Abstract
Microbial oxidation is the only biological sink for atmospheric methane. We assessed seasonal changes in atmospheric methane oxidation and the underlying methanotrophic communities in grassland near Giessen (Germany), along a soil moisture gradient. Soil samples were taken from the surface layer (0–10 cm) of three sites in August 2007, November 2007, February 2008 and May 2008. The sites showed seasonal differences in hydrological parameters. Net uptake rates varied seasonally between 0 and 70 μg CH4 m−2 h−1. Greatest uptake rates coincided with lowest soil moisture in spring and summer. Over all sites and seasons, the methanotrophic communities were dominated by uncultivated methanotrophs. These formed a monophyletic cluster defined by the RA14, MHP and JR1 clades, referred to as upland soil cluster alphaproteobacteria (USCα)-like group. The copy numbers of pmoA genes ranged between 3.8 × 105–1.9 × 106 copies g−1 of soil. Temperature was positively correlated with CH4 uptake rates (P<0.001), but had no effect on methanotrophic population dynamics. The soil moisture was negatively correlated with CH4 uptake rates (P<0.001), but showed a positive correlation with changes in USCα-like diversity (P<0.001) and pmoA gene abundance (P<0.05). These were greatest at low net CH4 uptake rates during winter times and coincided with an overall increase in bacterial 16S rRNA gene abundances (P<0.05). Taken together, soil moisture had a significant but opposed effect on CH4 uptake rates and methanotrophic population dynamics, the latter being increasingly stimulated by soil moisture contents >50 vol% and primarily related to members of the MHP clade.
Similar content being viewed by others
Introduction
Methane is the second most important anthropogenic greenhouse gas after CO2, with a 25 times larger 100-year radiative forcing than CO2 (IPCC, 2007). The atmospheric CH4 concentration has increased from pre-industrial 0.7 to currently about 1.8 parts per million of volume (ppmv) (Blunier et al., 1993). The global budget of atmospheric methane is in the order of 500–600 Tg CH4 yr−1 (Wang et al., 2004). Tropospheric photochemical oxidation of CH4 is the major sink (>80% of the total) of atmospheric methane; additional sinks are diffusion into the stratosphere and microbial oxidation in soils (Conrad, 2009). Methane oxidation by methanotrophic bacteria in aerated upland soils is estimated to be in the range of 15 to 45 Tg yr−1 (Wuebbles and Hayhoe, 2002), equaling 3–9% of the annual sink. Understanding the activity and population dynamics of atmospheric methane oxidizers is important because the soil uptake of atmospheric CH4 is close to the methane source–sink imbalance that gave rise to the increase in atmospheric methane concentration of about 1% per year over the last century (Dörr et al., 1993). Typical soil ecosystems that are permanently or periodically a sink for atmospheric methane are, for example, forests, grasslands and meadows, savannah and desert.
There is increasing evidence that as-yet uncultivated types of methanotrophs are abundant in upland soils, besides Methylocystis spp. Some Methylocystis strains have been shown to oxidize atmospheric methane for extended periods without decline in their activity (Knief and Dunfield, 2005; Baani and Liesack, 2008). Two of the uncultured groups are upland soil cluster alphaproteobacteria (USCα) and gammaproteobacteria (USCγ), as revealed by cultivation-independent surveys of pmoA diversity. pmoA encodes the β-subunit of particulate methane monooxygenase, the key enzyme in methane oxidation. In the pmoA tree, pmoA of USCα groups with pmoA from type II methanotrophs and Methylocapsa, while pmoA of USCγ is affiliated with pmoA from type I methanotrophs (Knief et al., 2003). USCα sensu stricto is defined by the pmoA clade RA14 (Holmes et al., 1999; Henckel et al., 2000; Bourne et al., 2001; Knief et al., 2003; Jaatinen et al., 2004). Two additional pmoA clades of uncultivated methanotrophs form a monophyletic cluster with USCα: JR1 and MHP (Chen et al., 2008; Degelmann et al., 2010). JR1, also termed cluster 5, was first detected in California upland grassland soil (Horz et al., 2005; Knief et al., 2006; Degelmann et al., 2010), while MHP was initially detected in moorlands in the United Kingdom (Chen et al., 2008). In this study, we will refer to the monophyletic lineage composed of RA14, JR1 and MHP as the USCα-like group.
Environmental factors affecting the soil methane sink have been assessed comprehensively during the last two decades, particularly in forest soils (reviewed in Dunfield, 2007). However, little is known about seasonal population dynamics of methanotrophic communities in upland soils and how the composition and abundance of atmospheric methane oxidizers are affected by environmental variables (Kolb, 2009). In this study, seasonal changes in net CH4 uptake rates and in the composition and abundance of the underlying methanotrophic communities were assessed in relation to relevant soil variables. Non-grazed, extensively managed grassland near Giessen (Germany) was selected for analysis. Long-term records have shown that carbon and nitrogen contents, and grass species are uniformly distributed across the entire study area. However, the hydrological conditions are a major factor that varies across the study area, which is due to a slight slope inclination and changes in soil texture (Jäger et al., 2003; Kammann et al., 2008, 2009).
In a previous study, the specific affinity of atmospheric CH4 oxidation was highest in the soil surface layer (0–10 cm) and decreased with depth (Horz et al., 2002). Therefore, our research focused on methanotroph dynamics in the surface layer, using a combination of process-oriented measurements and molecular techniques in field and laboratory studies. Field studies involved measurements of methane uptake and continuous recording of temperature, volumetric soil moisture content and methane concentration in the soil surface layer. In addition, pH, gravimetric moisture content, carbon/nitrogen contents and potential methane oxidation rates were determined at samplings. Changes in methanotroph diversity and abundance were assessed by terminal restriction fragment length polymorphism (T-RFLP) analysis, cloning and Sanger sequencing, 454 pyrosequencing and quantitative real-time polymerase chain reaction (PCR) of pmoA genes.
