Abstract
To understand soil microbial community stability and temporal turnover in response to climate change, a long-term soil transplant experiment was conducted in three agricultural experiment stations over large transects from a warm temperate zone (Fengqiu station in central China) to a subtropical zone (Yingtan station in southern China) and a cold temperate zone (Hailun station in northern China). Annual soil samples were collected from these three stations from 2005 to 2011, and microbial communities were analyzed by sequencing microbial 16S ribosomal RNA gene amplicons using Illumina MiSeq technology. Our results revealed a distinctly differential pattern of microbial communities in both northward and southward transplantations, along with an increase in microbial richness with climate cooling and a corresponding decrease with climate warming. The microbial succession rate was estimated by the slope (w value) of linear regression of a log-transformed microbial community similarity with time (time–decay relationship). Compared with the low turnover rate of microbial communities in situ (w=0.046, P<0.001), the succession rate at the community level was significantly higher in the northward transplant (w=0.058, P<0.001) and highest in the southward transplant (w=0.094, P<0.001). Climate warming lead to a faster succession rate of microbial communities as well as lower species richness and compositional changes compared with in situ and climate cooling, which may be related to the high metabolic rates and intense competition under higher temperature. This study provides new insights into the impacts of climate change on the fundamental temporal scaling of soil microbial communities and microbial phylogenetic biodiversity.
Similar content being viewed by others
Introduction
One of the great scientific challenges in 21st century ecology is to understand the response, succession and stability of biological communities to potential threats from anthropogenic disturbances, such as climate change (Thomas et al., 2004; Bellard et al., 2012; Fussmann et al., 2014). Given the fundamental role of microbial communities in biogeochemical cycling, their responses to climate change may lead to ecosystem structure repercussions and feedback to the climate system (Wardle et al., 2004; Bardgett et al., 2008; Castro et al., 2010; Singh et al., 2010; Gutknecht et al., 2012). Previous studies indicated that temperature is an important determinant of soil microbial composition (Castro et al., 2010; Vanhala et al., 2011; Zhou et al., 2012), diversity (Pold and DeAngelis, 2013) and ecological functions, such as soil respiration (Schindlbacher et al., 2011; Zhao et al., 2014), organic matter decomposition (Conant et al., 2011) and nitrification (Long et al., 2012; Zhao et al., 2014). However, information is still lacking on the relationship between microbial community succession and multiple environmental changes resulting from climate change.
The transplantation of soils between different geoclimatic regions provides a powerful approach to test the responses of microbial communities to climate change scenarios that simultaneously alter multiple factors (Bottomley et al. 2006; Waldrop and Firestone, 2006; St John et al., 2011; Vanhala et al., 2011; Zumsteg et al., 2013; Sun et al., 2014; Zhao et al., 2014). In general, an increase in temperature and precipitation is expected in southward transplantation (climate warming), whereas a decrease in these parameters is expected in northward transplantation (climate cooling) at northern latitudes along the coastal areas. The soils transplanted to the north/south acquire colder/warmer climate features. Thus, we were able to analyze the climatic effects on the sensitivity of microbial communities. The climatic effect on microbial communities simulated by soil transplantation has been reported, but is still under debate. For example, soils transplanted from north to south resulted in a loss of microbial biomass and changes in the microbial community structure and functions in response to the warmer climate, which indicates that exposure to a temperature increase of 4.5 °C for 2 years was sufficient to change the soil biology (Vanhala et al., 2011). Waldrop and Firestone (2006) found that microbial biomass, composition, enzyme activities and respiration decreased when soils under the oak canopies were transplanted to an open grassland environment with higher maximum temperature and lower soil water content. Bottomley et al. (2006) also reported that the fungal and bacterial community structures of forest soil changed significantly after transfer from forests to meadows after 2 years of incubation. Similarly, our previous studies indicated that transplanting either black soil to warmer regions (Zhao et al., 2014) or red soil to colder regions (Liu et al., 2014) caused significant changes in the microbial functional gene diversity and patterns. By contrast, Lazzaro et al. (2011) reciprocally transplanted soil samples from two different unvegetated glacier forefields (calcareous and siliceous) and found that bacterial communities were more significantly affected by seasons than by the transplantation after 15 months of incubation. Another study indicated that the bacterial community composition in grassland soil (as assessed by the phospholipid-derived fatty acid guilds) appeared to be resistant when soils were transplanted to a conifer ecosystem (Balser and Firestone, 2005). The contrasting results require further studies to examine the effects of transplantation on the microbial community structure and the mechanisms involved.
Species temporal turnover is defined as the number of species eliminated and replaced per unit time (MacArthur and Wilson, 1967; Magurran, 2004; Hatosy et al., 2013). Changes in species number can be described using the time–decay relationship, which is a common model to describe the changes in community similarity over time (Nekola and White, 1999). Rates of temporal turnover vary in relation to ecosystem types (Korhonen et al., 2010), local environmental factors (Werner et al., 2007), disturbances (Svensson et al., 2009) and temporal scales (Hatosy et al., 2013). Temporal turnover has been well documented in plant and animal communities, but information on microbial temporal turnover is insufficient. Only a few studies, which were primarily based on engineered systems for relatively short periods (from minutes to months), used the time–decay relationship to test whether theoretical predictions of community assembly and dynamics are applicable to microbial communities (Oliver et al., 2012; Shade et al., 2013). Recently, Hatosy et al. (2013) studied bacterial temporal beta-diversity across different temporal scales in three marine microbial communities and found that turnover at different temporal scales appeared to be driven by different factors. The variation of soil microbial temporal turnover with climate change and the underlying mechanisms remain unknown.
