Abstract
Dehalococcoides ethenogenes strain 195 (DE195) was grown in a sustainable syntrophic association with Desulfovibrio vulgaris Hildenborough (DVH) as a co-culture, as well as with DVH and the hydrogenotrophic methanogen Methanobacterium congolense (MC) as a tri-culture using lactate as the sole energy and carbon source. In the co- and tri-cultures, maximum dechlorination rates of DE195 were enhanced by approximately three times (11.0±0.01 μmol per day for the co-culture and 10.1±0.3 μmol per day for the tri-culture) compared with DE195 grown alone (3.8±0.1 μmol per day). Cell yield of DE195 was enhanced in the co-culture (9.0±0.5 × 107 cells per μmol Cl− released, compared with 6.8±0.9 × 107 cells per μmol Cl− released for the pure culture), whereas no further enhancement was observed in the tri-culture (7.3±1.8 × 107 cells per μmol Cl− released). The transcriptome of DE195 grown in the co-culture was analyzed using a whole-genome microarray targeting DE195, which detected 102 significantly up- or down-regulated genes compared with DE195 grown in isolation, whereas no significant transcriptomic difference was observed between co- and tri-cultures. Proteomic analysis showed that 120 proteins were differentially expressed in the co-culture compared with DE195 grown in isolation. Physiological, transcriptomic and proteomic results indicate that the robust growth of DE195 in co- and tri-cultures is because of the advantages associated with the capabilities of DVH to ferment lactate to provide H2 and acetate for growth, along with potential benefits from proton translocation, cobalamin-salvaging and amino acid biosynthesis, whereas MC in the tri-culture provided no significant additional benefits beyond those of DVH.
Similar content being viewed by others
Introduction
Dehalococcoides are, thus far, the only known bacteria capable of completely dechlorinating tetrachloroethene (PCE) and trichloroethene (TCE) to the benign product ethene (Freedman and Gossett, 1989; Maymó-Gatell et al., 1997; Holliger et al., 1999; Cupples et al., 2003; He et al., 2003b; Smidt and de Vos, 2004). They exhibit low growth rates, specific obligate nutrient requirements (hydrogen as electron donor, acetate as carbon source, cobalamin as co-factor) and non-robust physiology when grown in isolation, resulting in limited dechlorination activity and subculturing reproducibility (DiStefano et al., 1992; Maymó-Gatell et al., 1997; He et al., 2003a, 2003b). Consequently, developing methods to improve the robust growth and dechlorination activity of Dehalococcoides would be useful for developing improved bioremediation strategies.
Dehalococcoides are commonly found in microbial communities that contain other anaerobes, such as Desulfovibrio, Eubacterium, Acetobacterium, Citrobacter, Spirochetes and Clostridium. (Richardson et al., 2002; Ritalahti and Löffler, 2004; Duhamel and Edwards, 2006; Lee et al., 2006), which are able to ferment organic substrates into hydrogen and acetate. Studies of a previously described dechlorinating microbial community, ANAS, showed that it was dominated by three bacterial groups: Dehalococcoides, Desulfovibrio and Clostridium, with the hydrogenotrophic methanogen Methanobacterium congolense (MC) as the dominant archaeal species (Richardson et al., 2002). Previous studies have shown that Desulfovibrio vulgaris Hildenborough (DVH) can transform lactate to acetate and H2 when grown syntrophically with a H2-utilizing methanogen (McInerney and Bryant, 1981; Schink, 1997). It is possible that DVH might also have a role in syntrophic interactions with other hydrogenotrophic microorganisms such as Dehalococcoides.
Methanogens are commonly found to be growing concomitantly with Dehalococcoides within chloroethene-degrading communities. It has been hypothesized that methanogens have an important role in the anaerobic dechlorination by cometabolic processes (Vogel and McCarty, 1985; Fathepure and Boyd, 1988; Gantzer and Wackett, 1991). In addition, methanogens were found to compete for hydrogen with dehalogenators (Smatlak et al., 1996; Fennell et al., 1997; Yang and McCarty, 1998). Although there have been a number of studies evaluating the link between methanogens and Dehalococcoides (Fathepure and Boyd, 1988; Freedman and Gossett 1989; Smatlak et al., 1996; Fennell et al., 1997; Löffler et al., 1997; Yang and McCarty, 1998; Booker and Pavlostathis, 2000; Heimann et al., 2006), the role of methanogens in dechlorinating communities remains unclear.
We have previously generated defined consortia containing Dehalococcoides ethenogenes strain 195 (DE195) and Desulfovibrio desulfuricans, using lactate as the electron donor (He et al., 2007). In that single-transfer experimental co-culture, PCE was dechlorinated to vinyl chloride (VC) and ethene with concomitant growth of DE195, whose density was 1.5 times greater than when grown in isolation (He et al., 2007).
In this study, we constructed two defined DE195-containing consortia capable of sustained robust growth (>15 subcultures) with lactate as the electron donor and carbon source. These two consortia, a co-culture containing DE195 and DVH, and a tri-culture containing DE195, DVH and MC, dechlorinate TCE to VC and ethene consistently and reproducibly. Physiological characteristics were quantified and transcriptomic and proteomic analyses were applied to evaluate the effects of long-term syntrophic growth on this Dehalococcoides strain.
Materials and methods
Bacterial cultures and growth conditions
DE195 was grown in 160-ml serum bottles containing 100 ml defined medium as described previously (He et al., 2007), with H2:CO2 (20:80 v/v) headspace, 5 mM acetate, 78 μmol TCE and 100 μg l−1 cyanocobalamin (vitamin B12). Co-cultures containing DE195 and DVH (DE195/DVH) were grown in the same medium with the substitutions of 5 mM lactate for acetate and N2:CO2 (90:10 v/v) headspace for H2:CO2. The ratio of DE195 to DVH cells in the initial syntrophic co-culture was 10:1. Three parallel bottles were established as biological triplicates for each subculture. Subcultures were established with transfers of the co-culture (10% v/v) from one of the three bottles into three newly prepared bottles containing 90 ml fresh medium every 10 days for 20 subcultures (∼66 generations) before the described experiment. Another co-culture containing DVH and MC (DVH/MC) was prepared by inoculating 10 ml of each into 80 ml lactate medium. After three subcultures (10% v/v), the defined tri-culture (DE195/DVH/MC) was constructed by adding 10 ml of DVH/MC and 10 ml of DE195/DVH to 80 ml lactate medium in triplicate bottles. The tri-culture was maintained in the same medium as the co-culture, with the exception that TCE was added in ∼40 μmol doses to avoid MC inhibition. A DE195/DVH co-culture with ∼40 μmol TCE was also constructed as a control. Transfers of the co-culture (10% v/v) with low TCE dosage and the tri-culture (10% v/v) from one of the three bottles into three newly prepared bottles containing fresh medium were made every 10 days for 16 subcultures (∼53 generations) before the described experiment.