Materials and methods
Site description and soil sampling
The study area (Environmental Monitoring and Climate Impact Research Station Linden) is non-grazed, extensively managed grassland (50°32′N and 8°41.3′E; 172 m a.s.l.) near Giessen, Germany. Since 1996, granular mineral N-fertilizer (calcium ammonium nitrate, 40 kg N ha−1 yr−1) has been applied in mid-April each spring. Soil physical properties have been detailed previously (Horz et al., 2002; Kammann et al., 2008, 2009). The three sampling sites (I, II and III) were located 40–50 m apart from each other and differed in moisture content. Samples of the 0–10 cm surface layer were taken in 28 August 2007, 29 November 2007, 25 February 2008 and 20 May 2008. Triplicate soil cores taken from each sampling site sized 3 cm in diameter and were immediately stored on ice. Upon arrival in the laboratory, each core (3 replicate cores × 3 sites per sampling event) was homogenized manually by thorough physical mixing (Cavagnaro et al., 2007; Delmont et al., 2011). Aliquots were used immediately for measurements of methane oxidation potential, pH, and C, N and moisture contents. The remaining soil of each replicate core was stored at −80 °C until used for DNA extraction.
Analysis of soil physical parameters
Soil temperature was measured in 10 cm depth (sensor Pt 100 DIN 43760). Volumetric soil moisture was recorded daily with n=4 TDR-sensors per site (Imko, Ettlingen, Germany; type P2G, 0–15 cm depth). In addition, sample-specific gravimetric moisture content of the soil samples was determined as described earlier (Shrestha et al., 2007). Soil pH was measured by preparing a suspension of 1 g of wet soil in 9 ml of demineralized water (Chen et al., 2008). C and N contents were analyzed by dry combustion with a macro-analyzer (type Vario MAX; Elementar Analysen Systeme GmbH, Hanau, Germany).
In situ CH4 concentration and CH4 uptake rate measurements
For the in situ CH4 concentration measurement, soil air was sampled from depths of 5 and 10 cm in weekly intervals using permanently installed silicone soil-air samplers (detailed method description: Kammann et al., 2001a). Samplers were located approximately 1 m away from each sampling site.
In addition, permanently installed silicon soil-air sampling devices were used to weekly obtain representative vertical CH4 profiles (depth interval: 5, 10, 20, 30, 40 and 50 cm). The profiles were located approximately 10 m apart from sampling site I. Net CH4 fluxes were measured using a modified closed chamber method (Hutchinson and Mosier, 1981). Black, conical-shaped chambers (100 cm basical diameter, 80 cm top diameter and 38 cm height) equipped with a battery-driven ventilator and a small vent for pressure equilibration were placed in U-shaped stainless-steel frames filled with water (see Regan et al. (2011) for details). Measurements were made bi-weekly by withdrawing three gas samples from each chamber (tclosure=40 min) at times t0, tc/2 and tc, using 60-ml syringes (Becton/Dickinson Plastipak, Heidelberg, Germany) fitted with three-way stopcocks.
All gas samples were analyzed within 24 h after collection, using a GC equipped with an ECD and FID (FID flame temperature 280 °C). Methane uptake rates were calculated by linear regression and the ideal gas law with average chamber temperature and average air pressure during the cover period.
Measurement of potential methane oxidation rates
Potential methane oxidation rates were measured using samples from November 2007, February 2008 and May 2008. Homogenized soil of triplicate cores was mixed to produce a composite soil sample for each site and sampling event. Approximately 10 g of wet soil was mixed with 10 ml of distilled water in 120-ml serum vials (Bull et al., 2000; Horz et al., 2002). The vials were capped with butyl rubber stoppers, and CH4 was injected into the gas headspace of the vials to give final mixing ratios of 10, 100, 1000 and 10 000 ppmv. Duplicate flasks were prepared at each mixing ratio, resulting in a total of 24 flasks per sampling event (3 sampling sites × 2 replicates per measurement × 4 different CH4 mixing ratios), along with four duplicate blanks containing only water, one set for each of the four different CH4 mixing ratios. The serum vials were incubated on a rotary shaker (150 r.p.m.) at 25 °C in the dark. Methane consumption was measured at 0 h, 3 h, 1 day, 2 days and up to 21 days after CH4 addition. CH4 depletion in the duplicate flasks was used to calculate the linear regression. The potential CH4 oxidation rate of each measurement was determined from the slope of the regression line (Eller and Frenzel, 2001; Horz et al., 2002; Shrestha et al., 2010). After completion of the measurements, total DNA was extracted from the eight slurries made from the soil of site III of the February 2008 sampling, followed by 454 pyrosequencing as described below.
Molecular techniques
The pmoA-based analysis of methanotroph diversity and abundance involved extraction of total DNA, PCR of pmoA gene, cloning and Sanger sequencing, 454 pyrosequencing, T-RFLP fingerprinting and real-time PCR. Each replicate soil core was processed and treated independently, resulting in T-RFLP analysis and real-time PCR in data sets from three biological replicates for each site and sampling event. Detailed description of these methods and sequence data deposition is provided in Supplementary Information (Supplementary Text 1). Total DNA was extracted from each soil core, using the FastDNA SPIN kit for soil (MoBio Laboratories, Carlsbad, CA, USA). As the result of an initial survey (Supplementary Text 2), the primer set A189f/A650r was used throughout the study to amplify pmoA, resulting in an amplicon size of 502 bp. T-RFLP analysis was performed as described by Shrestha et al. (2008). The numbers of pmoA and bacterial 16S rRNA gene copies were quantified by real-time PCR using the primer sets A189f/A650r (pmoA) and 519f/907r (16S rRNA), respectively. Three clone libraries were constructed from composite DNA of triplicate cores of site III, each library representing one of the following three sampling events: August 2007 (65 clones), November 2007 (58 clones) and May 2008 (61 clones). The 184 pmoA clones were fully sequenced. Composite PCR products from each site and seasonal sampling were analyzed by 454 pyrosequencing. In addition, the methanotroph composition in slurries incubated at four different methane mixing ratios (10, 100, 1000 and 10 000 ppmv) was assessed by 454 pyrosequencing. After removal of short sequences (<250 bp in length), a total of 4681 (seasonal samplings) and 927 (slurries) pyrosequencing reads were used for cluster analysis. Comparative sequence analysis was performed using the ARB software package (available at http://www.arb-home.de; Ludwig et al., 2004) and a manually curated pmoA database containing >3000 sequences. Representative pmoA gene sequences obtained in this study have been deposited in the EMBL, GenBank and DDBJ nucleotide sequence databases under the accession numbers FR720089 to FR720307.