To understand soil microbial community stability and temporal turnover in response to climate warming and cooling, a soil transplantation experiment was conducted in three long-standing agricultural stations located over large transects from the Fengqiu station in central China (warm temperate zone) to the Yingtan station in southern China (middle subtropical zone) and the Hailun station in northern China (cold temperate zone). Soil samples were collected from the three stations annually from 2005 to 2011 to test the following hypotheses: (i) soil transplantations will significantly alter soil microbial temporal turnover and climate warming will lead to higher microbial succession rates; (ii) different microbial groups will show differential sensitivities to temperature change perturbations; (iii) temperature is the most important factor that influences the fundamental temporal scaling of microbial biodiversity. The microbial communities were analyzed by sequencing microbial 16S ribosomal RNA (rRNA) gene amplicons with the Illumina MiSeq technology (San Diego, CA, USA). Our results revealed a distinctly different pattern of microbial communities in the northward and southward transplantations. Climate warming caused higher microbial temporal turnover and less stability, which may be attributed to high metabolic rates and intensive competition under higher temperatures.
Materials and methods
Study sites and sample collections
The long-term soil transplant experiments were conducted in agricultural experimental stations of the Chinese Academy of Sciences at three sites: the Hailun station (site N, 126° 38′ E and 47° 26′ N) in the Heilongjiang Province of northern China, the Fengqiu station (site M, 114° 24′ E and 35° 00′ N) in the Henan Province of central China and the Yingtan station (site S, 116° 55′ E and 28° 15′ N) in the Jiangxi Province of southern China. In October 2005, blocks with a size of 1.4 m in length × 1.2 m in width × 1.0 m in depth were established in each station. These blocks were surrounded by 20-cm brickwalls and underlain with sand to isolate each from the surrounding environment. Soils from the central Fengqiu site, classified as Cambisol soil, were transported northward to the Hailun site and southward to the Yingtan site. Soil was excavated in five 0.2-m deep layers, with each layer sufficiently mixed and then repacked sequentially to maintain the soil stratification. At each site, triplicate soil blocks were established with maize cropping. Maize has been planted since 2006 at all three sites with regular fertilization of 150 kg hm−2 nitrogen, 75 kg hm−2 phosphorus and 60 kg hm−2 potassium in the forms of urea, (NH4)2HPO4 and KCl, respectively. Basal fertilizer was applied before planting (half of nitrogen, all phosphorus and all potassium). The second half of nitrogen fertilizer was applied as the top dressing at large trumpet stage of maize growth. No irrigation was applied. Maize was sowed and harvested once a year. The soil samples with maize cropping from these three sites were designated as in situ (original Fengqiu site), northward to the Hailun site and southward to the Yingtan Site. This experiment belongs to an integrated project (The Soil Reciprocal Transplant Experiment, SRTE), which serves as a platform for a number of studies that evaluate climate and cropping effects on soil microbial diversity and its agro-ecosystem function (Sun et al., 2013; Liu et al., 2014; Zhao et al., 2014). Triplicate soil samples from each station, one sample from each block, were collected annually in August to September from 2006 to 2011. In addition, samples of the original soil transported to each site were collected in 2005. Ten soil cores were composited from surface soil (0–20 cm) within each block and sealed in a polythene wrapper, then stored on ice and transported to the laboratory. Any visible living plant material (for example, roots) was manually removed from the composited soil in the lab. The soil was then divided into two subsamples and stored at either 4 °C for soil geochemical variable measurements or −80 °C for microbial community analysis. Soil geochemical variables were measured as follows: pH was determined with a glass electrode in water-to-soil ratio of 2.5:1 (v/w). Total nitrogen, nitrate (NO3−−N) and ammonium nitrogen (NH4+−N) were measured by the Kjeldahl method (Bremner, 1965). Total phosphorus and available phosphorus were extracted by sodium carbonate and sodium bicarbonate, respectively; both were determined by the molybdenum blue method (Olsen et al., 1954). Total potassium and available potassium were determined by flame photometry after extraction with sodium hydroxide and ammonium acetate, respectively (Kanehiro and Sherman, 1965). Soil organic matter (SOM) was measured by the potassium dichromate oxidation method (Allison, 1965). Soil electrical conductivity was determined with a soil conductivity meter. Cation exchange capacity (CEC) was measured in an ammonium acetate solution at pH 7 (Chapman, 1965). Climate attributes, including the annual average temperature, rainfall and relative humidity were obtained from the meteorological observation database of the experimental stations. The above ground biomass and grain weight of maize were immediately measured after harvest.
Illumina sequencing analysis of 16S rRNA gene amplicons
Microbial genomic DNA was extracted from 5 g of well-mixed soil for each sample by combining freeze-grinding and sodium dodecyl sulfate for cell lysis, and purified by agarose gel electrophoresis, followed by phenol–chloroform–butanol extraction as previously described (Zhou et al., 1996). The primers 515 F (5′-GTGCCAGCMGCCGCGG-3′) and 806 R (5′-GGACTACHVGGGTWTCTAAT-3′) targeting the V4 hypervariable regions of microbial 16S rRNA genes were selected (Caporaso et al., 2012). The forward and reverse primers were tagged with adapter, pad and linker sequences. Each barcode sequence (12 mer) was added to the reverse primer for pooling of multiple samples in one run of MiSep sequencing. All primers were synthesized by Eurofins/MWG (Huntsville, AL, USA).
PCR amplification was performed in triplicate using a Gene Amp PCR-System 9700 (Applied Biosystems, Foster City, CA, USA) in a total volume of 25 μl, which contained 2.5 μl of 10 × PCR buffer II, and 0.5 unit of AccuPrime Taq DNA Polymerase High Fidelity (Invitrogen, Carlsbad, CA, USA), 0.4 μm of each primer and 10 ng of template DNA. Thermal cycling conditions were as follows: an initial denaturation at 94 °C for 1 min, followed by 25 cycles at 94 °C for 20 s, 53 °C for 25 s and 68 °C for 45 s, with a final extension at 68 °C for 10 min.
Following amplification, 2 μl of the PCR product was used for agarose gel (1%) detection. The triplicate PCR reactions for each sample preparation were combined and quantified with PicoGreen. From each sample, 200 ng of the PCR product was collected and pooled with other samples for one sequencing run. The pooled mixture was purified with a QIAquick Gel Extraction Kit (QIAGEN Sciences, Germantown, MD, USA) and re-quantified with PicoGreen.
