Abstract
Metabolic specialization is a general biological principle that shapes the assembly of microbial communities. Individual cell types rarely metabolize a wide range of substrates within their environment. Instead, different cell types often specialize at metabolizing only subsets of the available substrates. What is the advantage of metabolizing subsets of the available substrates rather than all of them? In this perspective piece, we argue that biochemical conflicts between different metabolic processes can promote metabolic specialization and that a better understanding of these conflicts is therefore important for revealing the general principles and rules that govern the assembly of microbial communities. We first discuss three types of biochemical conflicts that could promote metabolic specialization. Next, we demonstrate how knowledge about the consequences of biochemical conflicts can be used to predict whether different metabolic processes are likely to be performed by the same cell type or by different cell types. We then discuss the major challenges in identifying and assessing biochemical conflicts between different metabolic processes and propose several approaches for their measurement. Finally, we argue that a deeper understanding of the biochemical causes of metabolic specialization could serve as a foundation for the field of synthetic ecology, where the objective would be to rationally engineer the assembly of a microbial community to perform a desired biotransformation.
Similar content being viewed by others
Main
Metabolic specialization is a general biological principle that applies across every domain of cellular life. Consider a microbial cell residing within the human gut. This cell encounters a myriad of different substrates that could be metabolized to satisfy its energetic and elemental requirements (Rambaud et al., 2006). Yet, even if this cell were near starvation, it would only metabolize a subset of the available substrates (Rambaud et al., 2006). What is the advantage of metabolizing only subsets of the available substrates rather than all of them? What are the underlying causes of metabolic specialization? Can we predict which substrates are likely metabolized by the same cell type and which are likely metabolized by different cell types? Thus far, there are few general principles and rules that address these questions.
The questions posed above have relevance for one of the most perplexing enigmas in microbial ecology: why are some microbial communities so incredibly diverse? Advances in molecular ecology revealed that a single liter of seawater or gram of soil contain thousands of different microbial taxa (Curtis et al., 2002; Gans et al., 2005; Huber et al., 2007). Yet, the mechanisms that promote these levels of diversity are not fully clear (Gudelj et al., 2010). Metabolic specialization provides one plausible explanation for how diversity could be promoted, and is therefore a likely general organizing principle that shapes the assembly of microbial communities.
A substantial body of research has greatly improved our understanding of the causes of metabolic specialization (Elena and Lenski, 2003; Kassen and Rainey, 2004; Gudelj et al., 2010). An important conclusion is that metabolic specialization readily evolves in nearly every conceivable environment and can often be explained by basic ecological and evolutionary principles. This research, however, has largely focused on the ecological and genetic causes of metabolic specialization rather than the biochemical causes (Gudelj et al., 2010). It is clear that biochemical conflicts exist between different metabolic processes and likely promote the evolution of metabolic specialization. Our limited understanding about these biochemical conflicts therefore represents a significant gap in our knowledge.
In this perspective manuscript, we argue that a better understanding of the biochemical conflicts that exist between different metabolic processes could help reveal the general principles and rules that govern the evolution of metabolic specialization. We begin by discussing three types of biochemical conflicts that could promote specialization. We then demonstrate how knowledge about the consequences of biochemical conflicts can be used to predict whether different metabolic processes are performed by the same cell type or by different cell types. We conclude by discussing the major challenges in identifying and assessing biochemical conflicts and propose several approaches for their measurement.
Biochemical conflicts that could promote metabolic specialization
Conflicts resulting from competition for intracellular resources
The synthesis and maintenance of metabolic enzymes requires the consumption of intracellular resources, including elemental building blocks (carbon and nitrogen), energy resources (ATP), mRNA synthesis machinery (RNA polymerase and sigma factors), protein synthesis machinery (amino acids, ribosomes, tRNAs and chaperones) and cellular space for housing enzymes. If one of these resources is limiting, then a cell that invests more resources in one metabolic process must invest fewer resources in other processes. Recent studies support the existence of such competitive resource conflicts (Dekel and Alon, 2005; Scott et al., 2010), but the specific intracellular resources that were under competition were not identified. Below we discuss two that might be of relevance.
