Abstract
Protein synthesis regulation via mammalian target of rapamycin complex 1 (mTORC1) signaling pathway has key roles in neural development and function, and its dysregulation is involved in neurodevelopmental disorders associated with autism and intellectual disability. mTOR regulates assembly of the translation initiation machinery by interacting with the eukaryotic initiation factor eIF3 complex and by controlling phosphorylation of key translational regulators. Collybistin (CB), a neuron-specific Rho-GEF responsible for X-linked intellectual disability with epilepsy, also interacts with eIF3, and its binding partner gephyrin associates with mTOR. Therefore, we hypothesized that CB also binds mTOR and affects mTORC1 signaling activity in neuronal cells. Here, by using induced pluripotent stem cell-derived neural progenitor cells from a male patient with a deletion of entire CB gene and from control individuals, as well as a heterologous expression system, we describe that CB physically interacts with mTOR and inhibits mTORC1 signaling pathway and protein synthesis. These findings suggest that disinhibited mTORC1 signaling may also contribute to the pathological process in patients with loss-of-function variants in CB.
Similar content being viewed by others
Introduction
The mammalian target of rapamycin (mTOR) signaling pathway regulates several essential cellular processes including cell growth, proliferation, autophagy, and protein synthesis.1 In the central nervous system, mTORC1 signaling is crucial since the early stages of neural development, controlling self-renewal and differentiation of neural progenitor stem cells,2, 3, 4 and in neurons, mTORC1-dependent translation is involved in synapse formation and plasticity.5 Therefore, dysfunctional mTORC1 signaling and dysregulated protein synthesis in neuronal cells have been associated with several neurodevelopmental disorders with intellectual disability and autism.6, 7, 8, 9, 10, 11
mTORC1 signaling modulates assembly of the translation initiation machinery by interacting with the eukaryotic initiation factor eIF3, a scaffold complex that brings all components of the translation initiation process into close proximity, and by coordinating the phosphorylation and activity of key translational regulators S6K1 and 4EBP1.12
We have recently identified two novel binding partners for eIF3, the neural Rho-GEF collybistin (CB) and the synaptic scaffold protein gephyrin,13 which are known to be required for GABAA receptors clustering and inhibitory synapse formation.14, 15, 16, 17, 18, 19 Importantly, it has previously been shown that gephyrin associates with mTOR in cortical neurons of rat embryos, and that mTOR signaling regulates gephyrin clustering.20, 21 Taken together, these findings suggest that CB might also interact with mTOR and modulate mTORC1 signaling and protein synthesis. Consequently, dysregulation of mTORC1 activity might contribute to some of the neurological abnormalities observed in patients with CB variants, such as intellectual disability, epilepsy, and anxiety.15, 22, 23, 24, 25 However, to our knowledge, there are no published studies addressing these questions.
In this study, we test these hypotheses using induced pluripotent stem cell (iPSC)-derived neural progenitor cells from a patient with a CB gene (ARHGEF9) deletion and from control individuals and strengthen our findings using a heterologous expression system (HEK293T cells).
Materials and methods
Subject and controls
The K401 patient is an 18-year-old Caucasian boy, the only child of healthy non-consanguineous parents. IQ test could not be taken properly owing to severe mental impairment. Information about the patient's behavior was carried out using the Vineland Adaptive Behavior Scales second-edition (VABS-II), which revealed severely deficient adaptive behavior (VABS-II composite score of 32), and the Childhood Autism Rating Scale (CARS), which revealed mild to moderate autism (CARS score of 34.5). Electroencephalogram showed focal epileptiform discharges, which were maximum over the left frontocentral head region. Patient’s seizures have been under control with medication (valproic acid). No dysmorphic features were observed. After written informed consent was obtained, blood samples were drawn from the K401 patient and from his parents, and skin fibroblast from the patient and from two age- and sex-matched control subjects (called as C1 and C2) were obtained from skin by the punch biopsy. Fragile X syndrome was excluded molecularly and conventional karyotyping analysis showed no alterations. The study was approved by the Human Research Ethics Committees of the Instituto de Biociências – Universidade de São Paulo and of the Instituto Israelita de Ensino e Pesquisa Albert Einstein.
DNA copy number variation assessment
Evaluation of copy number variations (CNVs) in genomic DNA extracted from peripheral blood leukocytes was performed initially using the Agilent's Human Genome CGH Microarray 44 K platform (Agilent Technologies, Santa Clara, CA, USA). Confirmation and refinement of CNVs was then performed using the CytoSNP 850K BeadChip (Illumina, San Diego, CA, USA), which contains 850 000 SNP probes covering the whole genome, with enriched coverage for 3262 well-known genes of constitutional and cancer diseases. Labeling, hybridization, and washing procedures followed the manufacturer’s instructions. Microarray slides were scanned using the iScan System (Illumina), and generated gtc files were loaded on the BlueFuse Multi Software v.3.2 (BlueGnome/Illumina, San Diego, CA, USA) to evaluate experimental quality and to CNV callings. Alterations had to encompass at least five consecutive probes with aberrant log2 values to be called, with a minimum size of 100 kb. The detected alterations were compared to CNVs reported in the Database of Genomic Variants (http://projects.tcag.ca/variation/) and in the DECIPHER database (http://decipher.sanger.ac.uk/). CNVs presenting frequencies >0.1% among the control individuals were disregarded. The genomic structure and gene content of the Xq11.1 microdeletion were interrogated using the UCSC genome browser (GRCh37/hg19; http://genome.ucsc.edu/). Data from this study were deposited in the DECIPHER database under the following accession number: 252484.