According to the MiSeq Reagent Kit Preparation Guide (Illumina, San Diego, CA, USA), the purified mixture was diluted and denatured to obtain the 8 pm sample DNA library and mixed with an equal volume of 8 pm PhiX (Illumina). Finally, 600 μl of the mixture library was loaded with read 1, read 2 and the index sequencing primers (Caporaso et al., 2012) on a 300-cycle (2 × 150 paired ends) kit and run on a MiSeq apparatus at the Institute for Environmental Genomics of the University of Oklahoma.
After assigning each sequence to its sample according to its barcode, allowing up to two mismatches, a total of 9 48 765 reads from both ends were obtained as a partitioned run for the total 63 samples. To increase reproducibility (Zhou et al., 2011b), deep sequencing was performed with these samples. These sequences were then trimmed using Btrim with threshold of quality scores higher than 20 over a 5 bp window size and a minimum length of 100 bp (Kong, 2011). Forward and reverse reads with at least a 50 bp overlap and <5% mismatches were joined using FLASH (Magoc and Salzberg, 2011). After removing the sequences with ambiguous bases (that is, N), the sequences with lengths between 245 and 258 bp were subjected to chimera removal by U-Chime (Edgar et al., 2011) using GreenGenes core 16S reference sequences. Operational taxonomic units (OTUs) were clustered using the Uclust program at the 97% similarity level (Edgar, 2010). Final OTUs were generated based on the clustering results, and taxonomic annotations were assigned to each OTU’s representative sequence by the RDP 16S Classifier (Wang et al., 2007). Singletons were removed for downstream analyses. Samples were rarefied at 10, 947 sequences per sample. The above mentioned steps were performed through the Galaxy pipeline at the Institute for Environmental Genomics, University of Oklahoma (http://zhoulab5.rccc.ou.edu/).
Time–decay relationship and other statistical analyses
We used linear regression to examine the relationship between the temporal distance among samples and similarity in microbial composition. The Bray–Curtis distance was used as a taxon-based metric of differences in community composition. Arrhenius (log–log) plot was used for modeling the species–time relationship in the form: ln(Ss)=constant−w ln(T), where Ss is the pairwise similarity in community composition, T is the time interval and w is a measure of the rate of species turnover across time. A one-sample t-test between the original slope and a mean of bootstrapped slopes by random pairing of the original set (permuted 999 times) was performed for testing the significance of w values (Horner-Devine et al., 2004; Zhou et al., 2008). The significance comparison of w values among different estimations was also achieved by bootstrapping (999 times), followed by a pairwise t-test.
The microbial distribution patterns of northward and southward transplants were determined by nonmetric multidimensional scaling (NMDS) (Kruskal, 1964). A dissimilarity test of the microbial community composition was performed using non-parametric multivariate statistical tests and analysis of similarities (Clarke, 1993). The Mantel and partial Mantel tests were used to calculate the correlations between environmental factors and the soil microbial community (Legendre and Legendre, 2012). BIO-ENV is an algorithm identifying the best subset of environmental variables such that the Euclidean distances of scaled environmental variables have the maximum (rank) correlation with community dissimilarities (Clarke and Ainsworth, 1993). Canonical correspondence analysis (CCA) and partial CCA (Legendre and Legendre, 2012) were also used to identify the effect of soil geochemical variables, plants and climate on the microbial community composition. All the above analyses were performed in R (version 3.0.2; http://www.r-project.org/) using the vegan (Oksanen et al., 2013) and ecodist (Goslee and Urban, 2007) packages. A circular maximum likelihood phylogenetic tree was constructed based on the sequences representative for each OTU as determined by Uclust. The tree was generated with MEGA 5 (Tamura et al., 2011) using a neighbor-joining method with a bootstrap value of 1000 displayed using iTOL (Letunic and Bork, 2011).
Results
Effect of transplants on plant and soil geochemical variables
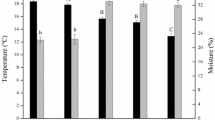
The average annual temperature in the Fengqiu station (in situ) is 14.0 °C, which decreases to 2.2 °C in the Hailun station (northward transplantation) and increases to 18.3 °C in the Yingtan station (southward transplantation) (Supplementary Figure S1a). The average annual precipitation was similar between Fengqiu and Hailun, but twice as high in Yingtan (Supplementary Figure S1b). The aboveground maize biomass fluctuated in each year after soil transplantation (Supplementary Figure S2a), which might be attributed to variation of meteorological characteristics such as temperature, precipitation and so on. However, yield production (seed weight) significantly decreased in both northward and southward transplants as compared with in situ (Supplementary Figure S2b), thereby indicating the significant impact of climate cooling or warming on crop yields, especially for warming-induced lower productivity. A few soil geochemical attributes also fluctuated after transplantation, such as pH, CEC and available potassium in northward transplantation and pH, CEC, SOM, available potassium, NO3− in southward transplantation (Supplementary Table S1).
Overall pattern of microbial succession
Soil microbial communities in the three locations were analyzed annually from 2005 to 2011 by sequencing microbial 16S rRNA gene amplicons with Illumina MiSeq technology. Interestingly, the northward transplant showed increased microbial richness in terms of the number of different OTUs (30, 870 OTUs in total with an average of 5035±313 OTUs) compared with that in situ (30, 109 OTUs in total with an average of 4804±264 OTUs), with the highest increase during the fourth year (15.6%, P<0.001) (Supplementary Figure S3). By contrast, southward transplant showed a decrease in microbial richness (29, 343 OTUs in total with an average of 4671±228 OTUs) (Supplementary Figure S3).