Statistical analysis
One-way analysis of variance (SPSS, version 11.5) was used for testing significant differences in the methane oxidation potentials and gene copy numbers between sampling sites and seasons. Spearman's rank-order correlation coefficient was used to test for correlations of moisture content with methane oxidation potential and pmoA gene copy number, using an online tool (http://faculty.vassar.edu/lowry/corr_rank.html). P-values ⩽0.05 were considered statistically significant.
All other statistical analyses were performed in R (version 2.9.1; R Development Core Team, Vienna, Austria) using vegan (Oksanen, 2008). Environmental variables (moisture content, pH, temperature and C/N ratio) were graphically correlated to T-RLFP data by constrained correspondence analysis. The significance test for the correlation between different environmental factors and changes in the T-RFLP patterns was performed by analysis of variance. Permutation test for constrained correspondence analysis was conducted under reduced model using 1000 permutations.
Results
Soil moisture content
In general, moisture content was always greater in site III than in sites I and II throughout the study period (Figure 1a). At the time when soil was sampled for molecular analysis (November 2007), site III exhibited moisture values near field capacity (60–70 vol%; Figure 1a), while the sites I and II did not (hatched background in Figure 1a; Supplementary Figure 1a).
Volumetric soil moisture (a), soil-air CH4 concentrations (b) and CH4 fluxes (c) in the three sampling sites in 2007 and 2008. (a) The volumetric soil moisture in 0–15 cm depth was measured daily at all three sites during the monitoring period. Hatched area indicates values around field capacity; values above 65 vol% indicate that field capacity was reached and the clayey soil swelled in response to high moisture content. Green arrows indicate biomass harvest dates. (b) Mean soil-air CH4 concentrations in 5 cm depth±s.e. (n=4 samplers per site). There was no significant difference in the CH4 concentration patterns between 5 and 10 cm depth. The concentrations at 10 cm depth were always slightly lower than at 5 cm depth (data not shown). (c) Mean monthly CH4 flux rates±s.e. (n=3 maxi-chambers per site) calculated from bi-weekly measurements. Negative rates indicate net CH4 uptake into the soil (CH4 oxidation dominates), while positive CH4 flux rates indicate net emission into the atmosphere (CH4 production dominates). (a–c) Red arrows mark the four dates at which soil was sampled for molecular analysis.
The soil moisture at all three sites showed clear seasonal patterns with low soil moisture during spring and summer (May–September) and relatively higher soil moisture during winter season (October–April) (Figure 1a). The soil moisture pattern reflects well the higher evapotranspiration rates during summer and at times when the grassland canopy is high (harvest dates (May/June and August/September)). However, it also reflects heavy rainfall events; for example, in July 2007, August 2007 and June 2008, when soil moisture showed a sudden increase (Supplementary Figure 2). April 2007 was the driest April ever recorded at this site since 1993 (0 mm), while April (spring) 2008 was wetter than the long-term average (Supplementary Figure 1b). During the following May, the soil dried as usual (Figure 1a).
In situ CH4 concentration and CH4 uptake rates
Soil-air CH4 concentrations at 5 and 10 cm depth in all sites were below-atmospheric (<1.8. ppmv) levels, with very few exceptions of above-atmospheric values (Figure 1b).
In general, in situ CH4 uptake rates (that is, negative values in Figure 1c) were always greater in site I than in sites II and III throughout the study period (January 2007–December 2008) (Figure 1c). The CH4 uptake rate was very low during winter (approximately 0 to −13 μg CH4 m−2 h−1), but was higher in summer (approximately −23 to −58 μg CH4 m−2 h−1), in good correspondence with soil moisture and temperature (Figures 1a and c). Moisture content was negatively and temperature positively correlated with the CH4 uptake rate (Figure 2; see also Supplementary Figure 3).
Correlation analyses between net CH4 fluxes (uptake rates) and (i) soil moisture and (ii) soil temperature. (a) Contour plot (smoothing procedure: running average) where the color indicates the magnitude of the CH4 flux rate (blue: negative rate=uptake; red=no uptake; zero was the ‘highest’ observed rate). (b) Regression analysis showing the relationship between plot-mean fluxes (combined data for sites I, II and III between January 2007 and December 2008) and plot-mean soil moisture and soil temperature (n=559; R2=0.47; P<0.001; see Supplementary Figure 3 for site-specific analyses).
pmoA sequence diversity
The methanotroph diversity in the seasonal samples was assessed using 184 pmoA Sanger sequences and 356 representative pmoA pyrosequences. Comparative analysis grouped all 540 pmoA sequences into 34 distinct clusters, using a nucleotide sequence identity of 92% as a cutoff in pairwise comparison. These clusters were considered to represent distinct genotypic populations. A single representative of each population-level cluster was used to define 12 species-level units (SL-1 to SL-12) based on 7% divergence of inferred amino-acid sequences, corresponding to 87% nucleotide sequence divergence for known methanotrophs (Degelmann et al., 2010). All species-level units were defined by sequence types obtained from both Sanger sequencing and 454 pyrosequencing, except for SL-8 and SL-10 that were detected only by 454 pyrosequencing.
Seven species-level units (SL-1 to SL-7) represented USCα-like populations related to the RA14, MHP and JR1 clades (Figure 3). The nucleotide and derived amino-acid sequence divergences between the three clades ranged from 77% to 80% (Supplementary Table 1). SL-3 could not be unambiguously assigned to either JR1 or MHP. SL-8 belonged to the Methylocystis/Methylosinus group. In the pmoA tree, SL-9 had a common branch point with the pmoA1 and pmoA2 sequences of Methylocystis/Methylosinus and with pmoA of Methylocapsa and the USCα-like cluster, but SL-9 could not be unambiguously assigned to one of these groups. SL-10 represents pmoA2 of type II methanotrophs. The branching of SL-11 and SL-12 relates to clusters 1 and 2, respectively, in Knief et al. (2006). In that study, these lineages were detected in deciduous forest soil. They could not be affiliated with one of the known methanotroph groups. In the pmoA tree, they branched in an intermediate position between pmoA sequences of described methanotrophs and amoA sequences of autotrophic ammonia oxidizers (see Figure 1 in Knief et al. (2006)).