Generation of iPSCs, iNPCs, and iNeurons
Control and K401 patient fibroblasts were reprogrammed with retrovirus vectors containing the Oct4, c-Myc, Klf4, and Sox2 human cDNAs obtained from Addgene (Cambridge, MA, USA) as previously described.26 iPSC colonies were propagated under feeder-free growth conditions on Matrigel-coated dishes (BD Biosciences, Franklin Lakes, NJ, USA) using mTeSR1 (StemCell Technologies, Vancouver, BC, Canada). At least two iPSC clones from each individual were generated. Aneuploidies in all selected iPSC clones were excluded using multiplex ligation-dependent probe amplification subtelomeric kits (P070-B2 and P036-E6; MRC-Holland, Amsterdam, the Netherlands). To obtain induced neural progenitor cells (iNPCs), the iPSCs were suspended to generate embryoid bodies and plated to produce neural rosettes, which were manually isolated and cultured onto polyornithine/laminin-coated dishes in NPC medium (DMEM/F12, 0.5 × N2, 1 × B27, 20 ng/ml FGF2 (Invitrogen, Carlsbad, CA, USA), 20 ng/ml EGF (Peprotech, Rocky Hill, NJ, USA)).27 Genome-wide CNV analysis with genomic DNA from K401- and control-derived iNPC clones was performed again (CytoSNP 850K BeadChip platform, Illumina) and only the same deletion affecting the CB gene was identified. For iNeuron differentiation, iNPCs were plated at low density in neural differentiation medium (DMEM/F12, 0.5 × N2, 1 × B27 (Invitrogen), 20 ng/ml GNDF (Peprotech), 200 nm ascorbic acid, and 1 μl retinoic acid (Sigma-Aldrich, St Louis, MO, USA)) onto polyornithine/laminin-coated plates.
To analyze mTORC1 signaling activation in K401 patient and control iNPCs, cells were cultured until 70–80% confluence in NPC medium and were starved in DMEM-F12 containing no supplements during 24 h before protein extraction.
Immunofluorescence
For immunofluorescence staining, cells were grown in eight-well LabTek II chamber slides (Nalgene/Thermo Fisher Scientific, Waltham, MA, USA) and fixed with 4% paraformaldehyde for 10 min. After permeabilization and blocking (5% donkey serum, 0.1% Triton in PBS) for 30 min at room temperature, cells were incubated with primary antibodies diluted in blocking solution overnight at 4 °C. Subsequently, cells were washed with PBS and incubated with secondary antibodies (1:250, Alexa 488/594-coated anti-mouse, anti-rabbit, or anti-goat; Invitrogen) for 1 h at room temperature. Nuclei were stained using DAPI (1:500; Invitrogen). Fluorescence images were taken using an Olympus FSX100 Microscope (Olympus, Waltham, MA, USA). The following primary antibodies were used: anti-Lin28 (1:200, no. 8706), anti-Oct4 (1:50, no. 5177), anti-Nanog (1:50, no. 8750), anti-Sox2 (1:200, no. 5067) (Cell Signaling Technology, Danvers, MA, USA), anti-nestin (1:200, no. MAB5326; Merck Millipore, Billerica, MA, USA), anti-Musashi (1:250, ab52865; Abcam, Cambridge, UK), anti-βIII-tubulin (1:2000, MMS-435P; Covance, Princeton, NJ, USA), anti-MAP2 (1:500, M9942; Sigma-Aldrich), and anti-prpS6S240/244 (1:800, no. 5364; Cell Signaling Technology).
HEK293T cells culture, transfection, and lysis
HEK293T cells were cultured and transfected with either pcDNA-CB (full-length human CB) or empty pcDNA plasmids as described previously.13 To analyze mTORC1 signaling activation, 24 h after transfection cells were starved in DMEM without fetal bovine serum during additional 24 h and were then lysed in CHAPS-containing buffer (40 mM HEPES (pH 7.5), 120 mM NaCl, 1 mM EDTA, 0.3% CHAPS, supplemented with protease and phosphatase inhibitors).