Further comparison of the microbial taxonomic composition at the phylum level showed that Proteobacteria was the dominant phylum present in all three sites, followed by Acidobacteria (Supplementary Figure S4). However, the southward transplant showed a distinct change in microbial taxonomic composition, such that the relative abundances of Proteobacteria and Bacteroidetes had significantly decreased (P<0.05) since 2006, whereas those of Acidobacteria and Actinobacteria had significantly increased (P<0.05) since 2007 (Supplementary Figure S4).
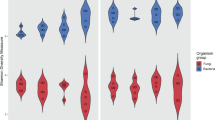
The overall pattern of microbial succession in the long-term soil transplant experiment is visualized on the first two coordinates of the nonmetric multidimensional scaling ordination based on the Bray–Curtis dissimilarity (Figure 1a). As expected, the close clustering of samples in 2005 indicated similar origins for the soils that contained similar microbial composition. A clearly different pattern was observed among samples in the three sites starting from 2006. The samples became distinctly separate from each other with time. This observation was confirmed by analysis of similarities, which showed that the microbial community structures were significantly different in each location from 2006 to 2011 (Supplementary Table S2). We calculated the pairwise distances between microbial communities in each year based on the Bray–Curtis dissimilarity, and the results are shown in Figure 1b. Obviously, the variations continuously increased between in situ soil and the northward transplant (r2=0.58, P<0.05). The correlations were even more significant between in situ soil and the southward transplant (r2=0.81, P<0.01), and between the northward and southward transplants (r2=0.91, P<0.01).
(a) Non-metric multidimensional scaling ordination based on Bray–Curtis distances showing the changes in microbial community composition with northward (square) and southward transplant (triangle) compared with microbial community in original Fengqiu station (circle). Color from deep to shallow represents the community succession from year 2005 to 2011. (b) Small differences among samples (based on Bray–Curtis dissimilarity) were linearly enlarged along as time elapsed. The lines denote the linear regression. Square symbols represent differences between northward and in situ samples; triangle symbols represent differences between southward and in situ samples; circle symbols represent differences northward and southward samples.
Changes in microbial temporal turnover with soil transplant
The slopes of microbial time–decay relationship were estimated by linear regression with log-transformed microbial community similarity (Figure 2). A significant time–decay relationship was observed for microbial communities in situ, with a low temporary turnover for the in situ samples (w=0.046, P<0.001) across 6 years of transplantation. Compared with the in situ soil, the slope was significantly steeper in the northward transplant (w=0.058, P<0.001), and even steeper in the southward transplant (w=0.094, P<0.001). Figure 3a shows the relationship between microbial temporal turnover and temperature. The northward transplant was accompanied by a larger change in temperature, that is, a decrease of 11.9 °C on the average, as compared with an increase of 4.3 °C for the southward transplant. However, climate warming significantly increased the microbial community temporal turnover in the southward transplant with higher fluctuation (w=0.088±0.017), as compared with the northward transplant (w=0.051±0.013) and in situ (w=0.039±0.008). To obtain general insights into the temporal change of microbial communities across different time periods, the temporal turnover of the microbial community was estimated yearly starting from 2005 (Figure 3b). The southward transplant showed a greater effect on microbial community temporal turnover than northward transplant most of the time.
The microbial time–decay relationships in taxonomic divisions were also estimated (Table 1). Considerable variations of w values were observed among different phyla. Both northward and southward transplants showed a significant increase in microbial temporal turnover at all phylum levels. The mean w values were 0.037±0.023, 0.061±0.027 and 0.112±0.055 among different phyla of the in situ soil, northward transplant and southward transplant, respectively. The phyla Actinobacteria and Firmicutes were found to be the most sensitive to either climate cooling or warming. Proteobacteria showed the least response to both northward and southward transplants. Interestingly, significant correlations were found between temporal turnover of all phyla and the taxonomic abundances (P<0.05) (Figure 3c). The turnovers of Proteobacteria, Bacteroidetes and Verrucomicrobia were negatively correlated with phylum abundance (r2 of 0.18–0.40, P<0.05). By contrast, the turnovers of Acidobacteria, Actinobacteria, Firmicutes and Planctomycetes were positively correlated with phylum abundance (r2 of 0.12–0.28, P<0.05).
Relative abundance of bacteria detected in situ and with soil transplant
A quite low number of microbial OTUs (78 OTUs belonging to nine phyla) were detected in all in situ samples and in soil transplants during the 6-year experiment. A phylogenetic tree of these bacteria was constructed using MEGA 5 (Figure 4). The relative abundances of bacteria in situ and with transplanting are indicated by bars. The results indicated that OTUs belonging to Acidobacteria Gp4 and Gp6, Arthrobacter, Fervidicoccus, Nitrospira, Sphingomonas, Sphingosinicella, Steroidobacter and Terrimonas were predominant. Of these OTUs, northward transplant caused an increase in abundance of 27 OTUs and a decrease of 51 OTUs. Southward transplant caused an increase in abundance of 42 OTUs and a decrease of 35 OTUs.
Linkage between microbial succession and environmental factors
CCA was performed to discern the linkage between microbial succession and the environmental factors (Supplementary Figure S5). The analysis predicts the principal coordinates using a linear combination of several environmental gradients. The variation in microbial phylogenetic community structure was explained by the variation in the identified environmental gradients (40.1% with the first two constrained axes). The CCA results indicated that temperature and precipitation were important environmental attributes that control the microbial community structure in the southward transplant. Soil geochemical variables, such as pH, CEC, soil electrical conductivity, total phosphorus, total potassium and plant biomass, were also weighted favorably (long vector arrow in Supplementary Figure S5) in the control of microbial variations.
To determine how environmental factors such as temperature, plant biomass and soil geochemical variables (pH, CEC, soil electrical conductivity, total potassium and total phosphorus), affected microbial composition during the 6-year experiment, Mantel and partial Mantel tests were performed (Table 2). Temperature was found to be the most important factor that caused microbial β-diversity of all soil samples, especially in the first year of soil transplanting (rm=0.708, P<0.01). Interestingly, we found that microbial community diversity in the northward transplant was mainly affected by soil geochemical variables (rm of 0.217–0.637, P<0.05), whereas that in the southward transplant was mostly influenced by temperature (rm of 0.477–0.880, P<0.05).