Maximum-likelihood tree showing the phylogenetic affiliation of pmoA sequences obtained from seasonally sampled grassland soil. Initially, pmoA trees were constructed based on 165-derived amino-acid sequence positions of 215 partial pmoA genes, of which 184 sequences were obtained in the course of this study by cloning and Sanger sequencing. Tree topologies were similar regardless of the method (Tree-Puzzle, Neighbor-Joining or Maximum Likelihood) used for tree construction. Therefore, the maximum-likelihood tree was used to insert a total of 356 shorter sequences (85–159 deduced amino acids) into the existing pmoA tree. Sequences were inserted using parsimony without allowing changes in the tree topology. Sequences obtained in the course of this study were grouped into 12 species-level (SL) units based on a sequence identity threshold of 93%. All species-level units, except for SL-3, SL-4 and SL-7, contain published pmoA sequences. Predicted T-RF lengths based on MspI are shown next to the sequences. The numbers of population-level clusters assigned to the different USCα-like species-level units, SL-1 to SL-7, are given within parenthesis. The framed box shows the branching pattern and T-RF sizes within the MHP clade (SL-4 and SL-5) in more detail. The number of pyrosequencing reads assigned to each of the non-USCα-like species-level units, SL-8 to SL-12, was (in total) <70. The 454 pyrosequencing was also used to detect methanotrophs in samples incubated at 10, 100, 1000 and 10 000 ppmv CH4 (measurement of methane oxidation potential). The total numbers of pyrosequencing reads assigned to either type II methanotrophs or USCα-like populations are shown in relation to the different methane concentrations. The amoA sequence of Nitrosomonas communis (AF272399) was used as outgroup. The scale bar represents 10% sequence divergence.
Notably, 167 of 184 pmoA sequences (91%) obtained from site III by cloning and Sanger sequencing belonged to USCα-like populations, while 10 pmoA sequences were assigned to SL-9 and the remaining seven sequences belonged to SL-11 and SL-12. None of these 184 pmoA sequences grouped with pmoA of type II methanotrophs of the Methylocystis/Methylosinus group or with pmoA of type I methanotrophs. The overwhelming dominance of USCα-like populations was confirmed by 454 pyrosequencing for all sites and seasons, with 4497 of 4681 pmoA sequences (96%) being assigned to the RA14, MHP and JR1 clades. The proportion of USCα-like pmoA sequences ranged from 90% to 99% for the different sites and seasons, except for site I in May 2008. For the latter, the 454 library contained, in addition to USCα-like pmoA sequences (66%), a high percentage of pmoA sequences related to SL-11 and SL-12 (34%).
pmoA-based T-RFLP analysis
A total of seven different T-RFs were detected across all sites and seasons (Figure 4). The profiles were highly similar for all sites in August 2007, but had changed drastically for site III, and less pronounced for sites I and II, in November 2007 and February 2008. The different T-RFs were assigned to particular pmoA clades and species-level units by using the pmoA data set obtained from site III by cloning and Sanger sequencing. These assignments were consistently confirmed by the pmoA data set obtained by 454 pyrosequencing (compare Figure 4 with Figure 3).
Bar diagram of pmoA-based T-RFLP fingerprint patterns obtained for each sampling site (I, II and III) and date (August and November 2007, and February and May 2008). The percentage abundances (mean±s.d.; n=3) of seven distinguishable T-RFs are indicated by different color. See Figure 3 for affiliating particular T-RFs with defined methanotroph populations.
In November 2007 and February 2008, the major changes in the T-RFLP profiles of site III were related to an increase in the relative abundances of the 34-, 159- and 446-bp T-RFs, almost covering 90% of total T-RF abundance. The 159- and 446-bp T-RFs were unambiguously assigned to members of the MHP clade (framed box in Figure 3). The 34-bp T-RF could not be assigned unambiguously. PmoA sequences related to RA14 (SL-1, SL-2), SL-3, MHP (SL-4) and SL-9 contributed to the 34-bp T-RF.
The T-RFLP pattern from site III changed between February and May 2008, defined by an increase in the relative abundances of the 208- and 506-bp T-RFs (note that 506 bp is the experimentally determined T-RF size and relates to no-restriction site in the 502-bp pmoA amplicon). These T-RFs were minor components in the T-RFLP patterns obtained from previous sampling events (Figure 4). Neither Sanger nor 454 pyrosequencing identified pmoA sequences that could be assigned to the 208- and 506-bp T-RFs.
Seasonal changes in the relative abundances of T-RFs at sites I and II were clearly less pronounced than at site III, although the 159-, 208- and 446-bp T-RFs appeared and disappeared in T-RFLP patterns obtained from sites I and II at the different samplings (Figure 4).
Ordination analysis of T-RFLP patterns
The triplicate T-RFLP data sets obtained for each site and season were analyzed by ordination-based constrained correspondence analysis to assess the influence of environmental factors on the composition of the methanotrophic community. Constrained correspondence analysis confirmed that T-RFLP fingerprinting is a highly reproducible method and showed that data analysis was not strongly affected by spatial heterogeneities in methanotroph community composition between the three soil cores sampled and processed independently for each site and season (Supplementary Figure 4). Analysis of variance showed that moisture content significantly correlated with community change (R2: 0.73; P<0.001), but none of the other environmental factors, including temperature, pH and C/N ratio. The soil temperatures on the four sampling days were 15.7–17.0 °C (28 August 2007), 3.0–3.9 °C (29 November 2007), 4.6–5.3 °C (25 February 2008) and 13.3–14.7 °C (20 May 2008). The temperatures differed by no more than ±1 °C between the three sites for a given sampling day. The pH varied between 6.1 and 7.1, and the C/N ratio was between 9 and 12.
