Abstract
Purpose:
Phelan–McDermid syndrome is a developmental disability syndrome with varying deletions of 22q13 and varying clinical severity. We tested the hypothesis that, in addition to loss of the telomeric gene SHANK3, specific genomic regions within 22q13 are associated with important clinical features.
Methods:
We used a customized oligo array comparative genomic hybridization of 22q12.3-terminus to obtain deletion breakpoints in a cohort of 70 patients with terminal 22q13 deletions. We used association and receiver operating characteristic statistical methods in a novel manner and also incorporated protein interaction networks to identify 22q13 genomic locations and genes associated with clinical features.
Results:
Specific genomic regions and candidate genes within 22q13.2q13.32 were associated with severity of speech/language delay, neonatal hypotonia, delayed age at walking, hair-pulling behaviors, male genital anomalies, dysplastic toenails, large/fleshy hands, macrocephaly, short and tall stature, facial asymmetry, and atypical reflexes. We also found regions suggestive of a negative association with autism spectrum disorders.
Conclusion:
This work advances the field of research beyond the observation of a correlation between deletion size and phenotype and identifies candidate 22q13 loci, and in some cases specific genes, associated with singular clinical features observed in Phelan–McDermid syndrome. Our statistical approach may be useful in genotype–phenotype analyses for other microdeletion or microduplication syndromes.
Genet Med 2014:16(4):318–328.
Similar content being viewed by others
Introduction
Phelan–McDermid syndrome (PMS, MIM 606232) is a rare condition associated with deletions of chromosome 22q13. Common features of the syndrome include developmental delay, absent or impaired speech, neonatal hypotonia, autistic-like behaviors, and mild dysmorphic features.1,2 Loss of one copy of SHANK3 (SH3 and multiple ankyrin repeat domains 3), a gene in the telomeric portion of 22q13.33, is likely responsible for some of the neurological features of the PMS phenotype.3,4,5 An additional candidate gene is IB2 (islet-brain 2), also known as MAPK8IP2 (mitogen-activated protein kinase 8-interacting protein 2). IB2 maps 70 kb proximal to SHANK3, is deleted in most PMS patients, and may play an important role in synaptic stability and neuronal transmission.6
Understanding the etiology of PMS is complicated by the dual observations that some individuals with small deletions are severely affected3,5,7 and yet when assessing individuals across the full spectrum of PMS, those with larger deletions tend to be more severely affected than those with smaller deletions.4,8,9 This study explores the contribution of genomic regions proximal to SHANK3 as contributing to the phenotypic variability in the syndrome. A previous study conducted on the present cohort identified clinical features associated with deletion size.10 This work addresses the hypothesis that specific genomic regions of chromosome 22q13 are associated with PMS-related clinical features.
Materials and Methods
Patients
The cohort included 70 individuals, 41 females and 29 males, with deletion sizes ranging from 0.2 to 9.2 Mb and a median deletion size of 5.5 Mb. Ages spanned from 5 months to 40 years, with a median age of 5.3 years and a mean age of 7.8 years. Most subjects attended one or more PMS family support conferences held between 2001 and 2008. Blood samples were collected at these meetings or were collected by personal physicians and sent to the investigators. Information on physical features was obtained from standardized physical examinations performed at the conferences by trained clinical geneticists (n = 54) or medical record review (n = 3). Information on developmental milestones, medical history, and behavior was obtained from parent-completed medical history questionnaires (n = 60), most of which were completed with the assistance of the investigators. Level of developmental delay was analyzed as a rank score of 1–7 based on the parental classification of the child’s delay from “mild” to “profound.” Speech, autism spectrum disorders (ASDs), and age when learning to walk were analyzed for those older than 3 years of age. The data were analyzed as “ever” having a given condition compared to “never” having the condition. In cases where a patient participated multiple times over the years, and there were any discrepancies in responses, the positive history was used in analysis. This study was approved by the Institutional Review Board of Self Regional Healthcare (Greenwood, SC), and all participants’ parents or guardians provided informed consent. We excluded from analysis the individuals with chromosomal anomalies other than simple terminal deletions on 22q13 ascertained by our arrays, previous cytogenetic test results if available, or parent report (38 out of 108 individuals assessed). Genome-wide array comparative genomic hybridization (CGH) was not performed in this study. All participants have a terminal deletion encompassing the SHANK3 gene and all but two are also missing one copy of IB2.