DNA extraction and cell growth quantification
Cells from 1.5 ml culture sampled from each biological replicate were collected by centrifugation (21 000 × g for 10 min at 4 °C), and genomic DNA was extracted using the DNeasy Blood and Tissue Kit (Qiagen, Valencia, CA, USA). Quantitative PCR (qPCR) using SYBR Green-based detection reagent (Applied Biosystems, Foster City, CA, USA) was applied to quantify 16S rRNA genes. Briefly, each 20-μl reaction mix contained 2.5 μl of genomic DNA sample or 10-fold serially diluted standard, 1 × fast SYBR Green master mix and 0.625 μM of forward and reverse primers (Supplementary Table S1). Genomic DNA of DE195, DVH and MC isolates were quantified using Nanodrop 3300 fluorometer (NanoDrop Technologies, Wilmington, DE, USA) according to the manufacturer's instructions and used as standards for qPCR. Cell density was determined using the equation given by Ritalahti et al. (2006). According to the USDOE IMG website (http://img.jgi.doe.gov/cgi-bin/pub/main.cgi), the genome sizes for DE195 and DVH are 1.5 and 3.8 Mbp, respectively. Because of the lack of genomic information for MC, we assumed the genome size for MC to be 2 Mbp according to the average genome size of sequenced hydrogenotrophic methanogens.
Cell collection for transcriptomic and proteomic analysis
Cells were all collected during exponential phase when ∼90% of TCE was dechlorinated (that is, DE195: day 7; DE195/DVH fed by 78 μmol TCE: day 5; DE195/DVH fed by 40 μmol TCE and DE195/DVH/MC: day 3). To collect sufficient material for transcriptomic microarray analysis, 15 subcultures were inoculated and grown from the triplicate bottles of each culture (that is, pure DE195, DE195/DVH and DE195/DVH/MC). Then, for each biological triplicate, cells from five bottles were collected by vacuum filtration (100 ml culture per filter). Each filter was placed in a 2-ml microcentrifuge tube, frozen with liquid nitrogen and stored at −80 °C until processing. For proteomic analysis, a total of 12 bottles of DE195 and DE195/DVH were inoculated and grown, and cells were collected by centrifugation (12 000 × g for 5 min at 4 °C) in a nitrogen-flushed 250-ml centrifuge bottle. Supernatant was discarded, and cell pellets were frozen at −80 °C and shipped overnight on dry ice to Oak Ridge National Laboratory, where they were pooled before dividing into three analytical replicates for further analysis.
RNA extraction
RNA was extracted using the Qiagen RNeasy Mini Kit (Qiagen) according to the manufacturer's instructions with the addition of a bead-beating step using 1 g 100 μm diameter zirconia-silica beads (Biospec Products, Bartlesville, OK, USA) after addition of the lysis buffer. Contaminating DNA in the RNA samples was removed by DNase I treatment using a DNA-free kit (Applied Biosystems) according to the manufacturer's instructions. RNA was quantified on the Nanodrop 3300 fluorometer using the Quant-iT RiboGreen RNA Assay kit (Invitrogen, Molecular Probes, Carlsbad, CA, USA) according to the manufacturer's instructions.
Transcriptomic microarray
The Affymetrix (Santa Clara, CA, USA) GeneChip microarrays targeting the genome of DE195 were designed and applied as described previously by West et al. (2008) and Johnson et al. (2008). The microarray platform (GPL6336) was deposited to the NCBI Gene Expression Omnibus database (Johnson et al., 2008). cDNA was synthesized from 10 μg RNA, then fragmented, labeled and hybridized to each array. The hybridized arrays were stained, washed and then scanned with an Affymetrix Scan 3000 scanner. All procedures were performed according to the protocol outlined in Section 3 of the Affymetrix GeneChip Expression Analysis Technical Manual (http://www.affymetrix.com). Three replicate arrays were analyzed for each condition.
Microarray data analysis
The microarray analyses were performed using the R statistical program (http://www.r-project.org; R 2005) with packages available from Bioconductor version 1.9 (http://www.bioconductor.org; Gentleman et al., 2004) as described by Johnson et al. (2008, 2009). The Benjamini and Hochberg procedure (Benjamini and Hochberg, 1995) was applied to control the false discovery rate <0.05. In addition, only genes with absolute hybridization signal intensities >200 for at least one condition and with more than twofold changes between two conditions could be considered to be significantly regulated and used for further analyses.
Validation of microarray data
Various transcripts that were observed to be significantly differentially transcribed from microarray results were quantified by reverse transcription-qPCR using the QuantiFast SYBR Green reverse transcription-PCR Kit (Qiagen) according to manufacturer's instructions. The same RNA samples were processed as used in the microarray analysis. Targeted genes and the appropriate primer sets are listed in Supplementary Table S1.