Quantitative PCR of pmoA gene
The pmoA and 16S rRNA gene copy numbers ranged from 3.8 × 105–1.9 × 106 and 1.5 × 108–9.7 × 108 copies g−1 of dry soil, respectively (Figure 5). There was a tendency toward an increase in pmoA and 16S rRNA gene copy numbers in all three sites from August 2007 to February 2008. This was followed by a significant decline in pmoA and 16S rRNA gene copy numbers in May 2008, except for the 16S rRNA gene copy number in site II. The seasonal changes in pmoA and 16S rRNA gene copy numbers were most pronounced at sampling site III.
Seasonal changes in (a) pmoA and (b) 16S rRNA gene abundances (mean±s.d.; n=3) in sites I, II and III. The pmoA gene copy numbers are shown in relation to the results of methane oxidation potential measurements at 100 ppmv CH4. The means were compared for site-specific differences and seasonal changes by one-way analysis of variance, followed by Duncan's post hoc test. Means with the same letters on top of the bars indicate no significant differences, while different letters indicate significant differences between the samples that were compared (P<0.01), that is, between the same site over the different seasons (seasonal changes, letters on left-hand side) and between different sites in the same season (site-specific differences, letters on right-hand side).
There was no significant difference (P>0.05) in pmoA gene copy numbers between the three study sites in August 2007 and February 2008. However, a significantly higher pmoA gene copy number (P<0.05) was evident for site III, compared with sites I and II, in samples from November 2007 and May 2008. The pmoA gene copy numbers were significantly higher (P<0.05) at site III in November 2007 and February 2008 than in August 2007 and May 2008.
Potential methane oxidation rates
The initial rates of methane oxidation potential calculated for the slurries incubated under a headspace of 10 and 100 ppmv CH4 were always higher in samples from site III than in those from sites I and II (Supplementary Table 2). Methane consumption immediately began after the incubation was started. The initial rates showed a positive correlation with soil moisture content (R2: 0.78; P<0.05) and seasonal changes in pmoA gene copy numbers (R2: 0.75; P<0.05) (Figure 5a).
The initial and induced potential methane oxidation rates calculated for the slurries of site III under a headspace of 1000 and 10 000 ppmv CH4 showed the same trend as the initial rates calculated for the slurries under a headspace of 10 and 100 ppmv CH4. They were always higher than those calculated for sites I and II, except for the initial rate of the November 2007 sample of site III under a headspace of 1000 ppmv CH4 (Supplementary Table 2). There was a sudden increase in the CH4 consumption rates after the initial phase of 4 days or shortly thereafter (Supplementary Figure 5).
Changes in the methanotrophic communities in response to the different headspace CH4 concentrations were assessed by 454 pyrosequencing, using exemplarily the duplicate slurries from site III of the February 2008 sampling after an incubation period of 5–15 days. A total of 927 pmoA sequences obtained by 454 pyrosequencing were analyzed by Blastclust, resulting in 67 representative pmoA sequences that were grouped into 15 population-level clusters. There was a clear correspondence between the decline in the frequency of pmoA sequences belonging to USCα-like populations and the increase in the CH4 concentration used to determine the methane oxidation potential (Figure 3). The composition of methanotrophs in the slurries incubated under a headspace of 10 and 100 ppmv CH4 remained similar to that of the original soil, with a slight increase in the relative abundance of Methylocystis spp. However, the slurries incubated under a headspace of 1000 or 10 000 ppmv CH4 were dominated by Methylocystis spp. related to SL-8 (Figure 3; Supplementary Figure 6), in good correspondence with the sudden increase in the CH4 consumption rates after an initial phase of 4 days. None of the pmoA sequences obtained from the methane-incubated slurries were related to type I methanotrophs.
Discussion
In this study, we linked the activity, composition and seasonal dynamics of methanotrophs inhabiting the surface layer of grassland soil. The grassland showed a typical seasonal pattern of atmospheric methane oxidation, primarily governed by soil moisture (Castro et al., 1994; Borken et al., 2000; Kammann et al., 2001b). Soil moisture was negatively correlated with CH4 uptake rates, while temperature showed a positive correlation. However, temperature had no effect on methanotrophic population dynamics. For sites I and III, the R2 values of the correlation between soil moisture and the CH4 fluxes were greater than those of the correlation between soil temperature and the CH4 fluxes, supporting the notion that soil moisture is the main driver of atmospheric methane oxidation, and soil temperature is its covariable. The situation was slightly different for site II, but the overall model for correlation analysis fitted to the data of this site less well than to those of sites I and III (Figure 2; Supplementary Figure 3).
Thus, the net uptake of atmospheric methane into the soil was high during the vegetative growth period when soil moisture was lowered by plant transpiration and minor or zero during winter (Figure 1c). In summer, soil moisture never fell below 20 vol%, which otherwise may have caused a strong decline in atmospheric methane oxidation due to increasing desiccation stress of bacterial cells (Torn and Harte, 1996; Dunfield, 2007). Depending on soil moisture and water table level, slight production of methane may have occurred in subsurface soil during winter time, as exemplified by mean soil-air CH4 profiles located close to, but not in one of the sampling sites (Supplementary Figure 7). High-affinity methane oxidation prevailed in the sampling sites throughout all seasons, as indicated by soil-air CH4 concentrations that were continuously below atmospheric levels, in particular at site III (Figure 1b). In fact, the more active state of the bacteria during winter was not only reflected by the increased changes in methanotroph diversity and abundance, but also by increased initial rates of the methane oxidation potential calculated for slurries under a headspace of 10 and 100 ppmv CH4 (Supplementary Table 2).
