Abstract
Purpose: Dihydropyrimidine dehydrogenase is a critical enzyme in the catabolism of 5-Fluorouracil, a drug frequently used in cancer therapy. Patients with deficient dihydropyrimidine dehydrogenase activity are at risk of developing severe 5-Fluorouracil–associated toxicity. Genetic analysis of the gene coding for dihydropyrimidine dehydrogenase has shown that mutations in exon 14, especially the splice-site mutation IVS14+1G→A, were associated with dihydropyrimidine dehydrogenase enzymatic deficiency.
Methods: We evaluated the frequency of mutations in exon 14 of dihydropyrimidine dehydrogenase (DPYD) gene in 73 unselected colorectal cancer patients treated with 5-Fluorouracil after surgery at a Portuguese Cancer Institute.
Results: Sequencing the entire exon 14 allowed the detection of mutations in two of the 73 patients (2.7%), namely two of the eight (25%) patients who presented grade 3-4 toxicity after 5-Fluorouracil chemotherapy. One patient was heterozygous for the splice-site mutation IVS14+1G→A, whereas the second patient was heterozygous for a novel missense mutation 1845G→T (E615D) in exon 14 of DPYD gene.
Conclusion: We conclude that mutations in exon 14 of DPYD gene are responsible for a significant proportion of life-threatening toxicity to 5-Fluorouracil, and should therefore be excluded before its administration to cancer patients.
Similar content being viewed by others
Main
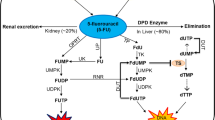
5-Fluorouracil (5-FU) is one of the most commonly used chemotherapeutic drugs for the treatment of malignant neoplasms, namely colorectal cancer. 5-FU is a pyrimidine analogue, giving rise to cytotoxic metabolites after metabolization by the pyrimidine pathways. One of them, the 5-fluoro-dUMP, inhibits thymidylate synthase, thus impairing DNA synthesis. Additional cytotoxic effects arise from 5-FU metabolites being incorporated into RNA and DNA.1
Dihydropyrimidine dehydrogenase (DPD) is the initial and rate-limiting enzyme in the catabolism of 5-FU. It has been reported that more than 80% of the administered 5-FU is catabolized by DPD,2–4 so the activity of this enzyme may be of paramount importance to predict the efficacy and toxicity of this drug. Deficiency in DPD enzyme activity has been correlated with considerable delay in 5-FU clearance from plasma.4,5 The important role of DPD in 5-FU–based chemotherapy has been demonstrated in cancer patients with major deficiency of this enzyme's activity. These patients may present severe toxicity after the administration of 5-FU, including diarrhea, neutropenia, and neurotoxicity, and death may occasionally occur.6–9 Population studies suggested that, although total deficiency is rare, as many as 3% to 5% of the population may have low enzyme levels and thus be at increased risk of severe toxicity if treated with 5-FU.10
Several of these patients are genotypically heterozygous for a mutant DPYD allele.11–16 To date, more than 30 variant DPYD alleles have been identified, some of which result in decreased activity of DPD enzyme.17–19 Analysis of the prevalence of the various mutations has shown that the splice-site mutation IVS14+1G→A is by far the most common.13,14 This mutation results in a 165-bp deletion (corresponding to exon 14) in the DPYD mRNA. The resulting protein product is truncated by 55 amino acids, and its catalytic activity is virtually absent. Some reports have shown that the splice-site mutation IVS14+1G→A is present in approximately 1% of the Caucasian population.17,18 In addition, a recent study showed that approximately 50% of the patients who had experienced severe 5-FU toxicity (grade 3-4) were heterozygous for this exon-skipping mutation.19
To evaluate the frequency of mutations in exon 14 of DPYD and its contribution to 5-FU toxicity in Portuguese colorectal cancer (CRC) patients treated with 5-FU, we have performed full sequencing of exon 14 in 73 CRC patients treated with this adjuvant chemotherapeutic regimen after surgery.