Cytoplasmic solvent capacity is one intracellular resource that could lead to competitive resource conflicts. The solvent capacity determines the maximum number of enzymes and other macromolecules that can be contained within the cell (Zhou et al., 2008). If the solvent capacity is exceeded then the biochemical and biophysical properties of macromolecules can change, often with detrimental effects (Zhou et al., 2008). Recent combinations of experiments and modeling of Escherichia coli showed that cells operate near their solvent capacity when grown with high substrate supply (Beg et al., 2007). Under these conditions, cells that are provided with mixtures of different substrates produce enzymes for metabolizing only the most productive substrate (Beg et al., 2007), thus potentially leaving the less productive substrates available for uptake by different cell types. A likely explanation for this type of specialization is that, because the solvent capacity is near saturation, cells that produce more enzymes for less productive pathways must produce fewer enzymes for the most productive pathway. Competition for solvent capacity could therefore promote metabolic specialization under specific substrate supply conditions.
RNA polymerase is another intracellular resource that could lead to competitive resource conflicts. RNA polymerase interacts with different sigma factors to regulate the transcription of different genes (Gruber and Gross, 2003). Studies with E. coli showed that the rpoS and rpoD sigma factors compete for limited RNA polymerase (Ferenci, 2005). rpoS regulates stress-response genes whereas rpoD regulates carbon metabolism genes that determine the metabolic versatility of E. coli. Increasing the expression of rpoS and stress-response genes must therefore coincide with decreasing the expression of rpoD and restricting the metabolic versatility of E. coli, thus resulting in metabolic specialization. Competition for limited RNA polymerase by different sigma factors could therefore promote metabolic specialization under specific stress conditions.
Conflicts resulting from inhibition
The production of inhibitory intermediates and endproducts could also lead to biochemical conflicts between different metabolic processes (Fay, 1992; Pfeiffer and Bonhoeffer, 2004; Costa et al., 2006; MacLean and Gudelj, 2006). One example is the antagonistic effect of oxygenic photosynthesis on nitrogen fixation. Oxygen is produced during photosynthesis but inhibits nitrogen fixation by irreversibly inactivating nitrogenase, which makes it challenging for a cell to perform both processes simultaneously (Fay, 1992). A number of strategies have evolved to overcome this conflict, such as differentiation into different cell types, where one type photosynthesizes and another type fixes nitrogen (Fay, 1992).
Inhibitory conflicts between metabolic processes could also occur if intracellular intermediates are growth inhibiting (Pfeiffer and Bonhoeffer, 2004; Costa et al., 2006). Theoretical models that consider growth-inhibiting intermediates predict that cross-feeding metabolic specialists are more likely to evolve as the inhibiting effects increase (Pfeiffer and Bonhoeffer, 2004; Costa et al., 2006). The explanation for this is that performing an additional metabolic conversion step of a substrate incurs two costs: the cost of synthesizing proteins for the additional step and the cost of producing additional intracellular intermediates. If these costs exceed the energetic gain from the additional step, then it is beneficial to not perform that step and only partially consume the substrate, thus allowing the intermediates to escape and be metabolized by other cell types. Although quantitative measures of these costs are lacking, these predictions are consistent with many empirical observations. For example, consortia of partially consuming cross-feeding cell types are often responsible for degrading pollutants that produce toxic intermediates (De Souza et al., 1998; Pelz et al., 1999).
Conflicts for enzyme specificity
If the same enzyme interacts with different substrates, such as occurs with some membrane transporters (Saier, 2000), then conflicts could occur for enzyme specificity. In this scenario, improving the specificity for one substrate reduces the specificity for another substrate, thus leading to metabolic specialization. Although specificity conflicts are intuitive and provide an explanation for why most enzymes have only a few substrates, there is little experimental support for these conflicts. For example, artificial selection was used to select for improved utilization of alternative substrates for three different enzymes in E. coli (Aharoni et al., 2005). After random mutagenesis and selection, improved utilization of the alternative substrates did not result in reduced utilization of the primary substrates, suggesting that specificity conflicts were not important for these enzymes (Aharoni et al., 2005).