Proteomic analysis
DE195 and DE195/DVH cell samples were analyzed via two-dimensional nanoES 2d LC-MS/MS as described in VerBerkmoes et al. (2009). Briefly proteomes were digested in peptides with sequencing grade trypsin (Promega, Madison, WI, USA) then separated via two-dimensional high-performance liquid chromatography (SCX/RP) into a nanoelectrospray source connected to a Linear ion Trap Orbitrap with full scans at 30 K in Orbitrap and five subsequent data dependent MS/MS scans in the linear ion trap. MS/MS spectra were searched with SEQUEST (Eng et al., 1994) against a database created with both DE195 and DVH predicted protein sequences, common contaminants as well as common lab protein standards. Peptide identifications were filtered and sorted into proteins with DTASelect (Tabb et al., 2002) as described previously (VerBerkmoes et al., 2009). Contrast (Tabb et al., 2002) was used to display all proteins across runs, differentially expressed proteins were identified based on the following criteria (Chourey et al., 2006): at least under one condition, >40% sequence coverage, more than five unique peptides and ⩾2-fold difference in spectral counts identified between DE195/DVH and DE195 isolate. Data identification as well as the actual MS/MS spectra from every peptide and accessory scores are available at http://compbio.ornl.gov/Dehalo195_CoCulture/.
Cobalamin measurements
Lactobacillus delbrueckii subsp. lactis was used in a biological assay for cobalamin as described in the Official Methods of Analysis of AOAC International (Horwitz, 2000). The cells were rinsed three times in tris-EDTA buffer before analysis. A standard curve was generated by adding vitamin B12 to mineral salts medium at concentrations of 0.2, 0.4, 0.6, 0.8, 1 and 2 ng/sample, and adding 5 ml B12 assay medium (Sigma-Aldrich, St Louis, MO, USA) and 5 μl of cell culture (Supplementary Figure S1). The cells were incubated at 34 °C on a shaker for 3 days before quantification by qPCR of 16S rRNA gene using appropriate primers (Supplementary Table S1). Filtered supernatants of cultures were analyzed by adding various amounts to 5 ml of L. delbrueckii B12 assay medium (Supplementary Table S6).
Analytical methods
Chloroethenes and ethene were analyzed using gas chromatograph with a flame ionization detector as described by Lee et al. (2006). Hydrogen was analyzed by gas chromatography with a reductive gas detector (Trace Analytical, Menlo Park, CA, USA). Organic acids were analyzed by Waters (Milford, MA, USA) high-performance liquid chromatography equipped with a UV detector (set at 210 nm) as described previously (He et al., 2007).
Accession number
The microarray data in this study (GSE26815) were deposited in the National Center of Biotechnology Information Gene Expression Omnibus database.
Results
Syntrophic growth of DE195 in co- and tri-cultures
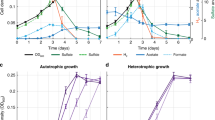
TCE degradation was substantially faster in the sytrophic cultures versus the isolate. That is, while it took 20 days for DE195 to dechlorinate 78 μmol TCE to VC and ethene when grown alone (ca. 3.8±0.1 μmol per day), it took only 7 days (ca. 11.0±0.01 μmol per day) in the co-culture, the DE195/DVH/MC tri-culture took 4 days to dechlorinate 40 μmol TCE (ca. 10.1±0.3 μmol per day), compared with 6 days (ca. 7.9±0.5 μmol per day) in the co-culture control (Figure 1). The amount of ethene produced from VC in the co-culture was higher compared with the isolate, and similar compared with the tri-culture (Figure 1). By the twentieth subculture of the co-culture, the syntrophic growth of DE195 was substantially more consistent and robust than growth in isolation. For example, whereas subculturing 10 vials of DE195 isolate resulted in only six successful cultures, all of the 10 vials inoculated with DE195/DVH subcultures grew successfully. Similar results were observed over multiple subsequent subcultures. The density of DE195 grown in the twentieth subculture of the co-culture was approximately two times higher than when DE195 was grown alone (Figure 2a; two-tailed student's t-test P=0.002). The density of DE195 grown in the sixteenth subculture of the tri-culture was lower than the co-culture control (Figure 2a; two-tailed student's t-test P=0.04). The growth of DVH in the tri-culture was fivefold greater than that in the co-culture control. The ratio of DE195 to DVH cells in co-culture remained ∼5:1 (Figure 2b) over multiple subsequent subcultures, whereas the ratio of DE195 to DVH cells in the tri-culture remained around 1:1.5 (Figure 2b). Further, the ratio of DE195 to MC and DVH to MC remained at ∼4:1 and 6:1, respectively (Figure 2b). The cell yield of DE195 in the co-culture was higher (9.0±0.5 × 107 cells per μmol Cl− released) than that of DE195 in pure culture (6.8±0.9 × 107 cells per μmol Cl− released; two-tailed student's t-test P=0.02), whereas the cell yield of DE195 in the tri-culture (7.3±1.8 × 107 cells per μmol Cl− released) had no significant difference from that of the co-culture control (10±1.6 × 107 cells per μmol Cl− released; two-tailed student's t-test P=0.12).
Temporal changes in the quantities of solvents for (a) DE195 fed ∼78 μmol TCE, (b) DE195/DVH fed ∼78 μmol TCE, (c) DE195/DVH fed ∼40 μmol TCE and (d) DE195/DVH/MC fed ∼40 μmol TCE. All measurements are averages from three biological replicates and error bars are the s.d.; (•) TCE, (▴) c-DCE, (♦) VC, (▪) ethene.
When DE195 was grown in isolation, 500 μmol acetate was added to the basal medium, far exceeding the stoichiometric requirement of 0.03 μmol for growing DE195 to 108 cells per ml and assuming a cell composition of C5H7O2N (Rittmann and McCarty, 2001; Cupples et al., 2003). In the co- and tri-cultures, lactate was provided as the electron donor and carbon source, with the expectation that DVH would ferment it into acetate and hydrogen with the stoichiometry given in Table 1, thus supporting the hydrogen and acetate requirements of DE195 and MC. However, the growth of DVH on lactate is only thermodynamically favorable when sufficient hydrogen is consumed by another strain (DE195 for dechlorination and/or MC for hydrogenotrophic methanogenesis (Table 1)), such that sustained survival of the constructed consortia requires syntrophic association among the species. The 500 μmol lactate provided per bottle was partially consumed in the co-culture but was completely consumed in the tri-culture (Figure 3), with near stoichiometric acetate production in both. The co-culture also generated 140 μmol (Figure 3a) hydrogen indicating that ∼40% of the lactate consumed provided electrons for dechlorination. In the tri-culture, hydrogen was completely consumed by day 6 and 250 μmol of methane was generated (Figure 3b), indicating that ∼90% of the lactate electrons were consumed by methanogenesis whereas only 10% were used for dechlorination. Although MC consumed most of the generated hydrogen in the tri-culture, the aqueous hydrogen concentration never dropped below 5 nM, remaining above the hydrogen threshold (2 nM) for dechlorination by Dehalococcoides (Yang and McCarty, 1998). Consequently, competition for hydrogen between MC and DE195 was not observed to affect dechlorination in this study.
