in these key aspects.
Dinucleotide content and tissue-specific gene expression patterns
More specific differences in methylation-dependent dinucleotide content between GKs, POs and CTs are summarized in Table 1. Frame-dependent dinucleotide alignment and evolutionary analysis have shown that CpG sites in GKs and POs fix missense mutations more often than do those in CTs, and that similar trends apply to whole-genome comparison of apoptosis vs repair genes.6 As the minimum sequence motif is a dinucleotide, we hypothesized that nonrandom alterations in dinucleotide patterns—including asymmetries due to transcription-coupled repair—may be selected from transgenerational variations, the frequency of which varies with germline gene methylation.58 With this in mind, directional analyses of methylation-dependent dinucleotides in the target gene sets reveal an asymmetry: not only are GKs and POs characterized by a higher CpG content than CTs but also by a selectively lower CpA—but not TpG—content, suggesting greater transcription-coupled repair of methylation-dependent mutation in GKs and POs than in CTs.53 This functional conclusion is supported by cross-species gene expression data indicating higher and more tissue-matched transcription frequencies of GKs and POs compared with CTs, with no similarity of either group to control HDs (P<0.03; Figure 2). We also used the R-package ‘pvclust’ to compute the corresponding P-values, based on both the approximately unbiased and bootstrap probability (BP), as shown in Supplementary Figures S5A and S5B. We submit that the topology depth difference in this expression data set supports our conclusion.
Gene expression profiles of caretakers (CTs), gatekeepers (GKs), proto-oncogenes (POs) and heart diseases (HDs) in human and mouse. The original data were downloaded from the University of California Santa Cruz genome site (http://genome.ucsc.edu). A hierarchical cluster analysis of gene data from gnfHumanAtlas2 and gnfMouseAtlas2 (Su AI et al.52) was used for classification with MATLAB clustergram function, computing median values for respective tissues. We compared the patterns seen in various tissues in the four categories using non-parametric methods. Heatmap graph data were scaled from −1 to +1 for Euclid distance. This analysis confirms that gene expression patterns are significantly different between CTs and HDs and GK/POs, both in mouse and in human (P<0.025).
Untranslated region, promoter region and flanking region analyses
Functions of the 5′-and 3′-UTRs include transcriptional regulation, mRNA stability and translational efficiency.59 These UTRs remain under evolutionary selection pressure,60 in turn confirming that conserved sequence homologies reflect associated selectable phenotypes.61 Consistent with this, UTR sequences may modify TSG function through either antisense62 or translational inhibition mechanisms,63 and may directly regulate cell growth and death.64, 65 For these reasons, we compared UTR length and dinucleotide composition in the three defined classes of cancer genes. As shown in Supplementary Figure S2, both the 5′-and 3′-UTRs of CTs are lower in GC content than are those of GKs and POs (multiple Behrens–Fisher Test, CTs vs GKs, P<0.05, CTs vs POs, P<0.001, GKs vs POs, P>0.35). Similarly, both the 5′-and 3′-UTRs of CTs are shorter in length than those of GKs and POs (lower panels, multiple Behrens–Fisher Test, CTs vs GKs, P<0.025, CTs vs POs, P<0.001, GKs vs POs, P>0.30); this finding again suggests an underlying functional similarity between POs and GKs, given that genes functioning in growth and apoptosis tend to be characterized by longer structured 5′-UTRs.65 To compare the methylation-dependent dinucleotide composition in 5′-UTR and 3′-UTR, we analyzed the distribution of relevant dinucleotides. As shown in Supplementary Figures S3A and S3B, the distributions of CpG, TpG and CpA reveal significant differences (with P<0.001 cutoff) between CTs and GKs/POs with respect to 5′-UTR CpG and TpG, but not to CpA or 3′-UTR, suggesting that these differences may arise, in part, due to variations in transcription-coupled DNA repair. With respect to flanking sequences, all parameters are significant between CTs and GKs/POs for 5′ 1 kb flanking sequences, but for 3′ 1 kb flanking sequence only TpG and CpA are significant, consistent with 3′ attenuation of transcription shown in our earlier report.