Discussion
Given the central and global importance of microorganisms in environments, one of the fundamental objectives is to understand how biodiversity is accumulated and maintained across time and space (Hanson et al., 2012; Shade et al., 2013). Microbial community structures and their ecological functions are sensitive in response to global climate changes (Singh et al., 2010; Gutknecht et al., 2012; Luo et al., 2014). However, to date, insufficient information is available on the succession dynamics, stability and responses of microbial communities under the integrated effects of climate change, such as temperature, precipitation and vegetation. In this study, we hypothesized that when soils are exposed to colder/warmer climate, the structure of the soil microbial community significantly changes as the bacteria adapts to new climatic conditions across a time span of several years. Our results supported this hypothesis. The microbial community structure was originally the same, but was altered significantly in 6 years by soil transplantation (Figure 1). The difference between any two locations continuously increased with time, which implied that the group of microbes exposed to dramatic climate change is neither resistant nor resilient during the years of the study. Recovery may fail to occur after persistent disturbances caused by climate change, which was demonstrated in a previous study wherein the transplanted soil microbial community was closer to the microbial community in the local soil than that in the original location twenty years after soil transplantation (Sun et al., 2014). In addition, previous studies indicated that the presence of plants may enhance the resilience of soil microbial communities (de Vries et al., 2012) or even inhibit the effects of soil transplantation (Liu et al., 2014), which might be related to the high resource availability offered by the plant (de Vries and Shade, 2013). However, a clear differential pattern was still observed even with maize cropping in this study. The conflicting results indicated that plants may only dampen disturbance of soil transplantation to a certain extent for a short period. Such hypothesis highlights the need to understand climate-induced changes of interactions between plants and soil communities and the corresponding feedback.
The temporal turnover of the time–decay relationship is an important indicator of the succession dynamics of biological communities. The w value was low across the 6-year experiment, but we still observed a significantly linear decrease in the microbial similarity with time (log transformed) for the in situ samples and the northward and southward transplants. The results strongly support the claim that the community similarity decays over time (time–decay), which underlies key ecological principles; this decay appears to be universal in biology (Chytry et al., 2001; Korhonen et al., 2010; Gonzalez et al., 2012; Shade et al., 2013). We compared the microbial temporal turnover in different habitats and across different temporal scales (Supplementary Figure S6). Generally, the turnover values obtained in this study (0.026–0.114) had the same magnitude as those of other studies (0–0.3) (Oliver et al., 2012; Hatosy et al., 2013; Shade et al., 2013). The turnover values of microbes in these different studies were obtained with a variety of approaches at various temporal scales, from minutes to years. Consequently, exact comparisons at fine resolutions would be impractical. Generally, the temporal turnover of soil microbial communities is low as compared with that of larger organisms (Shade et al., 2013). This phenomenon might be attributed to the unique biology of microorganisms such as massive population sizes, high dispersal rates, rapid asexual reproduction and resistance to extinction. Empirical studies across a wide range of taxa showed that more diverse communities have greater temporal stability of their species composition (Shurin, 2007). Thus, the decline in turnover for microbes occurs when high diversity either facilitates colonization of new species or reduces extinction rates of extant species. Second, the much steeper slope observed in soil transplants than in situ soil indicated that dramatic climate change increased the microbial succession rate. This result confirmed our hypothesis that microbial temporal turnover is significantly altered by soil transplantation. The rapid elimination and replacement of species might indicate the decreased stability and reassembling process of microbial populations with dramatic climate change. This phenomenon could be explained by the species-sorting concept of the metacommunity framework, which assumes that microbial communities have the potential to adapt to new environmental conditions by adjusting their composition (Leibold et al., 2004; Székely et al., 2013). In addition, changes of microbial temporal turnover could also be due to differences in resident bacterial diversity present at the north and south sites. One would predict that higher colonization potential (for example, higher likelihood that more species could successfully establish in the transplanted soils) could therefore be a driving force behind the patterns observed.
Different subsets of the microbial community showed varying slopes of the time–decay relationships in response to climate change, thereby indicating the differences in sensitivity of the microbial taxonomic groups to disturbance. Some taxa, such as the phyla Actinobacteria and Firmicutes are more sensitive to climate change disturbances than others. The differential responses may affect the overarching resistance and resilience of the community by changing ecological interactions among species (de Vries and Shade, 2013). A critical challenge is to construct a molecular ecological network of microbial communities in response to changing environmental conditions (precipitation, temperature and nutrient input) (Zhou et al., 2010; 2011a). Furthermore, given that changes in microbial community composition are often associated with changes in functional capabilities (Fierer et al., 2007; Strickland et al., 2009), further studies linking the temporal patterns of microbial community with their ecological functional processes under climate change are necessary.
We also hypothesized that the impact of southward transplant (climate warming) on microbial community is more significant than that of northward transplant (climate cooling). First, soil transplantation in the opposite directions resulted in different alterations in microbial richness (OTU number), which mostly decreased in the southward transplant and increased in the northward transplant. A previous study also indicated that soils transplanted to warmer regions lost microbial biomass (Vanhala et al., 2011). This might be partially explained by the predator–prey relationships with temperature (Fussmann et al., 2014). Low temperature may decrease the number of predators such as nematodes in soil aggregates, thereby increasing the population of some soil bacterial species and vice versa.