Transcriptomic microarray validation and analysis
Copy numbers of mRNA from 22 genes targeted by the microarray were quantified by reverse transcription-qPCR and compared with signal intensities measured by the microarray (Supplementary Table S3). All genes showed the same direction of transcription regulation, validating the microarray results (Supplementary Table S3).
Transcriptomic microarray analysis of DE195 grown in the co-culture and in isolation identified 102 genes that were differentially transcribed (Figure 4, Supplementary Tables S4, S5). However, no DE195 genes were differentially transcribed between the co- and tri-culture conditions, indicating that the presence of MC in the constructed consortium did not significantly affect gene expression of DE195 (data not shown).
Plot of signal intensities of transcripts from DE195 grown alone versus signal intensities of transcripts from DE195/DVH (grey colored points represent statistically significant differential transcription, average intensity >200, P<0.05, more than twofold difference; genes significantly up-regulated (▴) or down-regulated (▾) in DE195/DVH compared with DE195. All measurements are averages from three biological replicates; left-upper corner: s.e.m. photo of DE195/DVH; right-lower corner: s.e.m. photo of DE195).
Cobalamin-associated genes
Cobalamins, including vitamin B12, are corrinoid-based essential co-factors of reductive dehalogenases (RDases; Smidt and de Vos, 2004). They cannot be synthesized de novo by DE195 (Seshadri et al., 2005) nor by other sequenced Dehalococcoides strains. The cobalamin co-factor salvage and transport genes (DET0650–0652/DET0684–0686), as well as the genes annotated to construct and attach the lower ligand base to cobyric acid (DET0657–0660/DET0691–0694) and its associated riboswitch were significantly down-regulated in the co-culture compared with in isolation (Figure 5c and Supplementary Table S4), suggesting that cobalamin might be produced and released in sufficient quantities by DVH to down-regulate the cobalamin-salvaging genes of DE195. Interestingly, DET0125 and DET0126, two putative cobalamin riboswitches that were shown previously to be down-regulated in DE195 by excess cyanocobalamin (Johnson et al., 2009), were also down-regulated in the co-culture. To test the cobalamin-production hypothesis, we measured the cobalamin concentrations in culture supernatants. The supernatant from the DE195 isolate contained about 50 μg l−1 cobalamin after 10 days, whereas the co-culture supernatant contained 60 μg l−1 cobalamin, which is statistically different from the DE195 isolate (two-tailed student's t-test P<0.01; both with 100 μg l−1 initial vitamin B12 amendment) and DVH grown alone without vitamin B12 amendment generated 10 μg l−1 cobalamin in the supernatant after 7 days of growth (Supplementary Table S6), demonstrating the ability of DVH to produce and release this essential co-factor.
Log2 signal intensity ratio between transcripts of DE195/DVH and DE195. All measurements are averages from three biological replicates and error bars are s.d.s. All x-axis labels are designated for DE195 gene loci (for example, DET0101). Dashed lines indicate the twofold difference in signal intensities. *Indicates genes not actively transcribed (signal intensity <200) in one of the two cultures.
Radical SAM domain superfamily
Proteins within this family have many functions in the biosynthesis of DNA precursors, vitamins and co-factors (Sofia et al., 2001) in biodegradation pathways. Among the 15 genes that are predicted to encode SAM proteins within the genome of DE195, 8 genes were actively transcribed and 4 genes (DET0622, DET1280, DET1368 and DET1629) were down-regulated in the co-culture (Figure 5c) whereas one gene (DET1314) was significantly up-regulated in the co-culture as compared with the DE195 isolate.
Genes associated with membrane-bound oxidoreductase complexes
Predicted oxidoreductase complexes identified in the genome of DE195 include as follows: molybdopterin oxidoreductase, putative formate dehydrogenase, nicotinamide adenine dinucleotide hydride-ubiquinone oxidoreductase (complex I, Nuo), as well as six hydrogenase complexes (that is, Hup, Hym, Hyc, Vhu, Ech and Hyp; Seshadri et al., 2005). In this study, the entire operon for the nicotinamide adenine dinucleotide hydride-ubiquinone oxidoreductase (DET0923–933) was significantly down-regulated in the co-culture compared with the isolate (Figure 5a, Supplementary Table S4). Within the six hydrogenase complexes, the Hym operon (DET0145–148) was observed to be significantly down-regulated, whereas a gene predicted to encode a HymA subunit but not associated with other hydrogenase genes (DET0446) was significantly up-regulated in the co-culture. Other genes within the hydrogenase complexes did not show significant regulation (Figure 5b), or were not actively transcribed in either culture condition (that is, Hyc operon).
RDases
The genome sequence of DE195 revealed 19 potential RDase genes (Seshadri et al., 2005), 5 of which (DET0079, DET0162, DET0180, DET0318 and DET1559), were actively transcribed in the isolate with an additional 4 genes up-regulated in the co-culture (DET0311, DET1171, DET1522 and DET1545). Although the transcript levels of tceA (DET0079) were at the same level between the isolate and co-culture, all the other actively transcribed RDase genes were up-regulated in the co-culture (Figure 5a).
Twin-arginine transport system
These proteins are important for secretion outside the cytoplasmic membrane of folded proteins such as RDases, hydrogenase complexes (for example, Hup) and the formate dehydrogenases (for example, putative formate dehydrogenase). The gene predicted to encode one of these proteins, TatC, a secretion-independent translocase (DET1599) was significantly down-regulated in the co-culture (Supplementary Table S4).