Non-negative matrix factorization
To assess this structural similarity between GKs and POs, we next proceeded beyond multiple pairwise comparisons. The term (NNMF) refers to a method in which algorithms in multivariate analysis are factorized into matrices by incorporating different constraints, for example, using principal component analysis, in which all elements must be ⩾0.66, 67, 68 We therefore conducted analysis of the coding sequence pattern with an NNMF method that is ideal for non-zero variables, in addition to a linear normalization and transform demonstrated by principal component analysis. We used NNMF to ask the question: given the dinucleotide composition of the 116 putative cancer genes, how many meta-genes and meta-dinucleotides have an informational content similar to the data matrix therein? As all the data were non-negative, we tested whether the three modules (CTs, GKs, POs) were differentially composed of methylation-sensitive (for example, CG) and non-sensitive (for example, AA, TT) dinucleotides for all of the eight-dimensional data including 5′-UTR, 3′-UTR, 5′ 1 kb flanking, 3′ 1 kb flanking, total dinucleotide, frame-specific and dinucleotides. Figure 3 shows the close similarity of GK and PO meta-genes, but also shows the clear difference of CTs.
Non-negative matrix factorization analysis of dinucleotide pattern analysis, performed in MATLAB 7.6 platform. Green, yellow and red color dots represent caretakers (CTs), gatekeepers (GKs) and proto-oncogenes (POs), respectively. The data were computed with Statistical toolbox 5.1 using default non-negative matrix factorization (NNMF) parameters. The x and y axes correspond to the first and second columns of matrix W (W1 and W2), which were in turn computed by the NNMF algorithm, O=W × M, where O is the object n by m matrix, W is n by x and M is x by m. Results were derived using MATLAB statistical toolbox NNMF functions.
Molecular evolutionary rate analysis
Statistical analysis of gene evolutionary rates for the three modules confirms that CTs are evolving significantly faster than either GKs or POs, both of which are under strong purifying selection (non-parametric test, P<0.0001 for dN and dN/dS, but not dS; Figure 4). This conclusion is reinforced across phylogeny (Supplementary Table 7, including HD data, and Supplementary Figure 4).
Molecular evolutionary analysis of caretakers (CTs), gatekeepers (GKs) and proto-oncogenes (POs). Data were downloaded from ENSEMBL (version 50 release) and measured using maximum likelihood method (PAML). Mean and inter-quartile ranges are shown. Statistics were derived using non-parametric tests (P<0.0001 for dN and dN/dS).90
Discussion
Understanding the context-dependent function of so-called ‘cancer genes’ is essential for progress in both normal cell biology and rational anticancer therapeutics. The interacting functional effects of gene networks, or epistasis, is an extension of the concept of allelic dominance, and can be quantitatively modeled using a mathematical approach.25 As ‘cancer genes’ (dominant or recessive) do not cause cancer most of the time, and many of these genes remain wild type in individual tumors, the possibility that normal growth regulatory genes of this class could be subverted to drive tumor microevolution is important to address. Another way of considering this issue involves defining the temporal sequence of genetic lesions that drive oncogenesis, and thus to infer the sequential downstream interactive effect(s) of each lesion on other as-yet-unmutated regulatory genes.69 Indeed, increasing evidence supports the importance of such epistatic processes for mammalian cancer development24, 70, 71, 72, 73 and, perhaps, also for transgenerational carcinogenesis.6, 74
The central finding of the present study is the unanticipated insight that the structural and functional attributes of GKs and POs—when compared with control subgroups of CT cancer suppressor genes and functionally unrelated but developmentally important HD genes—are strikingly similar, as indicated by congruences of evolutionary rate, expression level and breadth, gene length and methylation-dependent mutation confirmed by logistic regression and C-statistics. These data imply that GKs have undergone epigenetic evolution trajectories similar to those of POs, suggesting an explanation for the otherwise puzzling observation that wild-type GK (but not CT) expression is often selectively increased in tumors. Indeed, abundant evidence now confirms that gene behavior varies with the environment; for example, environmental DNA damage—for example, due to smoking75 or inflammation76—selects for TSG methylation in exposed target tissues, thus permitting the upregulation of heterologous wild-type prosurvival gene functions, and contributing to tumor evolution in a negatively selected manner. Consistent with this, recent work shows that the physiological ‘decision threshold’ of MYC to induce either proliferation or apoptosis77 can depend either on gene expression level23 and/or on regulatory interactions with modifier proteins.78 Conversely, clonal overexpression of POs such as HER2 in human tumors seems likely to depend upon previous inhibition of GK expression.17, 79 This epistatic phenomenon has been well described by Vogelstein and Kinzler80 who compared POs and GKs with ‘…electronic components whose effects depend on their placement within a circuit’, a view that is supported by our findings.
