Abstract
Ribosomal RNA (rRNA) genes are widely utilized in depicting organismal diversity and distribution in a wide range of environments. Although a few cases of lateral transfer of rRNA genes between closely related prokaryotes have been reported, it remains to be reported from eukaryotes. Here, we report the first case of lateral transfer of eukaryotic rRNA genes. Two distinct sequences of the 18S rRNA gene were detected from a clonal culture of the stramenopile, Ciliophrys infusionum. One was clearly derived from Ciliophrys, but the other gene originated from a perkinsid alveolate. Genome-walking analyses revealed that this alveolate-type rRNA gene is immediately adjacent to two protein-coding genes (ubc12 and usp39), and the origin of both genes was shown to be a stramenopile (that is, Ciliophrys) in our phylogenetic analyses. These findings indicate that the alveolate-type rRNA gene is encoded on the Ciliophrys genome and that eukaryotic rRNA genes can be transferred laterally.
Similar content being viewed by others
Main
Genes that are involved in transcription, translation and related processes, and interacted with many partner molecules are believed to be less transferable between different organisms (Rivera et al., 1998; Jain et al., 1999). Hence, lateral transfer of ribosomal RNA (rRNA) genes is generally considered not to occur. Indeed, the lateral transfer of eukaryotic rRNA genes has never been reported, whereas just a few cases of possible lateral transfer of rRNA genes between two closely related prokaryotes have been shown (for example, Mylvaganam and Dennis, 1992; van Berkum et al., 2003). Thus, rRNA gene sequences have been considered as very useful phylogenetic markers and have been utilized in numerous biological studies. Especially, sequence analyses of rRNA genes amplified from environmental DNA have become the gold standard for assessing the diversity and distribution of microbial eukaryotes in a wide range of environments (for example, López-García et al., 2001; Moon-van der Staay et al., 2001).
Recently, we established a clonal culture of Ciliophrys infusionum (deposited as NIES-3355) from the sediment of the lagoon ‘Kai-ike’ (Satsumasendai, Kagoshima, Japan). Ciliophrys infusionum is a heterotrophic microbial eukaryote belonging to Dictyocophyceae (Stramenopiles) and is widely distributed in marine environments. However, as we amplified 18S rRNA gene sequences from this microbial eukaryote to confirm its morphological identification, two distinct 18S rRNA gene sequences (that is, alveolate-type (AB846664: 4418-6616) and stramenopile-type (AB846665)) were unexpectedly obtained. Note that the alveolate-type sequence was obtained first by the PCR with the primer set ‘18S forward and reverse (Yabuki et al., 2010)’ and then the stramenopile-type sequence was obtained by the PCR with the primer set ‘Euk1A (Sogin and Gunderson, 1987) and EukB (Medlin et al., 1988)’. The stramenopile-type sequence reasonably branched with the known sequence of C. infusionum, whereas the alveolate-type sequence branched with perkinsids in Alveolata, microbial eukaryotes parasitic on fishes, bivalves and algae (Figure 1). As neither contaminated nor parasitic/endosymbiotic perkinsids were observed in light and transmission electron microscopy (Supplementary Figure SI 1), the alveolate-type 18S rRNA gene sequence also seemed to be derived from a part of the Ciliophrys genome. Polymorphism of the 18S rRNA gene has been reported for several eukaryotes such as early-branching fish and diatoms (Krieger and Fuerst, 2002; Alverson and Kolnick, 2005). However, the degree of their polymorphism was very low, suggesting that such polymorphism was generated during species diversification. Therefore, the polymorphism found in this study is apparently different from previous cases, and the possibility that the alveolate-type 18S rRNA gene was laterally transferred to the Ciliophrys genome was raised. To confirm the existence of the alveolate-type 18S rRNA gene in the Ciliophrys genome, we obtained both upstream- and downstream-flanking regions of this unexpectedly detected gene from a genome library prepared by restriction enzyme treatment followed by adapter ligation with a Straight Walk Kit (BEX co Ltd, Itabashi, Tokyo, Japan).
Maximum-likelihood tree of 18S rRNA gene sequences (25 alveolates and 26 stramenopiles, 1411 positions) reconstructed by RAxML 7.2.8 (Stamatakis, 2006) with a GTRGAMMAI model selected as the best-fit model by the modeltest 3.7 (Posada and Crandall, 1998). Bootstrap supports were calculated from the analyses of 100 replicates and more than 50% values are shown. Nodes supported by Bayesian posterior probabilities ⩾0.95 are highlighted with bold lines.