MATERIALS AND METHODS
Patients and toxicity evaluation
Seventy-three consecutive patients [34 males and 39 females; mean age, 59 (range, 31–85) years] with CRC, who were treated with potentially curative surgical resections followed by adjuvant 5-FU–based chemotherapy, were included in this study (Table 1). Informed consent was obtained from all patients studied. Chemotherapy toxicity was evaluated according to the National Cancer Institute toxicity scale.20 The main documented toxicities were hematological, gastrointestinal, mucocutaneous, and neurological.
Mutation analysis
Genomic DNA was isolated from peripheral mononuclear cells using a DNA Extraction Kit (Invitrogen) following the manufacturer's instructions. PCR amplification of exon 14 and its flanking intronic regions was performed using the following primer set16: DPD14F, 5′-GTAAGAAGGACCTCATAAAATATTGTC-3′; DPD14R, 5′-GAATTGGATGTTTAAATAAACATTCACCAAC-3′. Amplification of exon 14 was performed in a 25 μL reaction mixture containing 10 mmol/L Tris-HCL (pH 8.3), 50 mmol/L KCl, 1.5 mmol/L MgCl2, 10 pmol of each primer, 250 μmol/L of each deoxynucleotide triphosphate, 0.2 Units of Taq polymerase, and 50 ng of genomic DNA. After initial denaturation for 2 minutes at 95°C, amplification was performed for 35 cycles (1 minute 95°C, 1 minute 55°C, 1 minute 72°C). The expected fragment size with this primer set is 342 bp. The PCR products were separated on 2% agarose gels, stained with ethidium bromide, purified using a GFX PCR DNA and Gel Band Purification Kit (Amersham Pharmacia Biotech) and used for direct sequencing. Sequence analysis was performed on an Applied Biosystems model 310 automated DNA sequencer using the dye-terminator method.21
Statistical analysis
Comparisons of allelic frequencies between patients with and without grade 3-4 5-FU toxicity was performed with Fisher exact test. Two-tailed values of P < 0.05 were considered statistically significant.
RESULTS
Identification of mutations in exon 14 of DPYD
Analysis of exon 14 of the DPYD gene in 73 patients with CRC revealed the presence of mutations in two of them (2.7%). One patient was heterozygous for the splice-site mutation IVS14+1G→A and one patient was heterozygous for a novel missense mutation 1845G→T (E615D) in exon 14 of DPYD (Fig. 1). Therefore, the prevalence of each of these mutant alleles in colorectal patients was 0.7%.
Clinical evaluation of toxicity
Of the 73 patients, 65 presented grade 1-2 toxicity and eight (11%) suffered from grade 3 or 4 toxicity related to 5-FU chemotherapy (Table 2). The two mutations described above were found in two of the eight patients (25%) who developed grade 3 or 4 toxicity. A statistically significant association between exon 14 DPYD gene mutations and grade 3-4 5-FU toxicity was found (P = 0.01). The patient heterozygous for the splice-site mutation IVS14+1G→A was a man with a rectal carcinoma diagnosed at age 64 (pT3N1/10M0). He was submitted to anterior resection of the upper rectum and treated postoperatively with radiotherapy, followed by adjuvant chemotherapy with 5-FU-folinic acid (bolus). Therapy was stopped after the second cycle due to grade 4 febril neutropenia and grade 3 digestive toxicity. The patient heterozygous for the missense mutation 1845G→T was a woman with a rectal cancer diagnosed at age 37 (pT4N2M0). She was submitted to abdominoperineal resection of the lower third of the rectum. Radiotherapy with 5-FU chemotherapy sensibilization was started after surgery. After radiotherapy was completed, 5-FU folinic acid chemotherapy was administered (bolus). On the 6th day after chemotherapy, grade 4 mucositis was documented, followed by febril pancytopenia and sepsis due to Pneumocystis carinii. Mechanical ventilation support was necessary due to respiratory failure.