The consequences of biochemical conflicts
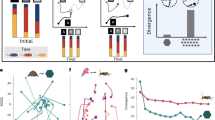
Even though the biochemical conflicts that exist between different metabolic processes are often unknown, knowledge about their consequences can provide important insights into the evolution of metabolic specialization. The consequence of a biochemical conflict is often antagonistic pleiotropy, which describes an outcome where a single genetic change has beneficial effects on some processes but correlated and detrimental effects on others (Cooper and Lenski, 2000). For example, if competitive resource conflicts exist between two processes, then a genetic change that increases the synthesis of one enzyme must simultaneously decrease the synthesis of other enzymes. These effects can be described graphically using constraint functions (Figure 1). A constraint function is a type of trade-off function that describes how the activity levels of different metabolic processes are connected and influenced by biochemical conflicts. Examples of constraint functions for two processes are shown in Figure 1. In these examples, the activity levels are plotted on independent axes for every possible phenotype. A line is then drawn that connects the maximal activities of all possible phenotypes. This line does not define all phenotypes that could be expressed by a single genotype. Instead, it delimits the set of all possible phenotypes that could evolve after long-term selection in any particular direction, and thus encompasses a wide range of genotypes.
The shape of a constraint function, which is a line that connects the maximal activities of all possible phenotypes, describes the consequences of biochemical conflicts on the activity levels of two different metabolic processes. A function consisting of vertical and horizontal lines with positive intercepts describes processes that are not in conflict with each other (solid lines). Note that the maximal activities of both processes are independent of each other. Alternatively, a function consisting of vertical and horizontal lines with zero intercepts describes processes that are in complete conflict with each other and cannot occur within the same cell type (short dashed lines). Lines between these two limiting cases describe processes that are in conflict with each other but can still be performed by the same cell type. Concave functions indicate weak conflicts because the sum of activities when performing both processes is always greater than the maximum activity when performing only one (long dashed line). Convex functions indicate strong conflicts because the sum of activities when performing both processes is always less than when performing only one (medium dashed line).
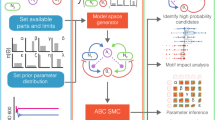
What use are constraint functions for understanding the evolution of metabolic specialization? If the shape of the constraint function is known for a set of metabolic processes, then simple evolutionary models can be used to predict whether specialization is likely to evolve (Doebeli, 2002; Gudelj et al., 2007; Figure 2). For the models discussed in Figure 2, metabolic generalists evolve for concave constraint functions (weak conflicts between processes) whereas metabolic specialists evolve for convex constraint functions (strong conflicts between processes). For the latter case, the evolution of metabolic specialization occurs in two phases, under the assumption that mutations have small pleiotropic effects. A generalist cell type first evolves that performs both processes. Selection then turns disruptive and the population splits into two coexisting cell types, where each type specializes at only one of the processes (Doebeli, 2002).
The shape of a constraint function can be used to predict whether metabolic specialization is likely to occur. Consider the concave and convex functions shown in (a). We used the modeling approach described by Doebeli (2002) to predict the evolutionary fate of the glucose and ribose metabolic pathways (b). The model predicts that the same cell type performs glucose and ribose metabolism if the function is concave, but that these pathways segregate into different cell types over evolutionary time if the function is convex (b). This result is not unique to parallel pathways and also occurs within a single pathway (c). Using the reduction of tetrachloroethene (PCE) to ethene as an example, our model predicts that the same cell type performs the complete pathway if the function is concave, but segregates into different cell types over evolutionary time if the function is convex (c). For (b, c), phenotypes are shown that are present in the population at frequencies >5%.
The mathematical models discussed above can thus be used to predict whether different metabolic processes are likely to be performed by the same cell type or by different cell types. These predictions are general in that they are independent of the kinetic parameters of the model (Doebeli, 2002), and are therefore applicable to any pair of processes. In the simple scenario depicted in Figure 2, the outcome of the model corresponds to what one might expect based on simple considerations: two processes will segregate into different cell types if they are in biochemical conflict. However, if more than two processes are involved (Doebeli and Ispolatov, 2010) or if ecological or genetic aspects are considered (Gudelj et al., 2007), then verbal arguments are insufficient, and mathematical modeling offers a rigorous and objective way to predict evolutionary outcomes.