Two impressions emerge from the present study: first, a cautionary emphasis on the misleading nature of classifications that ascribe a single invariant function to a given gene; and second, a model that puts epistasis at the heart of carcinogenesis. In this model, upstream CT gene dysfunction (whether germline or somatic) may short circuit the afferent limb of the DNA damage response, biasing GK function towards cell survival and away from apoptosis,81 in much the same way as somatic selection for senescent GK hypofunction permits oncogenic upregulation of wild-type POs.17 Although it is well known that DNA repair defects can potentiate cellular chemosensitivity to (unrepaired) oxidative damage,82 it is less appreciated that repair deficiencies may also impair apoptosis83, 84, 85, 86—presumably through failure to sense damage—thereby, exaggerating tolerance of unrepaired damage with consequent acceleration of carcinogenesis or tumor progression due to increased genetic instability.87, 88, 89
Similar to most studies using systems biology, our conclusions are limited by their inferential and indirect nature, though we note that clinical observations of GK overexpression in human tumors are consistent with the model proposed.12 It is not our contention that GKs are the same as oncogenes, as important differences can also be identified using our approach; for example, we have documented a significantly higher frequency of gene duplicates for POs than for GKs (data not shown). Nevertheless, on the basis of the present study, we submit that GKs are dual function genes, which share with POs a key prosurvival action that is normally subjected to purifying selection, but which may promote tumorigenesis in the abnormal epistatic context of neoplastic cells. Cancer microevolution may thus be accelerated by an age- and damage-driven cascade in which upstream TSG defects select for downstream wild-type GK overexpression. Paradoxically, in this transformed epistatic context, we caution that further upregulation of GK expression by cytotoxic DNA damage could promote tumor growth (that is, efficiently select for resistance) and thus inadvertently reduce patient survival. We therefore speculate that patients whose cancers are associated with GK upregulation may have the best survival outcome if managed with minimal cytotoxic intervention. Further studies—including, ideally, direct testing of the hypothesis using in vitro experimental systems in which wild-type GK expression is inducibly upregulated in reporter cell lines distinguished by different backgrounds of TSG dysfunction will be needed to test our conclusion that overexpressed GKs represent both a biomarker of heterologous suppressor gene defects, and a valid therapeutic target in such contexts.
References
Payne, S. R. & Kemp, C. J. Tumor suppressor genetics. Carcinogenesis. 26, 2031–2045 (2005).
Levitt, N. C. & Hickson, I. D. Caretaker tumour suppressor genes that defend genome integrity. Trends. Mol. Med 8, 179–186 (2002).
Berx, G. & Van Roy, F. The E-cadherin/catenin complex: an important gatekeeper in breast cancer tumorigenesis and malignant progression. Breast. Cancer. Res. 3, 289–293 (2001).
Clarke, A. R. Studying the consequences of immediate loss of gene function in the intestine: APC. Biochem. Soc. Trans. 33 (Part 4), 665–666 (2005).
Martinez, S. L. & Kolodner, R. D. Functional analysis of human mismatch repair gene mutations identifies weak alleles and polymorphisms capable of polygenic interactions. Proc. Natl Acad. Sci. USA. 107, 5070–5075 (2010).