Second, the southward transplant caused more significant changes in the microbial taxonomic composition as compared with the northward transplant, thereby indicating less ecological stability in the composition of microbial communities after persistent climate warming disturbance. The relative abundances of Proteobacteria and Bacteroidetes decreased, whereas those of Acidobacteria and Actinobacteria increased in the southward transplant. These observations are in accordance with previous findings that the most abundant microbial phyla, namely, Actinobacteria, Proteobacteria, Acidobacteria, Planctomycetes and Bacteroidetes, are significantly different between the warming treatment and the controls (Luo et al., 2014). Deslippe et al. (2012) also observed that climate warming led to a significant reduction in the evenness of microbial communities; it was associated with the significant increase in the dominance of Actinobacteria and the significant reduction of Gemmatimonadaceae and Proteobacteria. In addition, acid precipitation often occurs in the southern parts of China (Wang and Wang, 1995), thereby resulting in fluctuations of the soil pH (Krug and Frink, 1983), cation exchange equilibrium (Mcfee et al., 1977) and the chemical composition of soil water (Likens et al., 1996). All these changes may serve as environmental filters that require specific adaption strategies for microbial survival after southward transplantation (Fierer et al., 2007).
Third, we observed that southward transplantation had a more significant effect on microbial temporal turnover than northward transplantation. This observation is in accordance with previous findings, which stated that microbial community structures in soil transferred from a cold to a warm site induced high change rates because of higher microbial activity and faster species turnover than the reverse transfer (Zumsteg et al., 2013). Southward transplantation of soil causes a significant increase in temperature, which was found to be the most important factor in driving microbial temporal patterns. On the basis of the metabolic theories in ecology (Brown et al., 2004), higher temperature accelerates the consumption of substrate (Kirschbaum, 2004; Rousk et al., 2012) and, thus, may increase microbial internal competition because of lower substrate availability. The increased competition may cause higher temporal turnover and lower population numbers and population density with climate warming. However, further well-replicated, time-series experiments are required to elucidate the relative importance of temperature in controlling microbial community succession along a temperature gradient. Furthermore, aside from an increase in temperature, the southward transplantation of soil caused a significant loss in plant productivity, as well as changes of some soil physical and chemical factors (such as cation exchange capacity, soil organic matter and nitrate content).
In conclusion, despite the important roles of soil microbial communities in carbon and nitrogen cycling, as well as their intricate linkage with a variety of ecosystem functions, the stability and temporal succession in microbial communities under continuous disturbances caused by climate change are not fully understood. Microbial community structure was significantly altered by soil transplantation, which simulates climate change. Steeper temporal turnover was observed in northward and southward transplants, and warming posed a more significant impact on microbial succession dynamics. Soil transplant-induced changes in microbial community structure and temporal turnover may be important in predicting long-term ecosystem responses to global change.
References
Allison LE . (1965). Organic carbon. In: Black CA (ed). Methods of Soil Analysis, Part 2, Agronomy 9. American Society of Agronomy, Inc.: Madison, WI, USA, pp 1367–1389.
Balser TC, Firestone MK . (2005). Linking microbial community composition and soil processes in a California annual grassland and mixed-conifer forest. Biogeochemistry 73: 395–415.
Bardgett RD, Freeman C, Ostle NJ . (2008). Microbial contributions to climate change through carbon cycle feedbacks. ISME J 2: 805–814.
Bellard C, Bertelsmeier C, Leadley P, Thuiller W, Courchamp F . (2012). Impacts of climate change on the future of biodiversity. Ecol Lett 15: 365–377.
Bottomley PJ, Yarwood RR, Kageyama SA, Waterstripe KE, Williams WA, Cromack K Jr et al. (2006). Responses of soil bacterial and fungal communities to reciprocal transfers of soil between adjacent coniferous forest and meadow vegetation in the Cascade Mountains of Oregon. Plant Soil 289: 35–45.
Bremner JM . (1965). Total nitrogen, inorganic forms of nitrogen, organic forms of nitrogen, nitrogen availability indexes Black CA (ed). Methods of Soil Analysis, Part2, Agronomy 9. American Society of Agronomy, Inc.: Madison, WI, USA, pp 1149–1348.
Brown JH, Gillooly JF, Allen AP, Savage VM, West GB . (2004). Toward a metabolic theory of ecology. Ecology 85: 1771–1789.
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Huntley J, Fierer N et al. (2012). Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6: 1621–1624.
Castro HF, Classen AT, Austin EE, Norby RJ, Schadt CW . (2010). Soil microbial community responses to multiple experimental climate change drivers. Appl Environ Microbiol 76: 999–1007.
Chapman HD . (1965). Cation exchange capacity. In: Black CA (ed). Methods of Soil Analysis, Part2, Agronomy. American Society of Agronomy, Inc.: Madison, WI, USA, pp 891–900.
Chytry M, Sedlakova I, Tichy L . (2001). Species richness and species turnover in a successional heathland. Appl Veg Sci 4: 89–96.
Clarke KR . (1993). Non-parametric multivariate analysis of changes in community structure. Aust J Ecol 18: 117–143.
Clarke KR, Ainsworth M . (1993). A method of linking multivariate community structure to environmental variables. Mar Ecol-Prog Ser 92: 205–219.
Conant RT, Ryan MG, Ågren GI, Birge HE, Davidson EA, Eliasson PE et al. (2011). Temperature and soil organic matter decomposition rates – synthesis of current knowledge and a way forward. Global Change Biol 17: 3392–3404.
de Vries FT, Manning P, Tallowin JRB, Mortimer SR, Pilgrim ES, Harrison KA et al. (2012). Abiotic drivers and plant traits explain landscape-scale patterns in soil microbial communities. Ecol Lett 15: 1230–1239.
de Vries FT, Shade A . (2013). Controls on soil microbial community stability under climate change. Front Microbiol 4: 265.
Deslippe JR, Hartmann M, Simard SW, Mohn WW . (2012). Long-term warming alters the composition of Arctic soil microbial communities. FEMS Microbiol Ecol 82: 303–315.
Edgar RC . (2010). Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26: 2460–2461.
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R . (2011). UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27: 2194–2200.
Fierer N, Bradford MA, Jackson RB . (2007). Toward an ecological classification of soil bacteria. Ecology 88: 1354–1364.