Genes involved in amino acid biosynthesis
Several genes involved in amino acids biosynthesis were significantly differentially transcribed. Genes predicted to encode glutamate synthase and chorismate mutase/prephenate dehydratase (DET0038 and DET0461) were significantly down-regulated in the co-culture (Figure 5c and Supplementary Table S4). DET0461 is a gene involved in the biosynthesis of chorismate, a precursor of all aromatic amino acids (that is, tryptophan, tyrosine and phenylalanine). The other genes within the chorismate operon (DET0462–DET0468), except DET0464 and DET0468 were also down-regulated, but less than twofold difference (Figure 5c). The only amino acid synthesis gene significantly up-regulated in the co-culture was DET1484, predicted to encode indole-3-glycerol phosphate synthase, which is associated with tryptophan synthesis.
Stress-related genes
Genes significantly up-regulated in the co-culture included those predicted to encode glutaredoxin family protein reductase (DET0198, fourfold), ferredoxin-thioredoxin reductase (DET0199, threefold), superoxide dismutase (DET0956, fourfold), antioxidant alkyl hydroperoxide reductase (AhpC; DET1581, threefold) and the α-crystallin heat-shock family protein (DET0954, ninefold; Supplementary Table S5). α-Crystallin-related proteins have chaperone-like properties including the ability to prevent the precipitation of denatured proteins and to increase cellular tolerance to stress. DET1178 and DET1580, predicted to encode MarR and TetR transcriptional regulators, respectively, were significantly up-regulated in the co-culture (Supplementary Table S5), whereas another MarR family protein-encoding gene DET1536 was significantly down-regulated (Supplementary Table S4). MarR mediates response to multiple environmental stresses (Tropel and van der Meer, 2004) and the TetR family has been linked with cell density-sensing regulatory cascades (Ramos et al., 2005). Interestingly, the observed stress response was not accompanied by a reduction in apparent growth rate of DE195 grown in the co-culture.
Genes with unknown functions
The most up-regulated genes in the co-culture, but poorly transcribed in the pure culture were DET0765 (153-fold) and DET1322 (117-fold), whose associated functions have not been well annotated (Supplementary Table S5). Both genes are small but present in all Dehalococcoides species. The gene adjacent to DET0765 (DET0766) is predicted to encode a V-type H(+) translocating pyrophosphatase, suggesting a putative function in the regulation of proton transport. The gene adjacent to DET1322 (DET1323) is predicted to encode a dephospho-CoA kinase, involving in the synthesis of Coenzyme A, which is part of a pathway for acetate assimilation (Seshadri et al., 2005).
Proteomic analysis
A total of 610 and 530 proteins were positively detected among the 1249 and 1251 transcripts of open reading frames detected in the transcriptomes of DE195 and DE195/DVH, respectively. According to the proteome of DE195, genes in COG categories of translation, as well as nucleotide metabolism and transport were mostly expressed (>80%). Among the most abundant proteins are chaperonin GroEL (encoded by DET1428), formate dehydrogenase α-subunit (encoded by DET0187), TceA, as well as a cobalamin uptake and salvaging protein CobT (encoded by DET0657/0691). In the co-culture, 86 proteins were determined to be significantly up-regulated, whereas 34 proteins were significantly down-regulated (Supplementary Table S2) compared with the DE195 isolate. Overall, 24 up-regulated proteins were ribosomal proteins, indicating the robust growth of DE195 in the co-culture. Four hydrogenases, VhuA, HymB, HymC and Ech encoded by DET0615, DET0729, DET0730 and DET0866, respectively, were also found to be significantly up-regulated in the co-culture (Supplementary Table S2), whereas no significant differential transcription of these genes was found. Nevertheless, although transcripts for genes (DET0145–0148) predicted to encode HymA, HymB, HymC and HymD hydrogenases were found to be significantly down-regulated in the co-culture, no significant difference was found in the expression of corresponding proteins. Interestingly, cobalamin uptake and salvaging proteins CobT and CobU (encoded by DET0657/0691 and DET660/0694) were found to be less abundant in the co-culture than in the isolate (Supplementary Table S2), consistent with the transcriptomic data. Among the 19 RDases, only TceA and the RDase encoded by DET1559 were detected in both cultures, neither of which exhibited significant regulation. In contrast, transcriptomic data showed no significant regulation of tceA, but a more than twofold up-regulation of DET1559 and another highly transcribed RDase gene (DET1545) in the co-culture (Figure 5a, Supplementary Table S5).
Discussion
In this study, DE195 was sustainably grown with DVH in a syntrophic association as a co-culture, as well as with DVH and the hydrogenotrophic methanogen MC as a tri-culture using lactate as the sole energy and carbon source with TCE as the electron acceptor. In these syntrophic associations, DVH ferments lactate to acetate and hydrogen that becomes available for use by DE195 and MC as carbon source and electron donor, respectively. This syntrophy occurs at trace sulfate concentrations. This approach of defined syntrophic growth enabled direct experimental measurement of the specific effects of associated bacteria on the growth, activity and gene expression of Dehalococcoides, similar to the approach taken for the study of Syntrophomonas wolfei with Methanospirillum hungatei (Beaty and McInerney, 1989), for the study of D. vulgaris with methanogens (Bryant et al., 1977; Scholten et al., 2007; Stolyar et al., 2007; Walker et al., 2009) and for the study of Desulfovibrio sp. strain SULF1 and Desulfovibrio fructosivorans with a dehalorespiring bacterium Desulfitobacterium frappieri TCE1 (Drzyzga et al., 2001; Drzyzga and Gottschal, 2002).