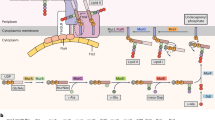
Consequently, a genome fragment (AB846664) containing the alveolate-type 18S rRNA gene (9709 bp in length) was obtained. BLAST searches indicated that two protein-coding genes (that is, ubc12 encoding ubiquitin-conjugating enzyme 12 (UBC12) and usp39 encoding ubiquitin specific peptidase 39 (USP39)) were located in the upstream-flanking region of the alveolate-type 18S rRNA gene. Four and 12 introns were identified in ubc12 and usp39, respectively, and the transcription of both genes was confirmed by RT–PCR with exactly matching primers (data not shown, see experimental procedures in supplement). The physical map of this genome fragment and the structures of ubc12/usp39 are shown in Figures 2a and b, respectively. In the downstream-flanking region, 5.8S rRNA and 28S rRNA genes were identified (Figure 2b). The transcription of the stramenopile-type 18S rRNA gene was confirmed by RT–PCR, whereas the transcription of neither the alveolate-type 18S rRNA nor its adjacent 28S rRNA genes was confirmed (Figure 2c and supplement: note that the primers used were verified to work by positive controls). These findings suggest that the alveolate-type 18S rRNA gene and its adjacent 28S rRNA gene are pseudogenes, although the possibility that these genes are transcribed under different situations (such as different culture conditions) cannot be completely excluded.
(a) Physical map of the genome fragment (AB846664) obtained by genome-walking analyses. Black arrows represent the coding regions of the predicted genes. usp39 is encoded on the strand opposite to other genes. (b) Structures of the transcription unit (18S, 5.8S and 28S rRNA genes), ubc12 and usp39 on the obtained genome fragment are separately shown. White arrows represent coding regions. The possible coding regions of rRNA genes were predicted by comparison with the rRNA gene sequence from Perkinsus atlanticus (AF509333). The sizes and positions of the introns in ubc12 and usp39 were identified by the comparisons between the sequences of genome DNA and RT–PCR products. (c) Summary of PCR amplification of stramenopile-type 18S, alveolate-type 18S and alveolate-type 28S rRNA genes using non-DNase-treated RNA, DNase-treated RNA and cDNA synthesized from the DNase-treated RNA as template. PCR primers exactly matching with the stramenopile-type 18S rRNA gene (S-type 18S F and S-type 18S R), alveolate-type 18S rRNA gene (A-type 18S F and A-type 18S R) and alveolate-type 28S rRNA gene (A-type 28S F and A-type 28S R) were independently designed, and their sequences are shown in the table. (d) Maximum-likelihood tree of 29 UBC12 and 6 UBC11 (outgroup) sequences reconstructed by RAxML 7.2.8 with an LGGAMMAF model selected as the best-fit model by Aminosan (Tanabe, 2011). Bootstrap supports were calculated from the analyses of 100 replicates and more than 50% values are shown. Nodes supported by Bayesian posterior probabilities⩾0.95 are highlighted with bold lines. (e) Maximum-likelihood tree of 33 USP39 sequences reconstructed by RAxML 7.2.8 with an LGGAMMAF model selected as the best-fit model by Aminosan. Bootstrap supports were calculated from the analyses of 100 replicates and more than 50% values are shown. Nodes supported by Bayesian posterior probabilities ⩾0.95 are highlighted with bold lines.
Phylogenetic analyses showed that both UBC12 and USP39 found in this study affiliated with stramenopile homologues (Figures 2d and e). Statistical support for the affiliation of USP39 from C. infusionum with the stramenopile homologues was very high (bootstrap probability (BP) of 95% and Bayesian posterior probability (BPP) of 1.00). Although the support for the affiliation of UBC12 from C. infusionum with the stramenopile homologues was moderate (BP of 66% and BPP of 0.98), its tight affiliation was confirmed by the approximately unbiased test (Supplementary Figure SI2). These results support the idea that the 9709 bp fragment is a part of the Ciliophrys genome and that the alveolate-type 18S rRNA sequence is actually encoded on the Ciliophrys genome. The 28S rRNA gene in the fragment was shown to form a clade with the homologue of Perkinsus with high support (Supplementary Figure SI3), suggesting that this 28S rRNA gene shares the same origin with the alveolate-type 18S rRNA gene. On the basis of our present findings it can be concluded that the lateral transfer of eukaryotic rRNA genes (18S, 5.8S and 28S rRNA genes) occurred from a perkinsid to Ciliophrys as a single evolutionary event.