All primer combinations and thermal profiles initially tested failed to consistently produce pmoA amplicons from total DNA of the three sampling sites, except when using the assay A189f/A650r (Supplementary Text 2). Sanger sequencing of pmoA clones and 454 pyrosequencing revealed that the three sampling sites were dominated by members of the pmoA clades RA14, JR1 and MHP throughout seasons. In a previous study, Methylocystis spp. were found to represent the dominant methanotroph population in soil of the Research Station Linden (Horz et al., 2002). However, pmoA detection was possible only after the incubation of soil under 5% (v/v) CH4 for 1 week or by re-amplification of first-round PCR products using the primer combination A189f/A682r. Thus, the findings by Horz et al. (2005) are in a good agreement with our observations. Methylocystis spp. dominated the soil slurries incubated under increased methane concentrations (1000 and 10 000 ppmv) or were detectable only at site III using the primer sets A189f/mb661r and A189f/A682r in semi-nested second-round PCR (Figure 3 and Supplementary Figure 6). Taken together, Horz et al. (2005) may not have detected USCα owing to the unavailability of the primer set A189f/A650r at the time of their study, or the dominance of the USCα-like group in this study may be due to a gradual change in the methanotroph composition towards USCα-like populations during the last 8–10 years, induced by a reduced annual input of nitrogen (from about 80–40 kg N ha−1 a−1) since 1996. The latter may have relieved USCα-like populations from inhibitory effects of nitrogen (Bodelier and Laanbroek, 2004; Dunfield, 2007) (see Supplementary Text 3 for additional details).
As compared to RA14, JR1 and MHP have only been detected in a limited number of soils but, like RA14, primarily in those that are shown or assumed to be a continuous or seasonal sink for atmospheric methane: forest, grassland, shrubland, Calluna-covered moorland and deglaciated soils (Horz et al., 2005; Knief and Dunfield, 2005; Knief et al., 2006; Ogram et al., 2006; Singh et al., 2007; Chen et al., 2008; Dörr et al., 2010; Bárcena et al., 2011). The detection of RA14, JR1 and MHP in our sampling sites may suggest that their members have similar ecophysiological roles, in correspondence to their monophyletic origin in pmoA-based trees (this study; Chen et al., 2008). Overall, these observations make it reasonable to combine RA14, JR1 and MHP into USCα-like group or populations, despite the fact that the pmoA sequence divergence between the three clades may be sufficient to consider genus-level distinction (Degelmann et al., 2010; and Supplementary Table 1). Another interesting observation is that particularly pmoA sequences of the MHP clade appear to be under very strong purifying natural selection acting to prevent amino-acid changes in particulate methane monooxygenase (Supplementary Table 1). Biased conclusion due to a limited sequence data set of 38 MHP-like pmoA sequences cannot be fully excluded, but is unlikely because the sequences were obtained from geographically different soils, including moorland (UK; Chen et al., 2008), meadow soil (Germany; this study), watershed soil (Georgia, USA; Ogram et al., 2006) and glacier forefield in Southeast Greenland (Denmark; Bárcena et al., 2011).
T-RFLP fingerprinting is a robust method to assess spatial differences and/or temporal changes in methanotroph diversity (for example, Horz et al., 2005; Singh et al., 2007; Shrestha et al., 2008, 2010; Krause et al., 2010; Ho et al., 2011). We used this method and real-time PCR of pmoA amplicons to monitor site-specific differences and seasonal changes in methanotroph diversity and abundance. Seasonal population dynamics were significantly affected by soil moisture but not by temperature, and most pronounced in site III between August and November 2007. More precisely, population dynamics were increasingly stimulated at soil moisture contents greater than about 50 vol% (compare Figure 1 with Figure 4) and coincided with a decline in net CH4 uptake. They were related to the emergence of new species-level populations within the MHP clade and a significant (three-fold) increase in pmoA gene copy numbers, of up to 1.8 × 106 copies g−1 of dry soil. These population dynamics may even be an underestimate of the real dynamics within the USCα-like group, considering that the USC-α-like diversity cannot be fully distinguished by pmoA-based T-RFLP profiling.
A reverse effect in the pmoA gene copy numbers occurred between February and May 2008. They decreased to 9 × 105 copies g−1 of dry soil, presumably triggered by the sharp decline in soil moisture to 20–30 vol% during the 5 weeks before sampling in May 2008, while net CH4 uptake increased again, likely due to better and deeper CH4 and O2 diffusion into the soil profile. The decrease in pmoA gene copy numbers coincided with a decrease in the relative abundance of the 446-bp T-RF (Figure 4). The methanotrophs defined by the 446-bp T-RF belong to the MHP clade and had developed in response to the increase in soil moisture.
Active MHP populations were first detected in nutrient poor Calluna-covered moorlands in the UK, with more than 40% of all mRNA-derived pmoA clone sequences obtained from this study site (Chen et al., 2008). Moisture content (89.25%), depth of maximum CH4 oxidation (10–15 cm) and CH4 fluxes (3.1 μmol m−2 h−1) were similar to those observed in our study for site III in autumn/winter 2007/2008. Based on CH4 flux data and the observation that the Calluna-covered peat soil could oxidize CH4 down to 20 ppmv, Chen et al. (2008) speculated that MHP methanotrophs may be adapted to low methane concentrations. Members of MHP were also detected in an acidic meadow soil that had a high affinity for CH4 (Km(app)=72 ppmv) (Knief et al., 2006). Thus, extending the ideas of Chen et al. (2008), it appears reasonable to assume that MHP methanotrophs are well adapted to hydromorphic soils that are exposed to strong variations in moisture content and water table levels. They may be active at ambient or, perhaps more likely, slightly elevated methane concentrations when increasing soil moisture begins to support soil methane production (Supplementary Figure 7). An alternative explanation may be that MHP methanotrophs used substrates other than methane for growth activity. Facultative methanotrophy has been suggested as one possible survival strategy of atmospheric methane oxidizers, such as USCα and USCγ (Dunfield, 2007). In fact, facultative particulate methane monooxygenase-based methanotrophy has recently been shown for Methylocapsa aurea and various Methylocystis spp. (Dunfield et al., 2010; Belova et al., 2011; Im et al., 2011), with acetate as the common substrate for growth. Intriguingly, seasonal changes in pmoA gene copy numbers in the wettest site III corresponded well to those of the bacterial 16S rRNA gene. The general stimulation of bacterial activity by increased soil moisture may have made substrates available to MHP methanotrophs other than methane.
In summary, our results provide first evidence that not only the net uptake of atmospheric methane, but also the composition and abundance of USCα-like populations are affected by seasonal changes in climatic variables. The question of whether slightly elevated methane concentrations or substrates other than methane, or a combination of both scenarios, promoted MHP population dynamics during the winter season cannot be resolved here and has to be addressed in future studies.