Sensitivity and specificity of DPYD mutations
Because two of the eight patients with grade 3-4 toxicity were shown to have DPYD mutations and all patients with grade 0-2 toxicity were negative, the sensitivity and specificity of the test is 25% and 100%, respectively.
Positive and negative predictive value
The positive predictive value of the test was 100% (2/2), whereas its negative predictive value was 92% (65/71).
DISCUSSION
Several population studies have demonstrated that total DPD deficiency is rare, but as many as 3% to 5% of the population may have low enzyme levels and thus be at increased risk for severe toxicity if treated with 5-FU.10,22 It has subsequently been demonstrated that a number of these individuals were genotypically heterozygous for a mutant DPYD allele,11–16 but data are scant on the frequency of mutations in DPYD among cancer patients. In this study, we showed that the prevalence of the mutation in exon 14 of DPYD in a CRC population treated with 5-FU was 2.7%. Exon 14 DPYD mutations showed a statistically significant association with grade 3-4 5-FU toxicity (P = 0.01). The frequency of DPYD in our colorectal cancer population was not significantly different from that reported by Ridge et al.15 (2.7 vs. 4%).
To date, more than 30 variant DPYD alleles have been identified, some of which clearly result in decreased DPD activity.17–19 In our group of patients, two different mutations were identified: the splice-site mutation IVS14+1G→A and the missense mutation 1845G→T. The mutation IVS14+1G→A is the most frequently detected in patients with complete DPD activity deficiency.13,14 This point mutation in the invariant GT splice donor site leads to the skipping of exon 14 in the DPYD mRNA, which results in a truncated protein by 55 amino acids with virtually no catalytic activity. This mutation was detected in one of eight patients with severe 5-FU toxicity, which is in line with other published studies.7,8,13–16 A few reports have recently described the prevalence of the exon 14–skipping mutation in colorectal cancer and healthy populations. Van Kuilenburg et al.18 reported a frequency of 0.9% in 1357 Caucasian blood donors. Furthermore, Raida et al.17 reported the IVS14+1G→A mutation in 0.94% of the alleles in 851 Caucasian blood donors. The allelic frequency found in the present study (0.7%) does not significantly differ from the findings in these two reports.
We identified a novel missense mutation, 1845G→T, in exon 14 of DPYD in a patient who suffered from severe 5-FU toxicity. Although DPD activity was not determined in this patient, the observed toxicity profile and the location of the mutation in an important functional domain of the protein (a uracil-binding site), suggest that this alteration may be pathogenic. A recent study demonstrated that missense mutations present in highly conserved regions of the DPYD gene may interfere with protein function.23 However, additional studies, such as determination of DPD activity and other functional studies, are necessary to determine the significance of this mutation.
Raida et al.17 has previously reported that 6 of 25 patients who experienced grade 3-4 toxicity upon 5-FU treatment carried the IVS14+1G→A mutation (five were heterozygous and one was homozygous for this splice-site mutation). In the present series, two of eight patients (25%) with grade 3-4 toxicity had mutations in exon 14 of DPYD, although only one (12.5%) was heterozygous for the IVS14+1G→A mutation. The fact that we detected a novel mutation (1845G→T) in exon 14 of DPYD indicates that full exon screening, not only the exon-skipping mutation, should be performed. Whereas our test revealed 100% positive predictive value and 100% specificity, six patients with severe toxicity to 5-FU treatment showed no mutations in exon 14 of DPYD, corresponding to a sensitivity of 25% and a negative predictive value of 92%. We cannot exclude the possibility that these patients are carriers of other loss-of-function mutations in the coding sequence or promoter region of DPYD.24 Alternatively, increased expression of enzymes of the anabolic pathway may be responsible for the observed increased sensitivity to 5-FU in patients without DPYD mutations.
In conclusion, the common use of 5-FU in cancer therapy and the relatively high frequency of mutations in exon 14 of DPYD in patients developing grade 3-4 toxicity, indicate that genetic screening for the presence of DPYD mutations in cancer patients before the administration of 5-FU is appropriate, in order to prevent serious and sometimes lethal 5-FU-related toxicities.