Generating hypotheses about biochemical conflicts
We have demonstrated that qualitative information about the consequences of biochemical conflicts (for example, the shapes of constraint functions) enable specific predictions about whether different metabolic processes are likely to be performed by the same cell type or different cell types (Figure 2). This raises the question about how such constraints can be measured. In the following section, we discuss two main approaches for obtaining such information.
Genome comparisons across organisms
A first approach that generates hypotheses about possible biochemical conflicts is to analyze the distributions of metabolic processes across microbial genomes. The complete sequences of several thousand microbial genomes are currently available, and automated gene annotation (Markowitz et al., 2006; Aziz et al., 2008) enables the rapid assessment of co-occurrence patterns of different processes across different genomes. One could thus identify combinations of processes that are more often found in the same cell type or in different cell types. The latter are candidates for processes that are in biochemical conflict.
One main advantage of this approach is that it considers information from a large number of species and strains, of which an increasing number are not experimental model systems and cannot be grown in the laboratory. It thus allows rapid and comprehensive assessment of co-occurrence patterns of a large number of metabolic processes. Although promising, this approach has several major caveats. First, the approach assumes that automated genome annotation is sufficient to predict the complete set of metabolic pathways of microorganisms. Although this is increasingly possible for model microorganisms, this remains challenging for non-model microorganisms that contain unusual and poorly characterized enzymes and pathways. Second, this approach does not consider gene expression. Cells could maintain incompatible pathways by expressing them at different times or in different environments. Finally, this approach does not consider the environment in which microorganisms live. The absence of co-occurrence of two processes might result from the absence of their substrates in the same environment rather than a biochemical conflict. The influence of the latter two confounding factors could be lessened by analyzing groups of microorganisms that live in similar environments. For example, one could analyze metagenome information collected from a single environmental sample, provided that genes within the metagenome can be accurately assigned to different cell types (Eisen, 2007).
Experimental evolution
A second and more direct approach to measure biochemical conflicts is through experimental evolution in the laboratory. If one is interested in measuring the constraint function between two metabolic processes, replicated populations can be selected to maximize the first process in isolation, the second process in isolation or both processes simultaneously. This would provide a measure of multiple points on the constraint function (Figure 1), which could then be used to estimate its shape.
Experimental evolution has, to the best of our knowledge, not been systematically used to investigate biochemical conflicts. Yet, we believe this method has potential. First, it offers experimental control. Confounding factors can be excluded by comparing different genotypes that have evolved under well-controlled conditions (Elena and Lenski, 2003; Jessup et al., 2004). Experiments can also be conducted such that the only target of selection is the rate at which a strain grows on one particular substrate or combination of substrates (Ibarra et al., 2002). Second, whole-genome resequencing and metabolic analyses can reveal detailed insights about the genetic changes that occurred over the course of experimental evolution (Herring et al., 2006; Barrick et al., 2009), and can thus help generate hypotheses about the molecular basis of biochemical conflicts. Although the timescale of laboratory evolution experiments does not extend to the timescales of evolutionary processes in natural environments, such experiments can nevertheless give insights into the initial phase of metabolic specialization. This initial phase can determine further evolutionary processes (Travisano et al., 1995; Le Gac and Doebeli, 2010; Khan et al., 2011), and studying this phase experimentally might thus provide information about how biochemical conflicts promote metabolic specialization.
Challenges with measuring biochemical conflicts
Conflicts likely affect many metabolic processes
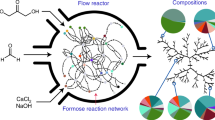
Our discussion of biochemical conflicts thus far has focused on constraining relationships between two metabolic processes (Figures 1 and 2). In reality, constraining relationships likely involve more than two processes (Pease and Bull, 1988) (Figure 3). To illustrate this, consider again one of the examples discussed above. We argued that RNA polymerase could be a limiting intracellular resource for which different sigma factors compete, and that this might lead to a conflict between stress response and metabolic versatility (Ferenci, 2005). One obvious resolution to this conflict is to produce more RNA polymerase. However, the production of RNA polymerase, and gene expression in general, is metabolically costly (Dekel and Alon, 2005), and this resolution might therefore have detrimental effects on other processes. A more realistic model would include a large number of metabolic processes and other cellular traits that are connected through complex interactions, both suppressive and facilitative.