Genetic analysis
Genetic deletions were measured from specimens of whole blood using a custom 4 × 44K 60-mer oligo array designed to cover 22q12.3-terminus by Oxford Gene Technology (Oxford, UK). Array CGH genomic coordinates of breakpoints were established according to the 2006 human genome build (NCBI 36/HG 18) with terminal deletion breakpoints ranging from chromosomal position 40.1 to 49.5 Mb. The terminus of chromosome 22 is located at position 49.69 Mb.
Statistical analysis
Statistical analyses using SAS 9.2 (http://www.SAS.com) were used to examine 22 clinical features identified from previous work10 as having different median deletion sizes (P < 0.10) between those with and without the given phenotype ( Tables 1 and 2 ). As a comparison, two phenotypes that were previously found to be unassociated with deletion size were also examined: microcephaly (P = 0.9556) and seizures (considered as parent reporting the child had at least one episode of seizure and also used an antiseizure medication, P = 0.5366).
Three exploratory and complementary methods were used to identify the genomic regions most associated with clinical features. The first two methods sought to identify the genomic breakpoint most associated with any given phenotype. The first method is called the “minimum P value” method.11,12 Sequentially at each breakpoint, the proportion of individuals with a given phenotype was compared between those with a given deletion size or larger to those with a smaller deletion size. This analysis was performed using an exact logistic regression model to adjust for age and gender covariates or by Fisher’s exact two-sided P value when the the sample size was too small to model. The genomic region bounded by the most distal and most proximal breakpoints which had a nominal P value <0.05 was identified as a genomic region of potential association. Unadjusted relative risk (RR) and 95% CIs were calculated at the minimum P-value cutpoint. Bonferroni-adjusted P values were also calculated to adjust for the fact that n−1 statistical tests were calculated for each phenotype.
Second, receiver operating characteristic (ROC) methods13,14,15 were used to examine sensitivity and specificity for all possible breakpoints using a logistic regression model. The Youden index,13 J = (sensitivity + specificity − 1), was calculated at each breakpoint. The breakpoint where the maximum Youden index was achieved was identified as the optimal cutpoint. The area under the curve (AUC) was calculated to determine whether genomic breakpoint position explained the data more than chance (AUC > 0.5). While an AUC value of 1.0 would indicate a perfect predictor, AUC values above 0.9 are considered to be highly accurate and AUC values from 0.7 to 0.9 can be considered moderately accurate.14,15 In addition, as part of this method, age and gender variables were included in the ROC analysis, and an AUC including these additional predictors was also calculated for each feature.
Third, the smallest genomic region of common deletion for each phenotype was also identified (the traditional approach). These three approaches were used simultaneously to narrow the search for potential genes most associated with a given phenotype.
Finally, we compared the prevalence of various conditions among the four cytogenetic bands in our region of interest (22q13.2, 22q13.31, 22q13.32, and 22q13.33).
Protein interaction networks to annotate 22q13 genes
To further identify 22q13 genes of interest in the regions highlighted by the association analysis, known genes related to ASDs,16,17,18,19 intellectual disability (ID),17,18,20,21 hypotonia, and head size were used as seeds in a protein interaction network to identify interacting partners located in the 22q13 deletion region under study (Supplementary Table S1 online). The OMIM database (www.omim.org) was searched for the terms “hypotonia”, “macrocephaly”, and “microcephaly” to identify seed genes for these phenotypes. The gene lists were submitted for each phenotype separately to the online gene interaction tool GeneMANIA22 to search against known protein–protein interaction databases. Genes identified as physical interacting partners with the seed genes were then compared with the list of protein coding genes on 22q13.
Results
Specific genomic regions were associated with each phenotype assessed ( Tables 1 and 2 , Figures 1 , 2 , 3 , Supplementary Figure S1 online). The location of minimum P value and maximum Youden index were almost always identical, and the location of the maximum Youden index was almost always within the range of significant P values obtained from the association analysis. In general, inclusion of age and gender in the logistic models had little impact on the association analysis or the predictive ability of the ROC analysis ( Table 1 ). Using the smallest common deletion as an indication of optimal cutpoint identified the same genomic regions as the association analysis for 7 of 22 features (macrocephaly, facial asymmetry, tall stature, full brow, strabismus, male genital anomalies, and delayed age at walking). In these cases, the RR was undefined (no cases observed below the optimal breakpoint). For the remaining 15 features, a genomic location was found to be associated with either increased risk of the feature (RRs ranging from 1.5 to 8.5) or decreased risk (RRs ranging from 0.3 to 0.4). For 13 features, deletion size was at least moderately predictive (AUC ≥ 0.7). The genomic regions significantly associated with these features are presented graphically in Figure 3 .