References
Schilsky RL . Biochemical and clinical pharmacology of 5-Fluorouracil. Oncology 1998; 12( suppl 7): 13–18.
Vincent M, Labianca R, Harper P . Which 5-Fluorouracil regimen? the great debate. Anticancer Drugs 1999; 10: 337–354.
Heggie GD, Somadsossi JP, Cross DS, Huster WJ, Diasio RB . Clinical pharmacokinetics of 5-Fluorouracil and its metabolism in plasma, urine, and bile. Cancer Res 1987; 47: 2203–2206.
Diasio RB, Lu Z . Dihydropyrimidine dehydrogenase activity and fluorouracil chemotherapy (editorial). J Clin Oncol 1994; 12: 2239–2242.
Fleming RA, Milano G, Thyss A, Etienne MC, Renée N, Schneider M et al. Correlation between dihydropyrimidine dehydrogenase activity in peripheral mononuclear cells and systemic clearance of fluorouracil in cancer patients. Cancer Res 1992; 52: 2899–2902.
Milano G, Etienne MC . Potential importance of dihydropyrimidine dehydrogenase (DPD) in cancer chemotherapy. Pharmacogenetics 1994; 4: 301–306.
Wei X, Mcleod HL, McMurrough J, Gonzalez FJ, Fernandez-Salguero PM . Molecular Basis of the human dihydropyrimidine dehydrogenase deficiency and 5-Fluorouracil toxicity. J Clin Invest 1996; 98: 610–615.
Van Kuilenburg ABP, Vreken P, Beex LVAM, Meinsma R, Van Lenthe H, De Abreu RA et al. Heterozygosity for a point mutation in a invariant splice donor site of dihydropyrimidine dehydrogenase and severe 5-Fluorouracil related toxicity. Eur J Cancer 1997; 13: 2258–2264,.
Kouwaki M, Hamajima N, Sumi S, Nonaka M, Sasaki M, Dobashi K et al. Identification of a novel mutation in the dihydropyrimidine dehydrogenase gene in a Japanese patient with 5-Fluorouracil toxicity. Clin Cancer Res 1998; 4: 2999–3004.
Etienne MC, Lagrange JL, Dassonville O, Fleming R, Thyss A, Renée N et al. Population study of dihydropyrimidine dehydrogenase in cancer patients. J Clin Oncol 1994; 12: 2248–2253.
Vreken P, Van Kuilenburg ABP, Meinsma R, De Abreu RA, Van Gennip AH . Identification of a four-base deletion (delTCAT 296–299) in the dihydropyrimidine dehydrogenase gene with variable clinical expression. Hum Genet 1997; 100: 263–265.
Vreken P, Van Kuilenburg ABP, Meinsma R, Van Gennip AH . Dihydropyrimidine dehydrogenase (DPD) deficiency: identification and expression of missense mutations C29R, R886H, R235W. Hum Genet 1997; 101: 333–338.
Van Kuilenburg ABP, Vreken P, Abeling NGGM, Bakker HD, Meinsma R, Van Lenthe H, De Abreu RA et al. Genotype and phenotype in patients with dihydropyrimidine dehydrogenase deficiency. Hum Genet 1999; 104: 1–9.
Van Kuilenburg ABP, Haasjes J, Richel DJ, Zoetekouw L, Van Lenthe H, De Abreu RA et al. Clinical implications of dihydropyrimidine dehydrogenase (DPD) deficiency in patients with severe 5-Fluorouracil-associated toxicity: Identification of new mutations in the DPD gene. Clin Cancer Res 2000; 6: 4705–4712.
Ridge SA, Sludden J, Wei X, Sapone A, Brown O, Hardy S et al. Dihydropyrimidine dehydrogenase pharmacogenetics in patients with colorectal cancer. Br J Cancer 1998; 77: 497–500.