Constraining relations are likely to involve more than two metabolic processes. For example, consider a situation where three processes are connected in a constraining relationship, but where an experimenter only measures two of them (process 1 and 2); the experimenter thus only observes the projections of the constraint functions in the horizontal plane (dashed lines). If the activity of process 3 is maintained constant during the experiment, the experimenter observes a concave constrain function between process 1 and process 2 (green dashed line). Changes in process 3 during the experiment can lead to the observation of qualitatively different constraint functions (red dashed line). Also, simultaneous improvements of process 1 and process 2 beyond their constraint function are possible at the cost of process 3 (orange).
One consequence of the involvement of multiple metabolic processes in constraining relationships is that constraints between two metabolic processes are not absolute; simultaneous improvements in both are possible at the cost of others (Figure 3). The combinations of processes that are observed in a particular cell type represent evolutionary compromises across many different selection pressures. If the environment changes and the strength of selection on some processes increases, then one would expect that it is possible to simultaneously improve these at the cost of other processes that are subject to weaker selection.
Another consequence of the involvement of multiple metabolic processes is that complex interactions might generally promote metabolic specialization. A recent theoretical study showed that the probability of diversification increases as constraining relationships involve more traits (Doebeli and Ispolatov, 2010). Although complex interactions between different metabolic processes are more difficult to experimentally measure, they might generally promote the diversification processes depicted in Figure 2.
Dependence on the environmental conditions
A second challenge with measuring constraining relationships is their dependence on environmental conditions. Constraints are ultimately based on resource conflicts and interactions between different metabolic processes; the environment will affect both of these aspects, and is thus expected to influence constraining relationships both quantitatively and qualitatively. Given the wide range of natural environments, how can we make progress in determining constraining relationships and understanding how they promote metabolic specialization? In our opinion, the main goal is to establish basic principles about how different environmental factors affect biochemical conflicts and influence the main constraining relationships between different processes.
The emerging concept is thus a dynamic view on biochemical conflicts between metabolic processes, on how these conflicts are influenced by environmental factors, and on how they promote the evolution of metabolic specialization and shape the assembly of microbial communities. Each process in a cell is connected to a large number of other processes, and many of these connections will depend on the environmental conditions. Mutations that increase the activity of one process are likely to affect other processes. There might be combinations of mutations that would improve single processes without substantial impairments of other processes, but these genotypes might take a long time to evolve, or they might not be easily accessible by consecutive mutational steps that are all individually beneficial (Weinreich et al., 2006). Such a situation can promote the emergence of mutants that specialize at metabolizing certain substrates. Genetic and ecological interactions might then lead to the consolidation of these differences and the evolution of specialized cell types that consume only subsets of the available substrates within their environment.
The foundation for synthetic ecology?
What is the broader importance of understanding the biochemical causes of metabolic specialization? A deeper understanding of these causes could serve as a foundation for establishing a field of synthetic ecology (Dunham, 2007). In our view, the objective of synthetic ecology would be to elucidate basic design principles that enable the rational engineering of the assembly of microbial communities to perform desired biotransformations. For example, consider a biological process where a microbial population is used to transform a substrate into an intermediate and then into a value-added end product, such as a pharmaceutical or bioenergy source. One strategy would be to engineer one cell type that catalyzes the complete pathway (Ro et al, 2006). An alternative strategy would be to engineer a community of specialized cell types, where one type transforms the substrate into the intermediate and another type then transforms the intermediate into the desired end product (Kato et al., 2005). We currently lack basic design principles that predict which strategy maximizes the performance of such a process. A better understanding of the causes of metabolic specialization will likely help elucidate such engineering design principles and contribute toward establishing a discipline of synthetic ecology.