Association and receiver operating characteristic (ROC) analysis for speech delay. The top left panel shows the distribution of chromosome 22q13 breakpoints by genomic position for those with sentences and those with absent speech. The top right panel shows the minus log10 P value of association between genomic breakpoint position and speech. The lower line represents a P value <0.05; the upper line is significant after Bonferroni correction (P < 0.0014). The bottom left panel shows the Youden index by breakpoint position. The bottom right panel shows the ROC curve with the area under the curve (AUC).
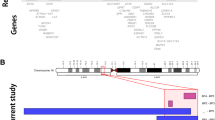
The Phelan–McDermid syndrome deletion region illustrating the genomic regions statistically associated with features of interest. The thin green bands represent regions associated at P < 0.05. The thick green bands represent the regions within 100 kb of the smallest P value and the maximum Youden index. Also shown are locations of known genes and miRNAs, locations of genes found to be associated with autism spectrum disorder (ASD), intellectual disability (ID), hypotonia, or macrocephaly in protein interaction databases using GeneMANIA.22 The interstitial deletions as given by Wilson et al.26 are also provided. FISH, fluorescence in situ hybridization.
For most phenotypes, the patients with the smallest terminal deletions (22q13.33) were less severely affected than those with the largest terminal deletions (22q13.2) ( Table 2 ).
Speech/language delay and developmental delay
The 3.6-Mb genomic region surrounding genomic position 43.9 Mb was associated with speech ability ( Table 1 , Figures 1 and 2 ). While all individuals presented with speech delay, there were differences in verbal communication abilities. Of the 50 individuals over the age of 3 years with information about speech development, 24 had absent speech (0 words), 14 had minimal speech (spoke 1–39 words, but no known sentences or phrases), and 11 had “sentences” (spoke 40 or more words or spoke in phrases or sentences). The subjects in the minimal speech group were not included in the association analysis to reduce misclassification and to better differentiate speech abilities between absent speech and verbal communication ability. Subjects with absent speech had deletion breakpoints ranging from position 40.4 to 49.4 Mb with a median deletion size of 7.0 Mb. Subjects with “sentences” had deletion breakpoints ranging from position 44.4 to 49.5 Mb, with a median deletion size of 3.3 Mb. The distribution of breakpoints is illustrated in Figures 1 and 2 . As shown in Table 1 and Figure 2 , deletion breakpoints at positions 41.9 to 45.5 Mb are significantly associated with speech ability, with the smallest P value (P = 0.0017) occurring at base position 43.9 Mb. None of the 15 individuals with deletion breakpoints at 43.9 Mb or more proximal had “sentences,” whereas 11 of the 20 (55%) subjects with deletions at 44.4 Mb or more distal had “sentences” (RR = 0; P < 0.05). The Youden index was maximum at cutpoint 43.9 Mb, identical to the location identified by the minimum P-value method ( Figure 2 ). The area under the curve (AUC) was 0.80 (“moderately accurate”; Figure 2 ). As shown in Table 2 , none of the 12 individuals with deletions in 22q13.2 was able to speak in sentences as compared with 50% with deletions of 22q13.31 or 60% with deletions occurring at 22q13.33.
Speech ability was also examined in relation to ASDs and degree of developmental delay. The proportion of subjects forming sentences was similar for those who were reported to have an ASD (3 out of 11) as compared with those who were not (8 out of 24, Fisher’s exact test P = 1.0). Speech ability was associated with parent report of degree of developmental delay. Those with sentences had a median developmental delay score of 3 (“moderate”), whereas those without sentences had a median developmental delay score of 6 (“severe to profound”, Wilcoxon rank-sum test P = 0.028). Deletion size and developmental delay score were significantly correlated (Spearman rank correlation coefficient ρ = 0.51; P = 0.0063).
Neonatal features
History of neonatal hypotonia and neonatal feeding problems, as reported by parents, were significantly associated with 22q13.31 to 22q13.32 deletion regions ( Tables 1 and 2 ; Figure 3 ; Supplementary Figure S1 online). In the case of neonatal hypotonia, the Youden index is maximum at chromosome 22 position 45.8 Mb, whereas position 47.4 Mb is the location of the minimum P value (P = 0.0002). Neonatal feeding problems were identified to have similar associated genomic regions.