References
Baani M, Liesack W . (2008). Two isozymes of particulate methane monooxygenase with different methane oxidation kinetics are found in Methylocystis sp. strain SC2. Proc Natl Acad Sci USA 105: 10203–10208.
Bárcena TG, Finster KW, Yde JC . (2011). Spatial patterns of soil development, methane oxidation, and methanotrophic diversity along a receding glacier forefield, Southeast Greenland. Arct Antarct Alp Res 43: 178–188.
Belova SE, Baani M, Suzina NE, Bodelier PLE, Liesack W, Dedysh SN . (2011). Acetate utilization as a survival strategy of peat-inhabiting Methylocystis spp. Environ Microbiol Rep 3: 36–46.
Blunier T, Chappellaz J, Schwander J, Barnola JM, Desperts T, Stauffer B et al. (1993). Atmospheric methane record from a Greenland ice core over the last 1000 years. Geophys Res Lett 20: 2219–2222.
Bodelier PLE, Laanbroek HJ . (2004). Nitrogen as a regulatory factor of methane oxidation in soils and sediments. FEMS Microbiol Ecol 47: 265–277.
Borken W, Brumme R, Xu YJ . (2000). Effects of prolonged soil drought on CH4 oxidation in a temperate spruce forest. J Geophys Res 105: 7079–7088.
Bourne DG, McDonald IR, Murrell JC . (2001). Comparison of pmoA PCR primer sets as tools for investigating methanotroph diversity in three Danish soils. Appl Environ Microbiol 67: 3802–3809.
Bull ID, Parekh NR, Hall GH, Ineson P, Evershed RP . (2000). Detection and classification of atmospheric methane oxidizing bacteria in soil. Nature 405: 175–178.
Castro MS, Peterjohn WT, Melillo JM, Steudler PA, Gholz HL, Lewis D . (1994). Effects of nitrogen fertilization on the fluxes of N2O, CH4, and CO2 from soils in a Florida slash pine plantation. Can J For Res 24: 9–13.
Cavagnaro TR, Jackson LE, Scow TM, Hristova KR . (2007). Effects of arbuscular mycorrhizas on ammonia oxidizing bacteria in an organic farm soil. Microb Ecol 54: 618–626.
Chen Y, Dumont MG, McNamara NP, Chamberlain PM, Bodrossy L, Stralis-Pavese N et al. (2008). Diversity of the active methanotrophic community in acidic peatlands as assessed by mRNA and SIP-PLFA analyses. Environ Microbiol 10: 446–459.
Conrad R . (2009). The global methane cycle: recent advances in understanding the microbial processes involved. Environ Microbiol Rep 1: 285–292.
Degelmann DM, Borken W, Drake HL, Kolb S . (2010). Different atmospheric methane-oxidizing communities in European beech and Norway spruce soils. Appl Environ Microbiol 76: 3228–3235.
Delmont TO, Robe P, Cecillon S, Clark IM, Constancias F, Simonet P et al. (2011). Accessing the soil metagenome for studies of microbial diversity. Appl Environ Microbiol 77: 1315–1324.
Dörr H, Katruff L, Levin I . (1993). Soil texture parameterization of the methane uptake in aerated soils. Chemosphere 26: 697–713.
Dörr N, Glaser B, Kolb S . (2010). Methanotrophic communities in Brazilian ferralsols from naturally forested, afforested, and agricultural sites. Appl Environ Microbiol 76: 1307–1310.
Dunfield PF . (2007). The soil methane sink. In: Reay D, Hewitt N, Smith K, Grace J (eds). Greenhouse Gas Sinks. CABI Publishing: Wallingford, UK, pp 152–170.
Dunfield PF, Belova SE, Vorob’ev AV, Cornish SL, Dedysh SN . (2010). Methylocapsa aurea sp. nov., a facultative methanotroph possessing a particulate methane monooxygenase, and emended description of the genus Methylocapsa. Int J Syst Evol Microbiol 60: 2659–2664.
Eller G, Frenzel P . (2001). Changes in activity and community structure of methane-oxidizing bacteria over the growth period of rice. Appl Environ Microbiol 67: 2395–2403.
Henckel T, Jäckel U, Schnell S, Conrad R . (2000). Molecular analyses of novel methanotrophic communities in forest soil that oxidize atmospheric methane. Appl Environ Microbiol 66: 1801–1808.
Ho A, Lüke C, Frenzel P . (2011). Recovery of methanotrophs from disturbance: population dynamics, evenness and functioning. ISME J 5: 750–758.
Holmes AJ, Roslev P, McDonald IR, Iversen N, Henriksen K, Murrell JC . (1999). Characterization of methanotrophic bacterial populations in soils showing atmospheric methane uptake. Appl Environ Microbiol 65: 3312–3318.
Horz HP, Raghubanshi AS, Heyer E, Kammann C, Conrad R, Dunfield PF . (2002). Activity and community structure of methane-oxidising bacteria in a wet meadow soil. FEMS Microbiol Ecol 41: 247–257.
Horz HP, Rich V, Avrahami S, Bohannan BJM . (2005). Methane-oxidizing bacteria in a California upland grassland soil: diversity and response to simulated global change. Appl Environ Microbiol 71: 2642–2652.
Hutchinson GL, Mosier AR . (1981). Improved soil cover method for field measurement of nitrous oxide fluxes. Soil Sci Soc Am J 45: 311–316.
Im J, Lee S-W, Yoon S, DiSpirito AA, Semrau JD . (2011). Characterization of a novel facultative Methylocystis species capable of growth on methane, acetate and ethanol. Environ Microbiol Rep 3: 174–181.
IPCC (2007). Climate change 2007: the physical science basis. In: Solomon S, Qin D, Manning M, Chen Z, Marquis M, Averyt KB, Tignor M, Miller HL (eds). Contribution of Working Group I to the Fourth Assessment Report of the Intergovernmental Panel on Climate Change. Cambridge University Press: Cambridge, UK and New York, NY, USA.