Fussmann KE, Schwarzmueller F, Brose U, Jousset A, Rall BC . (2014). Ecological stability in response to warming. Nat Clim Change 4: 206–210.
Gonzalez A, King A, Robeson MSI, Song S, Shade A, Metcalf JL et al. (2012). Characterizing microbial communities through space and time. Curr Opin Biotech 23: 431–436.
Goslee S, Urban D . (2007). The ecodist package for dissimilarity-based analysis of ecological data. J Stat Softw 22: 1–19.
Gutknecht JLM, Field CB, Balser TC . (2012). Microbial communities and their responses to simulated global change fluctuate greatly over multiple years. Global Change Biol 18: 2256–2269.
Hanson CA, Fuhrman JA, Horner-Devine MC, Martiny JBH . (2012). Beyond biogeographic patterns: Processes shaping the microbial landscape. Nat Rev Microbiol 10: 497–506.
Hatosy SM, Martiny JBH, Sachdeva R, Steele J, Fuhrman JA, Martiny AC . (2013). Beta diversity of marine bacteria depends on temporal scale. Ecology 94: 1898–1904.
Horner-Devine MC, Lage M, Hughes JB, Bohannan B . (2004). A taxa-area relationship for bacteria. Nature 432: 750–753.
Kanehiro Y, Sherman GD . (1965). Fusion with sodium carbonate for total elemental analysis. In: Black CA (ed). Methods of Soil Analysis, Part 2, Agronomy 9. American Society of Agronomy, Inc.: Madison, WI, USA, pp 952–958.
Kirschbaum MUF . (2004). Soil respiration under prolonged soil warming: are rate reductions caused by acclimation or substrate loss? Global Change Biol 10: 1870–1877.
Kong Y . (2011). Btrim: A fast, lightweight adapter and quality trimming program for next-generation sequencing technologies. Genomics 98: 152–153.
Korhonen JJ, Soininen J, Hillebrand H . (2010). A quantitative analysis of temporal turnover in aquatic species assemblages across ecosystems. Ecology 91: 508–517.
Krug EC, Frink CR . (1983). Acid-rain on acid soil: a new perspective. Science 221: 520–525.
Kruskal JB . (1964). Nonmetric multidimensional scaling: a numerical method. Psychometrika 29: 115–129.
Lazzaro A, Gauer A, Zeyer J . (2011). Field-scale transplantation experiment to investigate structures of soil bacterial communities at pioneering sites. Appl Environm Microbiol 77: 8241–8248.
Leibold MA, Holyoak M, Mouquet N, Amarasekare P, Chase JM, Hoopes MF et al. (2004). The metacommunity concept: a framework for multi-scale community ecology. Ecol Lett 7: 601–613.
Legendre P, Legendre L . (2012), Numerical Ecology. 3rd English Edition. Elsevier.
Letunic I, Bork P . (2011). Interactive Tree Of Life v2: online annotation and display of phylogenetic trees made easy. Nucleic Acids Res 392: W475–W478.
Likens GE, Driscoll CT, Buso DC . (1996). Long-term effects of acid rain: response and recovery of a forest ecosystem. Science 272: 244–246.
Liu S, Wang F, Xue K, Sun B, Zhang Y, He Z et al. (2014). The interactive effects of soil transplant into colder regions and cropping on soil microbiology and biogeochemistry. Environ Microbiol 17: 566–576.
Long X, Chen C, Xu Z, Linder S, He J . (2012). Abundance and community structure of ammonia oxidizing bacteria and archaea in a Sweden boreal forest soil under 19-year fertilization and 12-year warming. J Soil Sediment 12: 1124–1133.
Luo C, Rodriguez-R LM, Johnston ER, Wu L, Cheng L, Xue K et al. (2014). Soil microbial community responses to a decade of warming as revealed by comparative metagenomics. Appl Environ Microb 80: 1777–1786.
MacArthur RH, Wilson E . (1967) The Theory of Island Biogeography. Princeton University Press: Princeton, NJ, USA.
Magoc T, Salzberg SL . (2011). FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27: 2957–2963.
Magurran AE . (2004) Measuring Biological Diversity. Blackwell Publishing: Oxford, UK.
Mcfee WW, Kelly JM, Beck RH . (1977). Acid precipitation effects on soil pH and base saturation of exchange sites. Water Air Soil Poll 7: 401–408.
Nekola JC, White PS . (1999). The distance decay of similarity in biogeography and ecology. J Biogeogr 26: 867–878.
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’Hara RB et al. (2013). Reference Manual for Package ‘vegan’. Version 2.0-6 http://vegan.r-forge.r-project.org/.
Oliver A, Lilley AK, van der Gast CJ . (2012). Species-time relationships for bacteria. In: Ogilvie LA, Hirsch PR (eds). Microbial Ecological Theory: Current Perspectives. Caister Academic Press: Norfolk, UK.
Olsen SR, Cole C, Watanabe FS, Dean L . (1954) Estimation of Available Phosphorus in Soils by Extraction with Sodium Bicarbonate vol. 939 USDA Press: Washington, DC.
Pold G, DeAngelis KM . (2013). Up against the wall: The effects of climate warming on soil microbial diversity and the potential for feedbacks to the carbon cycle. Diversity 5: 409–425.
Rousk J, Frey SD, Bååth E . (2012). Temperature adaptation of bacterial communities in experimentally warmed forest soils. Global Change Biol 18: 3252–3258.
Schindlbacher A, Rodler A, Kuffner M, Kitzler B, Sessitsch A, Zechmeister-Boltenstern S . (2011). Experimental warming effects on the microbial community of a temperate mountain forest soil. Soil Biol Biochem 43: 1417–1425.
Shade A, Caporaso JG, Handelsman J, Knight R, Fierer N . (2013). A meta-analysis of changes in bacterial and archaeal communities with time. ISME J 7: 1493–1506.
Shurin JB . (2007). How is diversity related to species turnover through time? OIKOS 116: 957–965.