Abnormal growth
The presence of short stature (<5th percentile) and tall stature (>95th percentile) were moderately associated with distinct deletion regions ( Tables 1 and 2 ; Figure 3 ; Supplementary Figure S1 online). Macrocephaly was associated with the genomic position 44.5 to 46.6 Mb, with optimal cutpoint at position 44.6 Mb and overlapping the genomic region associated with tall stature. Having large or fleshy hands identified the same peak genomic region as macrocephaly. No genomic region was identified as being associated with microcephaly and the AUC for the ROC curve was 0.51 (similar to random chance; Tables 1 and 2; Supplementary Figure S1 online).
ASDs and aggressive behavior
The genomic region from position 41.9 to 46.6 Mb was found to be associated with reduced prevalence of parent-reported diagnosis of an ASD ( Tables 1 and 2 ; Figure 3 ). The graph depicting association statistics is broad and inconsistent (Supplementary Figure S1 online). Aggressive behavior and pinching behavior (toward themselves or others) were also associated with smaller deletions although the statistical support for these associations is less compared with ASDs as neither had an AUC >0.7.
Other features
Other features, including late walking, male genital anomalies, atypical reflexes, dolichocephaly, sacral dimple, bulbous nose, and full brow, were associated with specific genomic regions from 41.1 to 48.7 Mb ( Tables 1 and 2 and Supplementary Figure S1 online). No genomic region was associated with seizures and the AUC for the ROC curve was 0.56 (close to random chance; Supplementary Figure S1 online).
22q13 genes identified as interacting partners with known developmental disability genes
The use of GeneMANIA to search existing physical protein interaction databases identified several genes across the 22q13 deletion region (Supplementary Table S1 and Figure S2 online) not otherwise immediately known as being candidate genes. In particular, WNT7B and PARVB, both located in 22q13.31, were identified as interacting partners of proteins associated with ASDs, ID, and hypotonia, and WNT7B for macrocephaly.
Discussion
This study is the first to identify specific chromosome 22q13.2q13.32 genomic regions, in addition to the terminal 22q13.33 genomic region encompassing SHANK3, associated with key phenotypes in PMS. Strengths of this study design include a relatively large sample size for a rare condition, high resolution genotyping, and widely dispersed breakpoints allowing for resolution between individuals exhibiting different phenotypes. An additional strength is that we used well-established statistical techniques in a novel manner to a genotype–phenotype study to identify candidate genes. Traditionally, genotype–phenotype studies of chromosomal deletions look to identify the smallest region of overlap or the smallest common deletion in a small group of patients with a rare phenotype to identify candidate genes. This traditional method is useful for monogenic disorders with high penetrance and low variability but not as helpful for common or multifactorial phenotypes and genes with variable expressivity or incomplete penetrance. In the case of PMS, we find that some features are found in those with the smallest deletions, but are much more common in those with larger deletions. For instance, neonatal hypotonia and late walking were reported in 20% of those with deletions of just 22q13.33 yet were reported for more than 90% of those with the largest deletions (22q13.2). The traditional approach would ignore the difference in frequency and identify 22q13.33 as being the only candidate region. In previous works on PMS, there was a general impression that patients with larger deletions were more seriously affected, but genotype–phenotype studies were hampered by small sample size, low resolution genotyping, or reliance on statistical measures of linear association (correlation coefficients, linear regression),4,8,9,23,24 whereas the present study overcomes these limitations.
Because our patient cohort consists of those with terminal deletions and all patients are missing one copy of SHANK3, we cannot distinguish whether the more proximal genomic regions we identified have independent or additive effects along with SHANK3. The literature reports three cases of interstitial deletions which have intact SHANK3 and phenotypes similar to those in PMS,25,26 suggesting an independent role for genes in these genomic regions. In particular, two individuals had speech delay (two words each and no sentences), macrocephaly, tall stature, hypotonia, delayed walking, and developmental delay, yet had two copies of SHANK3.26 Their deletion breakpoints are reported to be between 40.42 and 44.00 Mb for one patient and between 41.22 and 45.37 Mb for the other. It should be noted that one of these interstitial deletions was inherited from a mother with only mild speech deficits, further demonstrating the phenotypic variability of these rearrangements. In addition, a positional effect of the deletion on SHANK3 expression, such as through disruption of an enhancer element, cannot be excluded. As shown in Figure 3 , CYB5R3, PARVB, and hsa-mir-1249 all map in the region deleted in these interstitial cases.