Zhao, Y. & Epstein, R. J. Programmed genetic instability: a tumor-permissive mechanism for maintaining the evolvability of higher species through methylation-dependent mutation of DNA repair genes in the male germ line. Mol. Biol. Evol. 25, 1737–1749 (2008).
Vanin, K., Scurry, J., Thorne, H., Yuen, K. & Ramsay, R. G. Overexpression of wild-type p53 in lichen sclerosus adjacent to human papillomavirus-negative vulvar cancer. J. Invest. Dermatol. 119, 1027–1033 (2002).
Milde-Langosch, K., Bamberger, A. M., Rieck, G., Kelp, B. & Loning, T. Overexpression of the p16 cell cycle inhibitor in breast cancer is associated with a more malignant phenotype. Breast. Cancer. Res. Treat. 67, 61–70 (2001).
Warneford, S., Townsend, M., Rowe, P. B., Dalla-Pozza, L. & Symonds, G. Overexpression of the retinoblastoma gene in a familial adrenocortical carcinoma. Cell. Growth. Differ. 2, 439–445 (1991).
Torbenson, M., Marinopoulos, S., Dang, D. T., Choti, M., Ashfaq, R., Maitra, A. et al. Smad4 overexpression in hepatocellular carcinoma is strongly associated with transforming growth factor beta II receptor immunolabeling. Hum. Pathol. 33, 871–876 (2002).
Tomlinson, J. S., Alpaugh, M. L. & Barsky, S. H. An intact overexpressed E-cadherin/alpha,beta-catenin axis characterizes the lymphovascular emboli of inflammatory breast carcinoma. Cancer. Res. 61, 5231–5241 (2001).
Isola, J., Visakorpi, T., Holli, K. & Kallioniemi, O. P. Association of overexpression of tumor suppressor protein p53 with rapid cell proliferation and poor prognosis in node-negative breast cancer patients. J. Natl. Cancer. Inst. 84, 1109–1114 (1992).
Kim, E., Giese, A. & Deppert, W. Wild-type p53 in cancer cells: when a guardian turns into a blackguard. Biochem. Pharmacol. 77, 11–20 (2009).
Epstein, R. J. Unlocking blockbusters: using Boolean text-mining to optimise clinical trial design and timeline for novel anticancer drugs. Cancer. Informatics. 7, 231–238 (2009).
Watt, F. M., Frye, M. & Benitah, S. A. MYC in mammalian epidermis: how can an oncogene stimulate differentiation? Nat. Rev. Cancer. 8, 234–242 (2008).
Epstein, R. J., Druker, B. J., Roberts, T. M. & Stiles, C. D. Modulation of a Mr 175 000 c-neu receptor isoform in G8/DHFR cells by serum starvation. J. Biol. Chem. 265, 10746–10751 (1990).
Huang, G. C., Hobbs, S., Walton, M. & Epstein, R. J. Dominant negative knockout of p53 abolishes ErbB2-dependent apoptosis and permits growth acceleration in human breast cancer cells. Br. J. Cancer. 86, 1104–1109 (2002).
Podsypanina, K., Politi, K., Beverly, L. J. & Varmus, H. E. Oncogene cooperation in tumor maintenance and tumor recurrence in mouse mammary tumors induced by Myc and mutant Kras. Proc. Natl Acad. Sci. USA. 105, 5242–5247 (2008).
Zhu, G. H., Wong, B. C., Ching, C. K., Lai, K. C. & Lam, S. K. Differential apoptosis by indomethacin in gastric epithelial cells through the constitutive expression of wild-type p53 and/or up-regulation of c-myc. Biochem. Pharmacol. 58, 193–200 (1999).
Saito, Y. & Ogawa, K. Wild type p53 and c-myc co-operation in generating apoptosis of a rat hepatocellular carcinoma cell line (FAA-HTC1). Oncogene. 11, 1013–1018 (1995).