In DE195/DVH and DE195/DVH/MC, the dechlorination rates of DE195 were enhanced 2- to 3-fold over the isolate and cell yields were enhanced in the co-culture by ∼1.5 times. In addition, the ratio of ethene to VC was higher in the co- and tri-cultures and the subculturing was more reproducibly successful, four phenomena that are commonly observed when Dehalococcoides strains are grown in microbial communities (Maymó-Gatell et al., 1997; He et al., 2007). The complete dechlorination but lower cell yield in the tri-culture compared with the co-culture indicated that DE195 uncoupled TCE dechlorination from net cell growth. Johnson et al. (2008) demonstrated that DE195 uncouples dechlorination from net cell growth associated with stress and transition into a stationary phase. The uncoupling in this study may be associated with the competition for H2 in the tri-culture caused by MC, although competition for hydrogen between MC and DE195 was not observed to adversely affect the dechlorination rate in this study. Therefore, the presence of the methanogen did not provide benefits beyond those of DVH. Although the presence of methanogens has been shown to promote reductive dechlorination in some communities (Vogel and McCarty, 1985, Heimann et al., 2006), such an enhancement was not observed for Dehalococcoides in this study. Further, competition for hydrogen between MC and DE195 was not observed to adversely affect dechlorination in this study. The increase in cell density of DE195 grown in co-culture is similar to the increase previously reported for the single-transfer co-culture containing Desulfovibrio desulfuricans strain Essex 6 and 195 (He et al., 2007). A previous study with a co-culture containing Desulfovibrio sp. strain SULF1 and a dehalorespiring bacterium D. frappieri TCE1 showed that at low sulfate concentrations and relatively high PCE concentrations, SULF1 was outnumbered by TCE1 with a protein level ratio about 1:10 (Drzyzga et al., 2001). Because SULF1 and TCE1 are similar in biovolume, the cell number ratio of SULF1 to TCE1 should also be about 1:10, which is similar to the results in this study. Interactions among organisms in anaerobic environments where trophic hierarchies occur with functionally different members of the community providing substrates and essential co-factors for each other and removing inhibitory metabolic products are well known in both the environment and in engineered systems such as anaerobic digesters (Schink, 1997; Rittmann and McCarty, 2001).
One interesting observation that may be related to the increase in dechlorination activity and robustness of DE195 grown in the co- and tri-culture is the increased concentration of the corrinoid co-factor in the cultures containing DVH. Although the vitamin B12 concentration detected in the co-culture was only 10 μg l−1 higher than that of the isolate (60 μg l−1 for the co-culture versus 50 μg l−1 for the isolate), the amount of vitamin B12 required by Dehalococcoides might be uptaken and utilized immediately, such that what was detected in the supernatant is the B12 concentration at steady-state. It is not specifically known what form of corrinoid DE195 prefers. DVH possesses the full set of genes required for the biosynthesis of adenosylcobalamin, a derivative of vitamin B12 (Rodionov et al., 2004), however, mechanism for the transport of adenosylcobalamin has not been identified. Because of the important role of cobalamin as a co-factor for the RDases, the uptake and potential transformation of this corrinoid needs to be elucidated. Both transcriptomic and proteomic analyses showed that cobalamin-associated genes, including genes predicted to encode a corrinoid ABC-type transport system (DET0650–0655/DET0684–0689) and a corrinoid salvage system (DET0657–0660/DET0691–0694), were significantly down-regulated in the co-culture, which suggests that DE195 may exert less energy-salvaging corrinoids when DVH is present. DET0650–0654 were also found to be significantly down-regulated when DE195 is grown with excess vitamin B12, as well as with spent medium of a Dehalococcoides-containing enrichment culture (ANAS) compared with DE195 grown with limited vitamin B12 (Johnson et al., 2009). Interestingly, another significantly down-regulated gene in both this study and the study by Johnson et al. (2009) was DET0126, which is annotated as an anthranilate phosphoribosyltransferase (TrpD) and reported to have an upstream putative cobalamin-binding riboswitch (Johnson et al., 2009). Although the function of DET0126 remains unclear, its differential regulation within different growth conditions suggests that it is of biological importance to DE195.
Because the radical SAM proteins have functions in the biosynthesis of vitamins and co-factors, the down-regulation of several SAM protein-encoding genes could also indicate an increase in the availability of the cobalamin co-factor for DE195 grown with DVH.
Another possible reason for more robust growth and faster dechlorination of DE195 grown in the co- and tri-cultures is facilitated hydrogen transfer between DVH and DE195 because of physical proximity under the unshaken conditions. Membrane-bound oxidoreductase operons, such as putative formate dehydrogenase, nicotinamide adenine dinucleotide hydride-ubiquinone oxidoreductase, as well as Hym and Hup operon were downregulated in DE195 grown in the co-culture compared with the DE195 isolate. These operons are predicted to be components of the electron transport chain, and are important in hydrogen transfer for DE195 (Seshadri et al., 2005). Moreover, the downregulation of TatC gene, whose product functions in the secretion of folded proteins such as the putative hydrogenase complexes and putative formate dehydrogenase, could be a corresponding effect of a down-regulation of those proteins. The down-regulation of these genes suggests that the syntrophic growth of DE195 and DVH might have a different hydrogen transfer system than the DE195 isolate. Transcriptomic analysis of DVH syntrophically grown with a hydrogenotrophic methanogen compared with DVH grown in sulfate-limited monocultures showed that genes encoding hydrogenases Coo, Hyd and Hyn were among the most highly expressed and up-regulated genes (Walker et al., 2009). Therefore, differential regulation of genes encoding hydrogenases of DVH grown in DE195/DVH compared with the DVH isolate should be further investigated, to elucidate the interspecies hydrogen transfer during syntrophic growth.
Finally, aromatic amino acids (that is, tryptophan, tyrosine and phenylalanine) might have important roles in the robust syntrophic growth of DE195 with DVH. In the co-culture, the down-regulation of genes involved in the biosynthesis of chorismate, a precursor for aromatic amino acid biosynthesis, suggests that DVH might support the growth of DE195 by decreasing the need for de novo aromatic amino acid biosynthesis, consistent with our recent detailed amino acid study (Yi et al., 2010).
In summary, D. ethenogenes strain 195 exhibits faster dechlorination and more robust growth when growing syntrophically with DVH using lactate as the carbon and energy source than when grown in isolation on the gaseous substrate hydrogen. The difference in gene transcription and protein expression levels between DE195 grown in isolation and with DVH suggests that effective transfer of hydrogen, cobalamin and some amino acids may contribute to the enhanced dechlorination and robust growth of DE195 in the co-culture. These results provide an improved method for rapid and robust growth of Dehalococcoides strains using lactate as an inexpensive, widely available substrate.