References
Aharoni A, Gaidukov L, Khersonsky O, Gould SM, Roodveldt C, Tawfik DS . (2005). The ‘evolvability’ of promiscuous protein functions. Nat Genet 37: 73–76.
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA et al. (2008). The RAST server: rapid annotations using subsystems technology. BMC Genomics 9: 75.
Barrick JE, Yu DS, Yoon SH, Jeong H, Oh TK, Schneider D et al. (2009). Genome evolution and adaptation in a long-term experiment with Escherichia coli. Nature 461: 1243–1247.
Beg QK, Vazquez A, Ernst J, de Menezes MA, Bar-Joseph Z, Barabasi AL et al. (2007). Intracellular crowding defines the mode and sequence of substrate uptake by Escherichia coli and constrains its metabolic activity. Proc Natl Acad Sci USA 104: 12663–12668.
Cooper VS, Lenski RE . (2000). The population genetics of ecological specialization in evolving Escherichia coli populations. Nature 407: 736–739.
Costa E, Perez J, Kreft JU . (2006). Why is metabolic labour divided in nitrification? Trends Microbiol 14: 213–219.
Curtis TP, Sloan WT, Scannell JW . (2002). Estimating prokaryotic diversity and its limits. Proc Natl Acad Sci USA 99: 10494–10499.
De Souza ML, Newcombe D, Alvey S, Crowley DE, Hay A, Sadowsky MJ et al. (1998). Molecular basis of a bacterial consortium: Interspecies catabolism of atrazine. Appl Environ Microbiol 64: 178–184.
Dekel E, Alon U . (2005). Optimality and evolutionary tuning of the expression level of a protein. Nature 436: 588–592.
Doebeli M . (2002). A model for the evolutionary dynamics of cross-feeding polymorphisms in microorganisms. Popul Ecol 44: 59–70.
Doebeli M, Ispolatov I . (2010). Complexity and diversity. Science 328: 494–497.
Dunham MJ . (2007). Synthetic ecology: a model system for cooperation. Proc Natl Acad Sci USA 104: 1741–1742.
Eisen JA . (2007). Environmental shotgun sequencing: its potential and challenges for studying the hidden world of microbes. PLoS Biol 5: e82.
Elena SF, Lenski RE . (2003). Evolution experiments with microorganisms: The dynamics and genetic bases of adaptation. Nat Rev Genet 4: 457–469.
Fay P . (1992). Oxygen relations of nitrogen-fixation in cyanobacteria. Microbiol Rev 56: 340–373.
Ferenci T . (2005). Maintaining a healthy SPANC balance through regulatory and mutational adaptation. Mol Microbiol 57: 1–8.
Gans J, Wolinsky M, Dunbar J . (2005). Computational improvements reveal great bacterial diversity and high metal toxicity in soil. Science 309: 1387–1390.
Gruber TM, Gross CA . (2003). Multiple sigma subunits and the partitioning of bacterial transcription space. Annu Rev Microbiol 57: 441–466.
Gudelj I, Beardmore RE, Arkin SS, Maclean RC . (2007). Constraints on microbial metabolism drive evolutionary diversification in homogeneous environments. J Evol Biol 20: 1882–1889.
Gudelj I, Weitz JS, Ferenci T, Horner-Devine MC, Marx CJ, Meyer JR et al. (2010). An integrative approach to understanding microbial diversity: from intracellular mechanisms to community structure. Ecol Lett 13: 1073–1084.
Herring CD, Raghunathan A, Honisch C, Patel T, Applebee MK, Joyce AR et al. (2006). Comparative genome sequencing of Escherichia coli allows observation of bacterial evolution on a laboratory timescale. Nat Genet 38: 1406–1412.
Huber JA, Mark Welch D, Morrison HG, Huse SM, Neal PR, Butterfield DA et al. (2007). Microbial population structures in the deep marine biosphere. Science 318: 97–100.
Ibarra RU, Edwards JS, Palsson BO . (2002). Escherichia coli K-12 undergoes adaptive evolution to achieve in silico predicted optimal growth. Nature 420: 186–189.