Wagner, A. J., Kokontis, J. M. & Hay, N. Myc-mediated apoptosis requires wild-type p53 in a manner independent of cell cycle arrest and the ability of p53 to induce p21waf1/cip1. Genes. Dev. 8, 2817–2830 (1994).
Wang, Y., Ramqvist, T., Szekely, L., Axelson, H., Klein, G. & Wiman, K. G. Reconstitution of wild-type p53 expression triggers apoptosis in a p53-negative v-myc retrovirus-induced T-cell lymphoma line. Cell. Growth. Differ. 4, 467–473 (1993).
Murphy, D. J., Junttila, M. R., Pouyet, L., Karnezis, A., Shchors, K., Bui, D. A. et al. Distinct thresholds govern Myc's biological output in vivo. Cancer. Cell. 14, 447–457 (2008).
Sathyan, K. M., Nalinakumari, K. R. & Kannan, S. H-Ras mutation modulates the expression of major cell cycle regulatory proteins and disease prognosis in oral carcinoma. Mod. Pathol. 20, 1141–1148 (2007).
Cordell, H. J. Epistasis: what it means, what it doesn’t mean, and statistical methods to detect it in humans. Hum. Mol. Genet. 11, 2463–2468 (2002).
Shi, Y. Z., Hui, A. M., Li, X., Takayama, T. & Makuuchi, M. Overexpression of retinoblastoma protein predicts decreased survival and correlates with loss of p16INK4 protein in gallbladder carcinomas. Clin. Cancer. Res. 6, 4096–4100 (2000).
Wang, Y. C., Lin, R. K., Tan, Y. H., Chen, J. T., Chen, C. Y. & Wang, Y. C. Wild-type p53 overexpression and its correlation with MDM2 and p14ARF alterations: an alternative pathway to non-small-cell lung cancer. J. Clin. Oncol. 23, 154–164 (2005).
Ehrhart, J. C., Gosselet, F. P., Culerrier, R. M. & Sarasin, A. UVB-induced mutations in human key gatekeeper genes governing signalling pathways and consequences for skin tumourigenesis. Photochem. Photobiol. Sci. 2, 825–834 (2003).
Soreide, K., Janssen, E. A., Soiland, H., Korner, H. & Baak, J. P. Microsatellite instability in colorectal cancer. Br. J. Surg. 93, 395–406 (2006).
Kim, K. M., Salovaara, R., Mecklin, J. P., Jarvinen, H. J., Aaltonen, L. A. & Shibata, D. PolyA deletions in hereditary nonpolyposis colorectal cancer: mutations before a gatekeeper. Am. J. Pathol. 160, 1503–1506 (2002).
Helt, C. E., Wang, W., Keng, P. C. & Bambara, R. A. Evidence that DNA damage detection machinery participates in DNA repair. Cell. Cycle. 4, 529–532 (2005).
Li, L. & Zou, L. Sensing, signaling, and responding to DNA damage: organization of the checkpoint pathways in mammalian cells. J. Cell. Biochem. 94, 298–306 (2005).
Wee, K. B. & Aguda, B. D. Akt versus p53 in a network of oncogenes and tumor suppressor genes regulating cell survival and death. Biophys. J. 91, 857–865 (2006).
Chuang, H. Y., Hofree, M. & Ideker, T. A decade of systems biology. Annu. Rev. Cell. Dev. Biol. 26, 721–744 (2010).
Futreal, P. A., Coin, L., Marshall, M., Down, T., Hubbard, T., Wooster, R. et al. A census of human cancer genes. Nat. Rev. Cancer. 4, 177–183 (2004).
Vogelstein, B & Kinzler, K. W The Genetic Basis Of Human Cancer 2nd ed. (McGraw-Hill, New York, 2002).
Wood, R. D., Mitchell, M., Sgouros, J. & Lindahl, T. Human DNA repair genes. Science. 291, 1284–1289 (2001).
Wood, R. D., Mitchell, M. & Lindahl, T. Human DNA repair genes, 2005. Mutat. Res. 577, 275–283 (2005).