References
Beaty PS, McInerney MJ . (1989). Effects of organic-acid anions on the growth and metabolism of Syntrophomonas wolfei in pure culture and in defined consortia. Appl Environ Microbiol 55: 977–983.
Benjamini Y, Hochberg Y . (1995). Controlling the false discovery rate: a practical and powerful approach to multiple testing. JR Stat Soc Ser B 57: 289–300.
Booker RS, Pavlostathis SG . (2000). Microbial redulctive dechlorination of hexachloro-1, 3-butadiene in a methanogenic enrichment culture. Water Res 34: 4437–4445.
Bryant MP, Campbell LL, Reddy CA, Crabill MR . (1977). Growth of Desulfovibrio in lactate or ethanol media low in sulfate in association with H2-utilizing methanogenic bacteria. Appl Environ Microbiol 33: 1162–1169.
Chourey K, Thompson MR, Morrel-Falvey J, VerBerkmoes NC, Brown SD, Shah M et al. (2006). Global molecular and morphological effects of 24-hour chromium(VI) exposure on Shewanella oneidensis MR-1. Appl Environ Microbiol 72: 6331–6344.
Cupples AM, Sporeman AM, McCarty PL . (2003). Growth of a Dehalococcoides-like microorganism on vinyl chloride and cis-dichloroethene as electron acceptors as determined by competitive PCR. Appl Environ Microbiol 69: 953–959.
DiStefano TD, Gossett JM, Zinder SH . (1992). Hydrogen as an electron donor for dechlorination of tetrachloroethene by an anaerobic mixed culture. Appl Environ Microbiol 58: 3622–3629.
Drzyzga O, Gottschal JC . (2002). Tetrachloroethene dehalorespiration and growth of Desulfitobactetium frappieri TCE1 in strict dependence on the activity of Desulfovibrio fructosivorans. Appl Environ Microbiol 68: 642–649.
Drzyzga O, Gerritse J, Dijk JA, Elissen H, Gottschal JC . (2001). Coexistence of a sulphate-reducing Desulfovibrio species and the dehalorespiring Deulfitobacterium frappieri TCE1 in defined chemostat cultures grown with various combinations of sulphate and tetrachloroethene. Environ Microbiol 3: 92–99.
Duhamel M, Edwards EA . (2006). Microbial composition of chlorinated ethane-degrading cultures dominated by Dehalococcoides. FEMS Microbiol Ecol 58: 538–549.
Eng JK, McCormack AL, Yates III JR . (1994). An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J Am Mass Spectrom 5: 976–989.
Fathepure BZ, Boyd SA . (1988). Reductive dechlorination of perchloroethylene and the role of methanogens. FEMS Microbiol Lett 49: 149–156.
Fennell DE, Gossett JM, Zinder SH . (1997). Comparison of butyric acid, ethanol, lactic acid, and propionic acid as hydrogen donors for the reductive dechlorination of tetrachloroethene. Environ Sci Technol 31: 918–926.
Freedman DL, Gossett JM . (1989). Biological reductive dechlorination of tetrachloroethylene and trichloroethylene to ethylene under methanogenic conditions. Appl Environ Microbiol 55: 2144–2151.
Gantzer CJ, Wackett LP . (1991). Reductive dechlorination catalyzed by bacterial transition-metal coenzymes. Environ Sci Technol 25: 715–722.
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S et al. (2004). Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5: R80.
He J, Holmes V, Lee PKH, Alvarez-Cohen L . (2007). Influence of vitamin B12 and co-cultures on the growth of Dehalococcoides isolates in defined medium. Appl Environ Microbiol 73: 2847–2853.
He J, Ritalahti KM, Aiello MR, Löffler FE . (2003a). Complete detoxification of vinyl chloride (VC) by an anaerobic enrichment culture and identification of the reductively dechlorinating population as a Dehalococcoides species. Appl Environ Microbiol 69: 996–1003.
He J, Ritalahti KM, Yang KL, Koenigsberg SS, Löffler FE . (2003b). Detoxification of vinyl chloride to ethene coupled to growth of an anaerobic bacterium. Nature 424: 62–65.
Heimann AC, Batstone DJ, Jakobsen R . (2006). Methanosarcina spp. drive vinyl chloride dechlorination via interspecies hydrogen transfer. Appl Environ Microbiol 72: 2942–2949.
Holliger C, Wohlfarth G, Diekert G . (1999). Reductive dechlorination in the energy metabolism of anaerobic bacteria. FEMS Microbiol Rev 22: 383–398.
Horwitz W (ed.) (2000). Official Methods of Analysis of AOAC International, 17th edn. AOAC International: Gaithersburg, MD.
Johnson DR, Brodie EL, Hubbard AE, Andersen GL, Zinder SH, Alvarez-Cohen L . (2008). Temporal transcriptomic microarray analysis of Dehalococcoides ethenogenes strain 195 during the transition from exponential growth to the stationary phase. Appl Environ Microbiol 74: 2864–2872.
Johnson DR, Nemir A, Anderson GL, Zinder SH, Alvarez-Cohen L . (2009). Transcriptomic microarray analysis of corrinoid responsive genes in Dehalococcoides ethenogenes strain 195. FEMS Microbiol Lett 294: 198–206.
Lee PKH, Johnson DR, Holmes VF, He J, Alvarez-Cohen L . (2006). Reductive dehalogenase gene expression as a biomarker for physiological activity of Dehalococcoides spp. Appl Environ Microbiol 72: 6161–6168.
Löffler FE, Ritalahti KM, Tiedje JM . (1997). Dechlorination of chloroethenes is inhibited by 2-bromoethanesulfonate in the absence of methanogens. Appl Environ Microbiol 63: 4982–4985.
Maymó-Gatell X, Chien YT, Gossett JM, Zinder SH . (1997). Isolation of a bacterium that reductively dechlorinates tetrachloroethene to ethene. Science 276: 1568–1571.