Singh BK, Bardgett RD, Smith P, Reay DS . (2010). Microorganisms and climate change: terrestrial feedbacks and mitigation options. Nat Rev Microbiol 8: 779–790.
St. John MG, Orwin KH, Dickie I . (2011). No ‘home’ versus ‘away’ effects of decomposition found in a grassland-forest reciprocal litter transplant study. Soil Biol Biochem 43: 1482–1489.
Strickland MS, Lauber C, Fierer N, Bradford MA . (2009). Testing the functional significance of microbial community composition. Ecology 90: 441–451.
Sun B, Wang F, Jiang Y, Li Y, Dong Z, Li Z et al. (2014). A long- term field experiment of soil transplantation demonstrating the role of contemporary geographic separation in shaping soil microbial community structure. Ecol Evol 4: 1073–1087.
Sun B, Wang X, Wang F, Jiang Y, Zhang X . (2013). Assessing the relative effects of geographic location and soil type on microbial communities associated with straw decomposition. Appl Environ Microb 79: 3327–3335.
Svensson JR, Lindegarth M, Pavia H . (2009). Equal rates of disturbance cause different patterns of diversity. Ecology 90: 496–505.
Székely AJ, Berga M, Langenheder S . (2013). Mechanisms determining the fate of dispersed bacterial communities in new environments. ISME J 7: 61–71.
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S . (2011). MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28: 2731–2739.
Thomas CD, Cameron A, Green RE, Bakkenes M, Beaumont LJ, Collingham YC et al. (2004). Extinction risk from climate change. Nature 427: 145–148.
Vanhala P, Karhu K, Tuomi M, Bjorklof K, Fritze H, Hyvarinen H et al. (2011). Transplantation of organic surface horizons of boreal soils into warmer regions alters microbiology but not the temperature sensitivity of decomposition. Global Change Biol 17: 538–550.
Waldrop MP, Firestone MK . (2006). Response of microbial community composition and function to soil climate change. Microbial Ecol 52: 716–724.
Wang Q, Garrity GM, Tiedje JM, Cole JR . (2007). Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microb 73: 5261–5267.
Wang W, Wang T . (1995). On the origin and the trend of acid precipitation in China. Water Air Soil Poll 85: 2295–2300.
Wardle DA, Bardgett RD, Klironomos JN, Setala H, van der Putten WH, Wall DH . (2004). Ecological linkages between aboveground and belowground biota. Science 304: 1629–1633.
Werner EE, Yurewicz KL, Skelly DK, Relyea RA . (2007). Turnover in an amphibian metacommunity: the role of local and regional factors. OIKOS 116: 1713–1725.
Zhao M, Xue K, Wang F, Liu S, Bai S, Sun B et al. (2014). Microbial mediation of biogeochemical cycles revealed by simulation of global changes with soil transplant and cropping. ISME J 8: 2045–2055.
Zhou J, Bruns MA, Tiedje JM . (1996). DNA recovery from soils of diverse composition. Appl Environ Microb 62: 316–322.
Zhou J, Deng Y, Luo F, He Z, Tu Q, Zhi X . (2010). Functional molecular ecological networks. mBio 1: e00169–10.
Zhou J, Deng Y, Luo F, He Z, Yang Y . (2011a). Phylogenetic molecular ecological network of soil microbial communities in response to elevated CO2 . mBio 2: e00122–11.
Zhou J, Kang S, Schadt CW, Garten CT Jr . (2008). Spatial scaling of functional gene diversity across various microbial taxa. Proc Natl Acad Sci USA 105: 7768–7773.
Zhou J, Wu L, Deng Y, Zhi X, Jiang Y, Tu Q et al. (2011b). Reproducibility and quantitation of amplicon sequencing-based detection. ISME J 5: 1303–1313.
Zhou J, Xue K, Xie J, Deng Y, Wu L, Cheng X et al. (2012). Microbial mediation of carbon-cycle feedbacks to climate warming. Nat Clim Change 2: 106–110.
Zumsteg A, Baath E, Stierli B, Zeyer J, Frey B . (2013). Bacterial and fungal community responses to reciprocal soil transfer along a temperature and soil moisture gradient in a glacier forefield. Soil Biol Biochem 61: 121–132.
Acknowledgements
We thank Yueyu Sui for experiment management in Hailun Agricultural Ecology Experiment Station. This research was supported by Strategic Priority Research Program (B) of the Chinese Academy of Sciences (XDB15030200, XDB15010100), National Science Foundation of China (41271258, 41430856), Foundation for Distinguished Young Talents in State Key Laboratory of Soil and Sustainable Agriculture (Y412010008), the Office of the Vice President for Research at the University of Oklahoma and by the Collaborative Innovation Center for Regional Environmental Quality.
Author Contributions
All authors contributed intellectual input and assistance to this study and manuscript preparation. BS, JZ and YL developed the original framework. YL, YJ, FW, YY, KX, CW, YD, YQ and LW contributed reagents and data analysis. YL, KX and CW did GeoChip analysis. YL, BS and JZ wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Liang, Y., Jiang, Y., Wang, F. et al. Long-term soil transplant simulating climate change with latitude significantly alters microbial temporal turnover. ISME J 9, 2561–2572 (2015). https://doi.org/10.1038/ismej.2015.78
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2015.78
This article is cited by
-
Response of particle-attached and free-living bacterial communities to Microcystis blooms
Applied Microbiology and Biotechnology (2024)
-
Mineral-microbial interactions in nine-year organic fertilization field experiment: a mechanism for carbon storage in saline-alkaline paddy soil
Plant and Soil (2023)
-
Biochar derived from tobacco waste significantly reduces the accumulations of cadmium and copper in edible parts of two vegetables: an in-situ field study
Environmental Science and Pollution Research (2023)
-
Periphyton reduces cyanobacterial blooms by promoting potentially cyanobactericidal bacteria
Journal of Applied Phycology (2023)
-
Diurnal and seasonal gas exchange characteristics of Jatropha curcas leaves
Vegetos (2022)