Speech and language delay
The variability in speech ability in those with PMS and the often vague descriptions of this feature in the PMS literature hamper genotype–phenotype comparisons. Frequently no distinction is made between “absent” and “delayed” speech. The observed variability could be due to variable expressivity of the SHANK3 deletion, additional gene loss on 22q13 in combination with SHANK3 loss, or additional mutations or exposures. Even those with small deletions may still be significantly affected. For instance, one case report of a boy with a translocation disrupting SHANK3 had only a few words.3 Another case of a small deletion due to a translocation within SHANK3 was in a young woman whose language was “significantly delayed” at the age of 4 years, and at the age of 20 years, her “verbal language was simple and pronunciation so blurred that her speech was difficult to understand.” With the benefit of larger sample sizes and objectively categorized speech levels, we observed a distinct difference in speech ability by deletion size.
The genomic region associated with lack of speech contains an estimated 45 protein coding genes as well as miRNAs and other noncoding RNAs. The genes PARVB and WNT7B, found to be interacting partners of genes known to be associated with ASDs, ID, and hypotonia, are also in this region. The findings of severe speech impairment among two published cases with interstitial deletions overlapping our genomic region of interest and intact SHANK326 support the presence of genes affecting speech in this region. None of the genes in this region was found to be a transcriptional target of FOXP2, a transcription factor known to be associated with speech.27,28 Beyond FOXP2, little is known about genes related to speech, although recent studies have added CNTNAP2, CMIP, ATP2C2, RIT2, and SYT4 as potential genes of interest.29,30 Future research into speech and language abilities in PMS would benefit from having detailed evaluations to better characterize the types of language delay specific to this syndrome.
Abnormal growth
We recently reported that both tall (>95th percentile) and short stature (<5th percentile) as well as macrocephaly (>97th percentile) are more common in PMS than expected.31 This analysis provides evidence of distinct deletion regions associated with these growth parameters, which supports earlier findings by others.4,9
Autism spectrum disorders
In our analysis we noted that ASDs were associated with smaller deletions. SHANK3 mutations have been found to be associated with ASDs.32,33,34,35 In patients with PMS, the effect of SHANK3 may be attenuated as the deletion size increases and additional genes are codeleted. It may also be more difficult to evaluate ASDs in patients with severe developmental, speech, and motor impairments, which are associated with larger deletions. Future research is needed to better delineate the autism phenotype in PMS patients and those with SHANK3 deletions or mutations to better identify particular domains affected.
Review of potential genes of interest
A large number of potentially interesting candidate genes were identified in this analysis, most with expression observed in relevant tissue types as described in the EST profile database (http://www.ncbi.nlm.nih.gov/nucest). Few of these genes have been previously implicated as causative for human phenotypes. Of particular interest are NUP50 (nucleoporin 50), CERK (ceramid kinase), C22orf9 (also known as LOC23313 or KIAA0930), KIAA1644, PHF21B (PHD finger protein), ATXN10 (ataxin 10), FBLN1 (Fibulin-1), and CELSR1 (cadherin EGF LAG seven-pass G-type receptor 1). To prioritize candidate genes, we initially examined the prevalence of copy number variants encompassing these genes in controls as provided in the Database of Genomic Variants.36 However, this assessment method was problematic for PMS in that the primary candidate gene for PMS is the well-studied SHANK3, and SHANK3 itself has many entries in the Database of Genomic Variants of controls having duplications and deletions of the gene. A two-hit model may explain the observation that SHANK3 deletions are observed in some controls but nearly all cases of PMS. A two-hit model could also explain other candidate genes. Thus, we we did not use this information for prioritizing candidate genes on the 22q13 region. A more detailed review of these genes is provided in the Supplementary Materials online.