Jaatinen K, Knief C, Dunfield PF, Yrjälä K, Fritze H . (2004). Methanotrophic bacteria in boreal forest soil after fire. FEMS Microbiol Ecol 50: 195–202.
Jäger HJ, Schmidt SW, Kammann C, Grünhage L, Müller C, Hanewald K . (2003). The University of Giessen free-air carbon dioxide enrichment study: description of the experimental site and of a new enrichment system. J Appl Bot 77: 117–127.
Kammann C, Grünhage L, Jäger H-J . (2001a). A new sampling technique to monitor concentrations of CH4, N2O and CO2 in air at well-defined depths in soils with varied water potential. Eur J Soil Sci 52: 297–303.
Kammann C, Grünhage L, Jäger HJ, Wachinger G . (2001b). Methane fluxes from differentially managed grassland study plots: the important role of CH4 oxidation in grassland with a high potential for CH4 production. Environ Pollut 115: 261–273.
Kammann C, Hepp S, Lenhart K, Müller C . (2009). Stimulation of methane consumption by endogenous CH4 production in aerobic grassland soil. Soil Biol Biochem 41: 622–629.
Kammann C, Müller C, Grünhage L, Jäger HJ . (2008). Elevated CO2 stimulates N2O emissions in permanent grassland. Soil Biol Biochem 40: 2194–2205.
Knief C, Dunfield PF . (2005). Response and adaptation of different methanotrophic bacteria to low methane mixing ratios. Environ Microbiol 7: 1307–1317.
Knief C, Kolb S, Bodelier PLE, Lipski A, Dunfield PF . (2006). The active methanotrophic community in hydromorphic soils changes in response to changing methane concentration. Environ Microbiol 8: 321–333.
Knief C, Lipski A, Dunfield PF . (2003). Diversity and activity of methanotrophic bacteria in different upland soils. Appl Environ Microbiol 69: 6703–6714.
Kolb S . (2009). The quest for atmospheric methane oxidizers in forest soils. Environ Microbiol Rep 1: 336–346.
Krause S, Lüke C, Frenzel P . (2010). Succession of methanotrophs in oxygen–methane counter-gradients of flooded rice paddies. ISME J 4: 1603–1607.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar et al. (2004). ARB: a software environment for sequence data. Nucl Acids Res 32: 1363–1371.
Ogram A, Castro H, Stanley E, Chen W, Prenger J . (2006). Distribution of methanotrophs in managed and highly disturbed watersheds. Ecol Indicat 6: 631–643.
Oksanen J . (2008). Multivariate analysis of ecological communities in R: vegan tutorial. [WWW document]. URL http://cc.oulu.fi/~jarioksa/opetus/metodi/vegantutor.pdf. 2-13-2008. Ref Type: Internet Communication.
Regan K, Kammann C, Hartung K, Lenhart K, Müller C, Philippot L et al. (2011). Can differences in microbial abundances help explain enhanced N2O emissions in a permanent grassland under elevated atmospheric CO2? Global Change Biol 17: 3176–3186.
Shrestha M, Abraham WR, Shrestha PM, Noll M, Conrad R . (2008). Activity and composition of methanotrophic bacterial communities in planted rice soil studied by flux measurements, analyses of pmoA gene and stable isotope probing of phospholipid fatty acids. Environ Microbiol 10: 400–412.
Shrestha M, Shrestha PM, Frenzel P, Conrad R . (2010). Effect of nitrogen fertilization on methane oxidation, abundance, community structure, and gene expression of methanotrophs in the rice rhizosphere. ISME J 4: 1545–1556.
Shrestha PM, Noll M, Liesack W . (2007). Phylogenetic identity, growth-response time and rRNA operon copy number of soil bacteria indicate different stages of community succession. Environ Microbiol 9: 2464–2474.
Singh BK, Tate KR, Kolipaka G, Hedley CB, Macdonald CA, Millard P et al. (2007). Effect of afforestation and reforestation of pastures on the activity and population dynamics of methanotrophic bacteria. Appl Environ Microbiol 73: 5153–5161.
Torn MS, Harte J . (1996). Methane consumption by Montane soils: implications for positive and negative feedback with climatic change. Biogeochemistry 32: 53–67.
Wang JS, Logan JA, McElroy MB, Duncan BN, Megretskaia IA, Yantosca RM . (2004). A 3-D model analysis of the slowdown and interannual variability in the methane growth rate from 1988 to 1997. Global Biogeochem Cycles 18: B3011.
Wuebbles DJ, Hayhoe K . (2002). Atmospheric methane and global change. Earth-Sci Rev 57: 177–210.
Acknowledgements
The study was supported by the European Science Foundation (METHECO, EuroDiversity 018) and Deutsche Forschungsgemeinschaft (LI 455/3-1). Additional support was provided by the LOEWE Center for Synthetic Microbiology (SYNMIKRO). Bomba Dam is a recipient of an Alexander von Humboldt Fellowship. Claudia Kammann gratefully acknowledges a postdoctoral fellowship of the University College Dublin.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Shrestha, P., Kammann, C., Lenhart, K. et al. Linking activity, composition and seasonal dynamics of atmospheric methane oxidizers in a meadow soil. ISME J 6, 1115–1126 (2012). https://doi.org/10.1038/ismej.2011.179
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2011.179
Keywords
This article is cited by
-
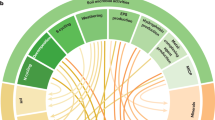
The methane-driven interaction network in terrestrial methane hotspots
Environmental Microbiome (2022)
-
Unique high Arctic methane metabolizing community revealed through in situ 13CH4-DNA-SIP enrichment in concert with genome binning
Scientific Reports (2022)
-
Greenhouse gas (CO2, CH4, and N2O) emissions after abandonment of agriculture
Biology and Fertility of Soils (2022)
-
Aerobic Methanotrophy and Co-occurrence Networks of a Tropical Rainforest and Oil Palm Plantations in Malaysia
Microbial Ecology (2022)
-
Nitrogen addition decreases methane uptake caused by methanotroph and methanogen imbalances in a Moso bamboo forest
Scientific Reports (2021)