Johnson MR, Hageboutros A, Wang K, High L, Smith JB, Diasio RB . Life-threatening toxicity in a dihydropyrimidine dehydrogenase-deficient patient after treatment with topical 5-Fluorouracil. Clin Cancer Res 1999; 5: 2006–2011.
Raida M, Schwabe W, Hausler P, Van Kuilenburg ABP, Van Gennip AH, Behnke D et al. Prevalence of a common point mutation in the dihydropyrimidine dehydrogenase (DPD) gene within the 5′-splice donor site of intron 14 in patients with severe 5-fluorouracil (5-FU)- related toxicity compared with controls. Clin Cancer Res 2001; 7: 2832–2839.
Van Kuilenburg ABP, Muller EW, Haasjes J, Meinsma R, Zoetekouw L, Watherham HR et al. Lethal outcome of patient with a complete dihydropyrimidine dehydrogenase (DPD) deficiency after administration of 5-fluorouracil: frequency of the common IVS14+1G >A mutation causing DPD deficiency. Clin Cancer Res 2001; 7: 1149–1153.
Van Kuilenburg ABP, Meinsma R, Zoetekouw L, Van Gennip AH . Increased risk of grade IV neutropenia after administration of 5-Fluorouracil due to a dihydropyrimidine dehydrogenase (DPD) deficiency: High prevalence of the IVS14+1G >A mutation. Int J Cancer 2002; 101: 253–258.
National Cancer Institute toxicity scale. Available at: http://ctep.cancer.gov/reporting/ctc.html. Accessed
Sanger F, Air GM, Barrel BG, Brown NL, Coulson AR, Fiddes CA et al. Nucleotide sequence of bacteriophage φX174. Nature 1977; 265: 678–695.
Lu Z, Zhang R, Diasio R . Dihydropyrimidine dehydrogenase activity in human peripheral blood mononuclear cells and liver: Population characteristics newly identified deficient patients, and clinical implication in 5-Fluorouracil chemotherapy. Cancer Res 1993; 53: 5433–5438.
Van Kuilenburg ABP, Dobritzsch D, Meinsma R, Haasjes J, Wathermen HR, Nowaczyk MJN et al. Novel disease-causing mutations in the dihydropyrimidine dehydrogenase gene interpreted by analysis of the three-dimensional protein structure. Biochem J 2002; 364: 157–63.
Collie-Duguid ESR, Jonhston SJ, Powrie RH, Milano G, Etienne MC, Rochat B et al. Cloning and initial characterization of the human DPYD gene promoter. Biochem Biophys Res Commun 2000; 271: 28–35.
Acknowledgements
The financial support of “Liga Portuguesa Contra o Cancro, Centro Regional do Norte” is gratefully acknowledged. This work was awarded a prize by “Comissão de Fomento da Investigação em Cuidados de Saúde (Ministério da Saúde P.I. N° 218/01).”
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Salgueiro, N., Veiga, I., Fragoso, M. et al. Mutations in exon 14 of dihydropyrimidine dehydrogenase and 5-Fluorouracil toxicity in Portuguese colorectal cancer patients. Genet Med 6, 102–107 (2004). https://doi.org/10.1097/01.GIM.0000118061.66602.A5
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1097/01.GIM.0000118061.66602.A5
Keywords
This article is cited by
-
Evaluation of predictive tests for screening for dihydropyrimidine dehydrogenase deficiency
The Pharmacogenomics Journal (2013)
-
Polymorphism of TS 3′-UTR predicts survival of Chinese advanced gastric cancer patients receiving first-line capecitabine plus paclitaxel
Clinical and Translational Oncology (2013)
-
Promoter methylation and large intragenic rearrangements of DPYD are not implicated in severe toxicity to 5-fluorouracil-based chemotherapy in gastrointestinal cancer patients
BMC Cancer (2010)
-
Dihydropyrimidine dehydrogenase polymorphisms and fluoropyrimidine toxicity: ready for routine clinical application within personalized medicine?
EPMA Journal (2010)