Sijmons, R. H. & Burger, G. T. Familial cancer database: a clinical aide-memoire. Fam. Cancer. 1, 51–55 (2001).
Doctor, K. S., Reed, J. C., Godzik, A. & Bourne, P. E. The apoptosis database. Cell. Death. Differ. 10, 621–633 (2003).
Familial cancer database [database on the Internet]. UICC Available from: http://facd.uicc.org/ (2006).
Ashburner, M., Ball, C. A., Blake, J. A., Botstein, D., Butler, H., Cherry, J. M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Thomas, P. D., Campbell, M. J., Kejariwal, A., Mi, H., Karlak, B., Daverman, R. et al. PANTHER: a library of protein families and subfamilies indexed by function. Genome Res. 13, 2129–2141 (2003).
Jabbari, K. & Bernardi, G. Cytosine methylation and CpG, TpG (CpA) and TpA frequencies. Gene. 333, 143–149 (2004).
Smith, N. G. & Hurst, L. D. Molecular evolution of an imprinted gene: repeatability of patterns of evolution within the mammalian insulin-like growth factor type II receptor. Genetics. 150, 823–833 (1998).
Epstein, R. J., Lin, K. & Tan, T. W. A functional significance for codon third bases. Gene. 245, 291–298 (2000).
Lin, K., Tan, S. B., Kolatkar, P. R. & Epstein, R. J. Nonrandom intragenic variations in patterns of codon bias implicate a sequential interplay between transitional genetic drift and functional amino acid selection. J. Mol. Evol. 57, 538–545 (2003).
Cooper, G. M., Brudno, M., Stone, E. A., Dubchak, I., Batzoglou, S. & Sidow, A. Characterization of evolutionary rates and constraints in three Mammalian genomes. Genome. Res. 14, 539–548 (2004).
Green, P., Ewing, B., Miller, W., Thomas, P. J. & Green, E. D. Transcription-associated mutational asymmetry in mammalian evolution. Nat. Genet. 33, 514–517 (2003).
Majewski, J. Dependence of mutational asymmetry on gene-expression levels in the human genome. Am. J. Hum. Genet. 73, 688–692 (2003).
Drummond, D. A. & Wilke, C. O. Mistranslation-induced protein misfolding as a dominant constraint on coding-sequence evolution. Cell. 134, 341–352 (2008).
Su, A. I., Wiltshire, T., Batalov, S., Lapp, H., Ching, K. A., Block, D. et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl Acad. Sci. USA. 101, 6062–6067 (2004).
Tang, C. S, Zhao, Y. Z, Smith, D. K & Epstein, R. J Intron length and accelerated 3′ gene evolution. Genomics. 88 (6), 682–689 (2006).
Chiaromonte, F., Miller, W. & Bouhassira, E. E. Gene length and proximity to neighbors affect genome-wide expression levels. Genome. Res. 13, 2602–2608 (2003).
Liao, B. Y., Scott, N. M. & Zhang, J. Impacts of gene essentiality, expression pattern, and gene compactness on the evolutionary rate of mammalian proteins. Mol. Biol. Evol. 23, 2072–2080 (2006).
Lercher, M. J., Urrutia, A. O., Pavlicek, A. & Hurst, L. D. A unification of mosaic structures in the human genome. Hum. Mol. Genet. 12, 2411–2415 (2003).
Acar, M., Mettetal, J. T. & van Oudenaarden, A. Stochastic switching as a survival strategy in fluctuating environments. Nat. Genet. 40, 471–475 (2008).
Mathieu, O., Reinders, J., Caikovski, M., Smathajitt, C. & Paszkowski, J. Transgenerational stability of the Arabidopsis epigenome is coordinated by CG methylation. Cell. 130, 851–862 (2007).
Tanguay, R. L. & Gallie, D. R. Translational efficiency is regulated by the length of the 3′ untranslated region. Mol. Cell. Biol. 16, 146–156 (1996).