Two of the genes that were identified using GeneMANIA protein interaction networks were PARVB (Beta parvin or affixin) and WNT7B (wingless-type MMTV integration site family). Both proteins were found to be physically interacting partners with proteins associated with ASDs, ID, and hypotonia, and WNT7B was also found to interact with macrocephaly-associated proteins (Supplementary Table S1 and Figure S2 online). PARVB is deleted in the regions associated with lack of ASDs, absent speech, touch sensitivity, hair-pulling behavior, large/fleshy hands, and male genital anomalies. It is also deleted in both cases of interstitial deletions.26 WNT7B is deleted in one case of interstitial deletion26 and is located in the regions associated with macrocephaly, large/fleshy hands, dysplastic toenails, dolichocephaly, bulbous nose, lack of ASDs, lack of aggression, lack of pinching behavior, absent speech, walking late, hair-pulling behavior, neonatal hypotonia, neonatal feeding problems, and male genital anomalies. PARVB and SHANK3 are both scaffolding proteins important in postsynaptic structures.37 PARVB binds ARHGEF6 (Rac/Cdc42 guanine nucleotide exchange factor (GEF) 6; also called alphaPIX).38 Mutations in the PARVB binding site on ARHGEF6 are associated with X-linked ID.39 ARHGEF6 heterodimerizes with ARHGEF7,38 a binding partner of SHANK3, a candidate gene for PMS as well as ASDs.3,5,32,33,34,35 Supplementary Figure S2 online illustrates the physical interactions between PARVB and SHANK3 as obtained from GeneMANIA. Thus, PARVB may have overlapping and interacting roles with SHANK3.37
Finally, we note the presence of several miRNAs in 22q13.31 including hsa-mir-1249 which is located within an intronic region of C22orf9. This miRNA is predicted to target the 22q13 genes SHANK3, PHF21B, and SERHL2 and also a number of other brain- and development-associated genes (TargetScan).40 Loss of hsa-mir-1249 may lead to misregulation of the preserved copy of SHANK3, affecting the severity phenotypes associated with SHANK3 haploinsufficiency. The fact that hsa-mir-1249 maps within the regions of interstitial deletions associated with PMS25,26 probably represents further evidence of the potential role played by these miRNAs in neurodevelopmental disorders.
The remaining genes, noncoding RNAs, and regulatory elements in genomic regions associated with various phenotypes, particularly in the chromosome 22q13.31 region, deserve further examination in future studies.
Limitations
There are several limitations in this study. All participants had been previously diagnosed with a 22q13 deletion, although the methods used to make this determination varied greatly including fluorescence in situ hybridization, chromosome analysis, and whole-genome arrays of varying levels of sensitivity. While all patients in this cohort have a confirmed 22q13 deletion as determined by our custom 22q13 array CGH and patients with a known structural anomaly other than a simple terminal deletion were excluded from our analysis, we did not conduct a genome-wide aCGH and cannot exclude the possibility that some individuals harbor balanced rearrangements or additional significant copy-number variations. Reporting of some features relied upon parent report and may be subject to recall or information bias. In particular, the results of standardized assessments for ASD and developmental delay were not available, and thus analyses relied upon parent reporting of these diagnoses. Association statistics are heavily influenced by sample size, and this study had small statistical power to examine associations with genomic regions near the telomere (the smallest deletions) and most proximal (those with deletions >8 Mb in size). No statistical adjustment was made for the fact that multiple phenotypes were examined in the same population.
Conclusion
We identified 22q13.2q13.32 genomic regions associated with severity of speech/language delay, neonatal hypotonia, delayed age at walking, hair-pulling behaviors, male genital anomalies, dysplastic toenails, large/fleshy hands, macrocephaly, short and tall stature, facial asymmetry, and abnormal reflexes. These results indicate that, although the terminal 22q13.33 region encompassing SHANK3 may be critical for some of the PMS phenotypes, additional, more proximal genomic regions are important to determine the severity of some symptoms and the variable occurrence of many secondary features of the syndrome. Therefore, we believe that future studies on the role of SHANK3, IB2, and other telomeric genes in PMS should also include efforts to determine the independent and additive effects of loss of 22q13.2q13.32 genes, noncoding RNAs, or regulatory elements and position effects.
Disclosure
The authors declare no conflict of interest.
References
Phelan K, McDermid HE . The 22q13.3 Deletion Syndrome (Phelan-McDermid Syndrome). Mol Syndromol 2012;2:186–201.
Bonaglia MC, Giorda R, Ciccone R, Zuffardi O . Chromosome 22q13 rearrangements causing global developmental delay and autistic spectrum disorder. In: Knight SJL (ed). Genetics of Mental Retardation. Karger: Basel, 2010:137–150.
Bonaglia MC, Giorda R, Borgatti R, et al. Disruption of the ProSAP2 gene in a t(12;22)(q24.1;q13.3) is associated with the 22q13.3 deletion syndrome. Am J Hum Genet 2001;69:261–268.