In the past few years, large-scale environmental rRNA gene surveys conducted with next generation sequencing technology have become common practice (for example, Bråte et al., 2010; Cheung et al., 2010). These large-scale surveys may detect transferred rRNA genes and such transferred rRNA genes may confuse our understanding of the true diversity and distribution of microbial eukaryotes, even if the frequency of lateral transfers of the rRNA gene is rare and the copy numbers of the transferred rRNA gene in environments are low. We agree that environmental rRNA gene surveys with PCR are still useful and effective to estimate the diversity/distribution of microbial eukaryotes. However, the fact that recovered rRNA gene sequences do not always reflect the actual existence of microbial eukaryotes corresponding to these sequences should be kept in mind based on our findings.
References
Alverson AJ, Kolnick L . (2005). Intragenomic nucleotide polymorphism among small subunit (18S) rDNA paralogs in the diatom genus Skeltonema (Bacillariophyta). J Phycol 41: 1248–1257.
Bråte J, Logares R, Berney C, Ree DK, Klaveness D, Jakobsen KS et al (2010). Freshwater Perkinsea and marine-freshwater colonizations revealed by pyrosequencing and phylogeny of environmental rDNA. ISMEJ 4: 1144–1153.
Cheung MK, Au CH, Chu KH, Kwan HA, Wong CK . (2010). Composition and genetic diversity of picoeukaryotes in subtropical coastal waters as revealed by 454 pyrosequencing. ISMEJ 4: 1053–1059.
Jain R, Rivera MC, Lake JA . (1999). Horizontal gene transfer among genomes: the complexity hypothesis. Proc Natl Acad Sci USA 96: 3801–3806.
Krieger J, Fuerst PA . (2002). Evidence of multiple alleles of the nuclear 18S ribosomal RNA gene in sturgeon (Family: Acipenseridae). J Appl Ichthyol 18: 290–297.
López-García P, Rodríguez-Valera F, Pedrós-Alió C, Moreira D . (2001). Unexpected diversity of small eukaryotes in deep-sea Antarctic plankton. Nature 409: 603–607.
Medlin L, Elwood HJ, Stickel S, Sogin ML . (1988). The characterization of enzymatically amplified eukaryotic 16S-like rRNA-coding regions. Gene 71: 491–499.
Moon-van der Staay SY, De Wachter R, Vaulot D . (2001). Oceanic 18S rDNA sequences from picoplankton reveal unsuspected eukaryotic diversity. Nature 409: 607–610.
Mylvaganam S, Dennis PP . (1992). Sequence heterogeneity between the two genes encoding 16S rRNA from the halophilic archaebacterium Haloarcula marismortui. Genetics 130: 399–410.
Posada D, Crandall KA . (1998). Modeltest: testing the model of DNA substitution. Bioinformatics 14: 817–818.
Rivera MC, Jain R, Moore JE, Lake JA . (1998). Genomic evidence for two functionally distinct gene classes. Proc Natl Acad Sci USA. 95: 6239–6244.
Sogin ML, Gunderson JH . (1987). Structural diversity of eukaryotic small subunit ribosomal RNAs. Evolutionary implications. Ann N Y Acad Sci 503: 125–139.
Stamatakis A . (2006). RAxML-VI-HPC: Maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22: 2688–2690.
Tanabe AS . (2011). Kakusan4 and Aminosan: two programs for comparing nonpartitioned, proportional and separate models for combined molecular phylogenetic analyses of multilocus sequence data. Mol Ecol Resour 11: 914–921.
van Berkum P, Terefework Z, Paulin L, Suomalainen S, Lindström K, Eardly BD . (2003). Discordant Phylogenies within the rrn Loci of Rhizobia. J Bacteriol 185: 2988–2998.
Yabuki A, Inagaki Y, Ken-ichiro I . (2010). Palpitomonas bilix gen. et sp. nov.: A novel deep-branching heterotroph possibly related to Archaeplastid or Hacrobia. Protist 161: 523–538.
Acknowledgements
We thank Dr JD Reimer (University of the Ryukyus) for critical comments and English-language corrections. We also thank Messrs N Ishihara and H Kishi (Satsumasendai city office) for their kind and technical helps for sampling in Kai-ike. AY was supported by Research Fellowship from the Japanese Society for the Promotion of Science (JSPS) for Young Scientists (No. 236484). This work was supported by grants from the JSPS awarded to KT (No. 24117527).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Yabuki, A., Toyofuku, T. & Takishita, K. Lateral transfer of eukaryotic ribosomal RNA genes: an emerging concern for molecular ecology of microbial eukaryotes. ISME J 8, 1544–1547 (2014). https://doi.org/10.1038/ismej.2013.252
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2013.252