Ponte, I., Monsalves, C., Cabanas, M., Martinez, P. & Suau, P. Sequence simplicity and evolution of the 3′ untranslated region of the histone H1o gene. J. Mol. Evol. 43, 125–134 (1996).
Geraldes, A. & Ferrand, N. A 7-bp insertion in the 3′ untranslated region suggests the duplication and concerted evolution of the rabbit SRY gene. Genet. Sel. Evol. 38, 313–320 (2006).
Kobayashi, M., Yamauchi, Y., Yamaguchi, K. & Tanaka, A. Transient expression assay for antisense RNAs using episomal replication of plasmids: effective reduction of retinoblastoma gene (Rb-1) product by its antisense RNA complementary to 3′-untranslated region. Antisense. Res. Dev. 5, 141–148 (1995).
Qu, X., Qi, Y. & Qi, B. Generation of multiple mRNA transcripts from the novel human apoptosis-inducing gene hap by alternative polyadenylation utilization and the translational activation function of 3′ untranslated region. Arch. Biochem. Biophys. 400, 233–244 (2002).
Schiavone, N., Rosini, P., Quattrone, A., Donnini, M., Lapucci, A., Citti, L. et al. A conserved AU-rich element in the 3′ untranslated region of bcl-2 mRNA is endowed with a destabilizing function that is involved in bcl-2 down-regulation during apoptosis. FASEB. J. 14, 174–184 (2000).
Morrish, B. C. & Rumsby, M. G. The 5′ untranslated region of protein kinase Cdelta directs translation by an internal ribosome entry segment that is most active in densely growing cells and during apoptosis. Mol. Cell. Biol. 22, 6089–6099 (2002).
Lee, D. D. & Seung, H. S. Learning the parts of objects by non-negative matrix factorization. Nature. 401, 788–791 (1999).
Brunet, J. P., Tamayo, P., Golub, T. R. & Mesirov, J. P. Metagenes and molecular pattern discovery using matrix factorization. Proc. Natl Acad. Sci. USA. 101, 4164–4169 (2004).
Devarajan, K. Nonnegative matrix factorization: an analytical and interpretive tool in computational biology. PLoS. Comput. Biol. 4, e1000029 (2008).
Steen, H. B. The origin of oncogenic mutations: where is the primary damage? Carcinogenesis. 21, 1773–1776 (2000).
Marino, S., Romelfanger, C., Yokota, Y. & Nusse, R. Wnt1 is epistatic to Id2 in inducing mammary hyperplasia, ductal side-branching, and tumors in the mouse. BMC. Cancer. 4, 91 (2004).
Martin, R. W., Orelli, B. J., Yamazoe, M., Minn, A. J., Takeda, S. & Bishop, D. K. RAD51 up-regulation bypasses BRCA1 function and is a common feature of BRCA1-deficient breast tumors. Cancer. Res. 67, 9658–9665 (2007).
Bernichtein, S., Petretto, E., Jamieson, S., Goel, A., Aitman, T. J., Mangion, J. M. et al. Adrenal gland tumorigenesis after gonadectomy in mice is a complex genetic trait driven by epistatic loci. Endocrinology. 149, 651–661 (2008).
Cybulski, C., Gliniewicz, B., Sikorski, A., Kladny, J., Huzarski, T., Gronwald, J. et al. Epistatic relationship between the cancer susceptibility genes CHEK2 and p27. Cancer. Epidemiol. Biomarkers. Prev. 16, 572–576 (2007).
Lam, M. Y., Heaney, J. D., Youngren, K. K., Kawasoe, J. H. & Nadeau, J. H. Trans-generational epistasis between Dnd1Ter and other modifier genes controls susceptibility to testicular germ cell tumors. Hum. Mol. Genet. 16, 2233–2240 (2007).
Liu, Y., Lan, Q., Shen, M., Jin, J., Mumford, J., Ren, D. et al. Aberrant gene promoter methylation in sputum from individuals exposed to smoky coal emissions. Anticancer. Res. 28, 2061–2066 (2008).