Wilson HL, Wong AC, Shaw SR, et al. Molecular characterisation of the 22q13 deletion syndrome supports the role of haploinsufficiency of SHANK3/PROSAP2 in the major neurological symptoms. J Med Genet 2003;40:575–584.
Delahaye A, Toutain A, Aboura A, et al. Chromosome 22q13.3 deletion syndrome with a de novo interstitial 22q13.3 cryptic deletion disrupting SHANK3. Eur J Med Genet 2009;52:328–332.
Giza J, Urbanski MJ, Prestori F, et al. Behavioral and cerebellar transmission deficits in mice lacking the autism-linked gene islet brain-2. J Neurosci 2010;30:14805–14816.
Misceo D, Rødningen OK, Barøy T, et al. A translocation between Xq21.33 and 22q13.33 causes an intragenic SHANK3 deletion in a woman with Phelan-McDermid syndrome and hypergonadotropic hypogonadism. Am J Med Genet A 2011;155A:403–408.
Jeffries AR, Curran S, Elmslie F, et al. Molecular and phenotypic characterization of ring chromosome 22. Am J Med Genet A 2005;137:139–147.
Dhar SU, del Gaudio D, German JR, et al. 22q13.3 deletion syndrome: clinical and molecular analysis using array CGH. Am J Med Genet A 2010;152A:573–581.
Sarasua SM, Dwivedi A, Boccuto L, et al. Association between deletion size and important phenotypes expands the genomic region of interest in Phelan-McDermid syndrome (22q13 deletion syndrome). J Med Genet 2011;48:761–766.
Altman DG, Lausen B, Sauerbrei W, Schumacher M . Dangers of using “optimal” cutpoints in the evaluation of prognostic factors. J Natl Cancer Inst 1994;86:829–835.
Williams BA, Mandrekar JN, Mandrekar SJ, Cha SS, Furth AF . Finding optimal cutpoints for continuous covariates with binary and time-to-event outcomes. Technical Report Series #79. Mayo Clinic: Rochester, MN, 2006.
Bewick V, Cheek L, Ball J . Statistics review 13: receiver operating characteristic curves. Crit Care 2004;8:508–512.
Hanley JA, McNeil BJ . The meaning and use of the area under a receiver operating characteristic (ROC) curve. Radiology 1982;143:29–36.
Greiner M, Pfeiffer D, Smith RD . Principles and practical application of the receiver-operating characteristic analysis for diagnostic tests. Prev Vet Med 2000;45:23–41.
Basu SN, Kollu R, Banerjee-Basu S . AutDB: a gene reference resource for autism research. Nucleic Acids Res 2009;37(Database issue):D832–D836.
Kou Y, Betancur C, Xu H, Buxbaum JD, Ma’ayan A . Network- and attribute-based classifiers can prioritize genes and pathways for autism spectrum disorders and intellectual disability. Am J Med Genet C Semin Med Genet 2012;160C:130–142.
Pinto D, Pagnamenta AT, Klei L, et al. Functional impact of global rare copy number variation in autism spectrum disorders. Nature 2010;466:368–372.
Sakai Y, Shaw CA, Dawson BC, et al. Protein interactome reveals converging molecular pathways among autism disorders. Sci Transl Med 2011;3:86ra49.
Lubs HA, Stevenson RE, Schwartz CE . Fragile X and X-linked intellectual disability: four decades of discovery. Am J Hum Genet 2012;90:579–590.
Chiurazzi P, Schwartz CE, Gecz J, Neri G . XLMR genes: update 2007. Eur J Hum Genet 2008;16:422–434.
Warde-Farley D, Donaldson SL, Comes O, et al. The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res 2010;38:W214–W220.
Koolen DA, Reardon W, Rosser EM, et al. Molecular characterisation of patients with subtelomeric 22q abnormalities using chromosome specific array-based comparative genomic hybridisation. Eur J Hum Genet 2005;13:1019–1024.
Philippe A, Boddaert N, Vaivre-Douret L, et al. Neurobehavioral profile and brain imaging study of the 22q13.3 deletion syndrome in childhood. Pediatrics 2008;122:e376–e382.
Fujita Y, Mochizuki D, Mori Y, et al. Girl with accelerated growth, hearing loss, inner ear anomalies, delayed myelination of the brain, and del(22)(q13.1q13.2). Am J Med Genet 2000;92:195–199.