McINerney MJ, Bryant MP . (1981). Anaerobic degredation of lactate by syntrophic associations of Methanosarcina barkeri and Desulfovibrio species and effect of H2 on acetate degredation. Appl Environ Microbiol 41: 346–354.
Ramos JL, Martínez-Bueno M, Molina-Henares AJ, Terán W, Watanabe K, Zhang X et al. (2005). The TetR family of transcriptional repressors. Microbiol Mol Biol Rev 69: 326–356.
Richardson RE, Bhupathiraju VK, Song DL, Goulet TA, Alvarez-Cohen L . (2002). Phylogenetic characterization of microbial communities that reductively dechlorinate TCE based upon a combination of molecular techniques. Environ Sci Technol 36: 2652–2662.
Ritalahti KM, Amos BK, Sung Y, Wu Q, Koenigsberg SS, Löffler FE . (2006). Quantitative PCR targeting 16S rRNA and reductive dehalogenase genes simultaneously monitors multiple Dehalococcoides strains. Appl Environ Microbiol 72: 2765–2774.
Ritalahti KM, Löffler FE . (2004). Populations implicated in the anaerobic reductive dechlorination of 1, 2-dichloropropane in highly enriched bacterial communities. Appl Environ Microbiol 70: 4088–4095.
Rittmann BE, McCarty PL . (2001). Environmental Biotechnology: Principles and Applications. McGraw-Hill: Boston, pp 570–629.
Rodionov DA, Dubchak I, Arkin A, Alm E, Gelfand MS . (2004). Reconstruction of regulatory and metabolic pathways in metal-reducing ä-proteobacteria. Genome Biol 5: R90.
Schink B . (1997). Energetics of syntrophic cooperation in methanogenic degradation. Microbiol Mol Biol Rev 61: 262–280.
Scholten JC, Culley DE, Brockman FJ, Wu G, Zhang WW . (2007). Evolution of the syntrophic interaction between Desulfovibrio vulgaris and Methanosarcina barkeri: involvement of an ancient horizontal gene transfer. Biochem Biophys Res Commun 352: 48–54.
Seshadri R, Adrian L, Fouts DE, Eisen JA, Phillippy AM, Methe BA et al. (2005). Genome sequence of the PCE-dechlorinating bacterium Dehalococcoides ethenogenes. Science 307: 105–108.
Smatlak CR, Gossett JM, Zinder SH . (1996). Comparative kinetics of hydrogen utilization for reductive dechlorination of tetrachloroethene and methanogenesis in an anaerobic enrichment culture. Environ Sci Technol 30: 2850–2858.
Smidt H, de Vos WM . (2004). Anaerobic microbial dehalogenation. Annu Rev Microbiol 58: 43–73.
Sofia HJ, Chen G, Hetzler BG, Reyes-Spindola JF, Miller NE . (2001). Radical SAM, a novel protein superfamily linking unresolved steps in familiar biosynthetic pathways with radical mechanisms: functional characterization using new analysis and information visualization methods. Nucleic Acids Res 29: 1097–1106.
Stolyar S, Van Dien S, Hillesland KL, Lie TJ, Leigh JA, Stahl DA . (2007). Metabolic modeling of a mutualistic microbial community. Mol Syst Biol 3: 92.
Tabb DL, McDonald WH, Yates III JR . (2002). DTASelect and contrast: tools for assembling and comparing protein identifications from shotgun proteomics. J Proteome Res 1: 21–26.
Tropel D, van der Meer JR . (2004). Bacterial transcriptional regulators for degradation pathways of aromatic compounds. Microbiol Mol Biol Rev 68: 474–500.
Verberkmoes NC, Russell AL, Shah M, Godzik A, Rosenquist M, Halfvarson J et al. (2009). Shotgun metaproteomics of the human distal gut microbiota. ISME J 3: 179–189.
Vogel TM, McCarty PL . (1985). Biotransformation of tetrachloroethylene to trichloroethylene, dichloroethylene, vinyl-chloride, and carbon-dioxide under methanogenic conditions. Appl Environ Microbiol 49: 1080–1083.
Walker CB, He ZL, Yang ZK, Ringbauer JA, He Q, Zhou JH et al. (2009). The electron transfer system of syntrophically grown Desulfovibrio vulgaris. J Bacteriol 191: 5793–5801.
West KA, Johnson DR, Hu P, DeSantis TZ, Brodie EL, Lee PKH et al. (2008). Comparative genomics of Dehalococcoides ethenogenes 195 and a Dehalococcoides-containing enrichment culture. Appl Environ Microbiol 74: 3533–3540.
Yang YR, McCarty PL . (1998). Competition for hydrogen within a chlorinated solvent dehalogenating anaerobic mixed culture. Environ Sci Technol 32: 3591–3597.
Yi S, Zhuang WQ, Feng X, Zinder SH, Tang YJ, Alvarez-Cohen L . (2010). Exogenous amino acid utilization by Dehalococcoides ethenogenes strain 195. 110th General Meeting of the American Society for Microbiology, San Diego, CA.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Men, Y., Feil, H., VerBerkmoes, N. et al. Sustainable syntrophic growth of Dehalococcoides ethenogenes strain 195 with Desulfovibrio vulgaris Hildenborough and Methanobacterium congolense: global transcriptomic and proteomic analyses. ISME J 6, 410–421 (2012). https://doi.org/10.1038/ismej.2011.111
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2011.111
Keywords
This article is cited by
-
Effect of wine pomace extract on dechlorination of chloroethenes in soil suspension
Bioresources and Bioprocessing (2023)
-
Substantial defluorination of polychlorofluorocarboxylic acids triggered by anaerobic microbial hydrolytic dechlorination
Nature Water (2023)
-
Key factors controlling microbial distribution on a DNAPL source area
Environmental Science and Pollution Research (2022)
-
Substrate-dependent competition and cooperation relationships between Geobacter and Dehalococcoides for their organohalide respiration
ISME Communications (2021)
-
Reconstruction and evaluation of oil-degrading consortia isolated from sediments of hydrothermal vents in the South Mid-Atlantic Ridge
Scientific Reports (2021)