Wang, F. Y., Arisawa, T., Tahara, T., Takahama, K., Watanabe, M., Hirata, I. et al. Aberrant DNA methylation in ulcerative colitis without neoplasia. Hepatogastroenterology. 55, 62–65 (2008).
Freie, B. W. & Eisenman, R. N. Ratcheting Myc. Cancer. Cell. 14, 425–426 (2008).
Gordan, J. D., Lal, P., Dondeti, V. R., Letrero, R., Parekh, K. N., Oquendo, C. E. et al. HIF-alpha effects on c-Myc distinguish two subtypes of sporadic VHL-deficient clear cell renal carcinoma. Cancer. Cell. 14, 435–446 (2008).
Feng, Q., Deftereos, G., Hawes, S. E., Stern, J. E., Willner, J. B., Swisher, E. M. et al. DNA hypermethylation, Her-2/neu overexpression and p53 mutations in ovarian carcinoma. Gynecol. Oncol. 111, 320–329 (2008).
Vogelstein, B. & Kinzler, K. W. Cancer genes and the pathways they control. Nat. Med. 10, 789–799 (2004).
Nowak, M. A., Michor, F., Komarova, N. L. & Iwasa, Y. Evolutionary dynamics of tumor suppressor gene inactivation. Proc. Natl Acad. Sci. USA. 101, 10635–10638 (2004).
de Boer, J., Andressoo, J. O., de Wit, J., Huijmans, J., Beems, R. B., van Steeg, H. et al. Premature aging in mice deficient in DNA repair and transcription. Science. 296, 1276–1279 (2002).
Robles, A. I., Wang, X. W. & Harris, C. C. Drug-induced apoptosis is delayed and reduced in XPD lymphoblastoid cell lines: possible role of TFIIH in p53-mediated apoptotic cell death. Oncogene. 18, 4681–4688 (1999).
Hinz, J. M., Helleday, T. & Meuth, M. Reduced apoptotic response to camptothecin in CHO cells deficient in XRCC3. Carcinogenesis. 24, 249–253 (2003).
Massey, A., Offman, J., Macpherson, P. & Karran, P. DNA mismatch repair and acquired cisplatin resistance in E. coli and human ovarian carcinoma cells. DNA. Repair. (Amst). 2, 73–89 (2003).
Gueven, N., Becherel, O. J., Howe, O., Chen, P., Haince, J. F., Ouellet, M. E. et al. A novel form of ataxia oculomotor apraxia characterized by oxidative stress and apoptosis resistance. Cell. Death. Differ. 14, 1149–1161 (2007).
Cao, L., Kim, S., Xiao, C., Wang, R. H., Coumoul, X., Wang, X. et al. ATM-Chk2-p53 activation prevents tumorigenesis at an expense of organ homeostasis upon Brca1 deficiency. EMBO. J. 25, 2167–2177 (2006).
Porcedda, P., Turinetto, V., Lantelme, E., Fontanella, E., Chrzanowska, K., Ragona, R. et al. Impaired elimination of DNA double-strand break-containing lymphocytes in ataxia telangiectasia and Nijmegen breakage syndrome. DNA. Repair. (Amst). 5, 904–913 (2006).
Kaminski, B. C., Grabenbauer, G. G., Sprung, C. N., Sauer, R. & Distel, L. V. Inter-relation of apoptosis and DNA double-strand breaks in patients with multiple primary cancers. Eur. J. Cancer. Prev. 15, 274–282 (2006).
Yang, Z . PAML: a program package for phylogenetic analysis by maximum likelihood. CABIOS 13, 555–556 (1997).
Acknowledgements
Author contributions: YZ performed the experimental analysis, assisted in the interpretation and statistics, and wrote the paper. RJE designed the experiments, wrote the paper, and finalized the paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Zhao, Y., Epstein, R. Unexpected functional similarities between gatekeeper tumour suppressor genes and proto-oncogenes revealed by systems biology. J Hum Genet 56, 369–376 (2011). https://doi.org/10.1038/jhg.2011.21
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2011.21