Jessup CM, Kassen R, Forde SE, Kerr B, Buckling A, Rainey PB et al. (2004). Big questions, small worlds: microbial model systems in ecology. Trends Ecol Evol 19: 189–197.
Kassen R, Rainey PB . (2004). The ecology and genetics of microbial diversity. Annu Rev Microbiol 58: 207–231.
Kato S, Haruta S, Cui ZJ, Ishii M, Igarashi Y . (2005). Stable coexistence of five bacterial strains as a cellulose-degrading community. Appl Environ Microbiol 71: 7099–7106.
Khan AI, Dinh DM, Schneider D, Lenski RE, Cooper TF . (2011). Negative epistasis between beneficial mutations in an evolving bacterial population. Science 332: 1193–1196.
Le Gac M, Doebeli M . (2010). Epistasis and frequency dependence influence the fitness of an adaptive mutation in a diversifying lineage. Mol Ecol 19: 2430–2438.
MacLean RC, Gudelj I . (2006). Resource competition and social conflict in experimental populations of yeast. Nature 441: 498–501.
Markowitz VM, Korzeniewski F, Palaniappan K, Szeto E, Werner G, Padki A et al. (2006). The integrated microbial genomes (IMG) system. Nucleic Acids Res 34: D344–D348.
Pease CM, Bull JJ . (1988). A critique of methods for measuring life-history trade-offs. J Evol Biol 1: 293–303.
Pelz O, Tesar M, Wittich RM, Moore ERB, Timmis KN, Abraham WR . (1999). Towards elucidation of microbial community metabolic pathways: unravelling the network of carbon sharing in a pollutant-degrading bacterial consortium by immunocapture and isotopic ratio mass spectrometry. Environ Microbiol 1: 167–174.
Pfeiffer T, Bonhoeffer S . (2004). Evolution of cross-feeding in microbial populations. Am Nat 163: E126–E135.
Rambaud JC, Buts JP, Corthier G, Flourie B . (2006) Gut Microflora: Digestive Physiology and Pathology. John Libbey Eurotext: Paris.
Ro DK, Paradise EM, Ouellet M, Fisher KJ, Newman KL, Ndungu JM et al. (2006). Production of the antimalarial drug precursor artemisinic acid in engineered yeast. Nature 440: 940–943.
Saier MH . (2000). Families of transmembrane sugar transport proteins. Mol Microbiol 35: 699–710.
Scott M, Gunderson CW, Mateescu EM, Zhang ZG, Hwa T . (2010). Interdependence of cell growth and gene expression: origins and consequences. Science 330: 1099–1102.
Travisano M, Mongold JA, Bennett AF, Lenski RE . (1995). Experimental tests of the roles of adaptation, chance, and history in evolution. Science 267: 87–90.
Weinreich DM, Delaney NF, Depristo MA, Hartl DL . (2006). Darwinian evolution can follow only very few mutational paths to fitter proteins. Science 312: 111–114.
Zhou HX, Rivas GN, Minton AP . (2008). Macromolecular crowding and confinement: Biochemical, biophysical, and potential physiological consequences. Annu Rev Biophys 37: 375–397.
Acknowledgements
We acknowledge the Swiss National Science Foundation for providing funding.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Johnson, D., Goldschmidt, F., Lilja, E. et al. Metabolic specialization and the assembly of microbial communities. ISME J 6, 1985–1991 (2012). https://doi.org/10.1038/ismej.2012.46
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2012.46
Keywords
This article is cited by
-
A dirty job: dishwasher wastewater reuse and upcycle through an ad hoc engineered microbial consortium
npj Clean Water (2023)
-
Multi-genome metabolic modeling predicts functional inter-dependencies in the Arabidopsis root microbiome
Microbiome (2022)
-
Initial community composition determines the long-term dynamics of a microbial cross-feeding interaction by modulating niche availability
ISME Communications (2022)
-
Nutritional stress induced intraspecies competition revealed by transcriptome analysis in Sphingomonas melonis TY
Applied Microbiology and Biotechnology (2022)
-
Gram-negative bacteria associated with a dominant arboreal ant species outcompete phyllosphere-associated bacteria species in a tropical canopy
Oecologia (2021)