Wilson HL, Crolla JA, Walker D, et al. Interstitial 22q13 deletions: genes other than SHANK3 have major effects on cognitive and language development. Eur J Hum Genet 2008;16:1301–1310.
Spiteri E, Konopka G, Coppola G, et al. Identification of the transcriptional targets of FOXP2, a gene linked to speech and language, in developing human brain. Am J Hum Genet 2007;81:1144–1157.
Vernes SC, Spiteri E, Nicod J, et al. High-throughput analysis of promoter occupancy reveals direct neural targets of FOXP2, a gene mutated in speech and language disorders. Am J Hum Genet 2007;81:1232–1250.
Bouquillon S, Andrieux J, Landais E, et al. A 5.3Mb deletion in chromosome 18q12.3 as the smallest region of overlap in two patients with expressive speech delay. Eur J Med Genet 2011;54:194–197.
Newbury DF, Fisher SE, Monaco AP . Recent advances in the genetics of language impairment. Genome Med 2010;2:6.
Rollins JD, Sarasua SM, Phelan K, DuPont BR, Rogers RC, Collins JS . Growth in Phelan-McDermid syndrome. Am J Med Genet A 2011;155A:2324–2326.
Durand CM, Betancur C, Boeckers TM, et al. Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nat Genet 2007;39:25–27.
Gauthier J, Spiegelman D, Piton A, et al. Novel de novo SHANK3 mutation in autistic patients. Am J Med Genet B Neuropsychiatr Genet 2009;150B:421–424.
Moessner R, Marshall CR, Sutcliffe JS, et al. Contribution of SHANK3 mutations to autism spectrum disorder. Am J Hum Genet 2007;81:1289–1297.
Boccuto L, Lauri M, Sarasua SM, et al. Prevalence of SHANK3 variants in patients with different subtypes of autism spectrum disorders. Eur J Hum Genet 2013;21:310–316.
Iafrate AJ, Feuk L, Rivera MN, et al. Detection of large-scale variation in the human genome. Nat Genet 2004;36:949–951.
Govek EE, Newey SE, Van Aelst L . The role of the Rho GTPases in neuronal development. Genes Dev 2005;19:1–49.
Rosenberger G, Jantke I, Gal A, Kutsche K . Interaction of alphaPIX (ARHGEF6) with beta-parvin (PARVB) suggests an involvement of alphaPIX in integrin-mediated signaling. Hum Mol Genet 2003;12:155–167.
Kutsche K, Yntema H, Brandt A, et al. Mutations in ARHGEF6, encoding a guanine nucleotide exchange factor for Rho GTPases, in patients with X-linked mental retardation. Nat Genet 2000;26:247–250.
Friedman RC, Farh KK, Burge CB, Bartel DP . Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 2009;19:92–105.
Acknowledgements
We thank the patients and family members who participated in this study and made this work possible. We thank the Phelan–McDermid Syndrome Foundation which organized the biannual family conferences where much of the study data collection took place. We thank Amy Lawton-Rauh and Charles E. Schwartz for helpful comments on the manuscript. We thank Gail Stapleton and Cindy Skinner who coordinated study operations. We dedicate this article to the late J.S.C. This work was supported by the Phelan–McDermid Syndrome Foundation (to S.M.S.); the Genetics Endowment of South Carolina; and the South Carolina Department of Disabilities and Special Needs.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Materials
(PDF 530 kb)
Rights and permissions
About this article
Cite this article
Sarasua, S., Dwivedi, A., Boccuto, L. et al. 22q13.2q13.32 genomic regions associated with severity of speech delay, developmental delay, and physical features in Phelan–McDermid syndrome. Genet Med 16, 318–328 (2014). https://doi.org/10.1038/gim.2013.144
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2013.144
Keywords
This article is cited by
-
Haploinsufficiency of Shank3 increases the orientation selectivity of V1 neurons
Scientific Reports (2022)
-
DNA methylation epi-signature is associated with two molecularly and phenotypically distinct clinical subtypes of Phelan-McDermid syndrome
Clinical Epigenetics (2021)
-
Speech and language phenotype in Phelan-McDermid (22q13.3) syndrome
European Journal of Human Genetics (2021)
-
Bringing everyone to the table – findings from the 2018 Phelan-McDermid Syndrome Foundation International Conference
Orphanet Journal of Rare Diseases (2020)
-
Transcriptional signatures of participant-derived neural progenitor cells and neurons implicate altered Wnt signaling in Phelan-McDermid syndrome and autism
Molecular Autism (2020)