Co-immunoprecipitation and immunobloting
For immunoprecipitation, iNPC lysates (500 μg) and 293T cell lysates (250 μg), prepared in CHAPS-containing buffer, were precleared with protein G-Sepharose (GE Healthcare, Little Chalfont, UK) and then incubated with either the antibody against mTOR protein (1:50 rabbit anti-mTOR antibody, no. 2972; Cell Signaling Technology) or an irrelevant isotype-matched control antibody overnight with rotation at 4 °C. Subsequently, 30 μl of protein G-Sepharose (50% (v/v)) was added and lysates were incubated for at least 6 h at 4 °C on a rotating device. The beads were then washed seven times with lysis buffer and bound proteins were eluted with SDS sample buffer, separated by SDS-PAGE, and transferred to nitrocellulose membranes by western blotting. Expression of either overexpressed CB or endogenous proteins was also analyzed by loading ~10–20 μg of cell lysates into the gel. Membranes were then blocked and incubated with primary antibodies overnight at 4 °C. Detection was performed using HRP-coupled anti-rabbit or anti-mouse secondary antibodies (1:2000; Cell Signaling Technology), ECL substrate (GE Healthcare), and conventional developing using X-ray films (Kodak, Rochester, NY, USA). The same immunoblot procedures were carried out to analyze: (i) the expression of endogenous CB and gephyrin in iPSCs, iNPCs, and iNeurons; (ii) the expression of endogenous PSD95 and GAD65 in iNeurons; and (iii) the mTORC1 signaling activation in 293T cells overexpressing CB and in iNPCs. The following primary antibodies were used: anti-CB (1:5000, no. 261003; Synaptic Systems, Göttingen, Germany), anti-gephyrin (1:200, sc-25311; Santa Cruz Biotechnology, Santa Cruz, CA, USA), anti-mTOR (1:1000, no. 2972), anti-S6K1 (1:1000, no. 9202), anti-eIF3H (1:1000, no. 3413), anti-PSD95 (1:1000, no. 3450), anti-GAD65 (1:1000, no. 5843), anti-pmTORS2448 (1:1000, no. 2971), anti-pS6K1T389 (1:1000, no. 9205), anti-prpS6S240/244 (1:5000, no. 5364) (all from Cell Signaling Technology), and anti-β-actin (1:15 000, A2228; Sigma-Aldrich) antibodies. When necessary, the intensity of the bands was determined by densitometry using NIH ImageJ software (http://rsbweb.nih.gov/ij/), and phosphorylated p-mTOR and p-S6K1 protein levels were quantified and normalized to the corresponding total t-mTOR and t-S6K1 levels, respectively; phosphorylated p-rpS6 protein levels were quantified and normalized to the corresponding β-actin levels (the anti-rpS6 antibody that we have available recognizes the same epitope sequence than anti-phospho-rpS6 antibody and could not be used as loading control).
SUnSet
Analysis of de novo translation was carried out using SUnSET (SUrface SEnsing of translation) technique, which involves the use of the compound puromycin, an aminonucleoside antibiotic, and an anti-puromycin antibody for immunological detection of puromycin-labeled peptides.28 Puromycin is a structural analog of aminoacyl-transfer RNAs, and, as such, can be incorporated into elongating peptide chains through peptide bond formation. However, the binding of puromycin to a nascent polypeptide chain prevents a new peptide bond from being formed and, as a consequence, prevents elongation. When used at very low amounts, which do not inhibit the overall rate of protein synthesis, the rate at which puromycin-labeled peptides are formed reflects the overall rate of mRNA translation. To examine mTORC1-dependent alterations in protein synthesis, iNPCs were starved in DMEM-F12 containing no supplements during 24 h, incubated with rapamycin (250 nm) for 2 h, and subsequently with 10 μg/ml of puromycin (Sigma-Aldrich) for 10 min. Detection of puromycin incorporation was carried out by immunoblotting with anti-puromycin antibody (1:25 000, 12D10; Merck Millipore).
Statistical analysis
For mTORC1 signaling analysis in HEK293T cells, Student's two-tailed t-test was performed using GraphPad Prism Software version 5.0 (GraphPad Prism Software, La Jolla, CA, USA). Differences were considered significant when P-value was <0.05.
Results
Xq11.1 microdeletion containing the entire CB gene in a patient with severe cognitive impairment
We report here a male patient, called as K401, who presented with severe intellectual disability, epilepsy, and mild to moderate autism. Using two array platforms (array-CGH platform (Agilent Technologies) and high-density SNP-based array (Illumina)), we identified in this patient a de novo 216.7 kb microdeletion of Xq11.1 (chrX.hg19:g. 62856174_63072861) including the entire CB gene (ARHGEF9) (Figure 1). This was the only aberration that affects gene function detected in the patient.
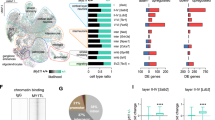
Characterization of Xq11.1 microdeletion encompassing the CB gene in K401 patient. SNP-array (CytoSNP 850K BeadChip platform) result of chromosome X in the K401 patient showing the deletion at Xq11.1. Bellow, enlarged view of the distal Xq11.1 region extracted from the UCSC genome browser (GRCh37/hg19; http://genome.ucsc.edu/) showing its gene content, evolutionary conservation, and the positions of common single-nucleotide polymorphisms (SNPs). The 216.7 kb deletion (segment between chrX.hg19:g.62856174_63072861) detected in the patient K401 including the entire CB gene (ARHGEF9) is represented by red bar (DECIPHER database (http://decipher.sanger.ac.uk/) accession number 252484). EST, expressed sequence tag.
Generation of iPSCs and neural progenitor cells from K401 patient and control individuals
To verify whether CB physically interacts with mTOR in neuronal cells and to begin to test the hypothesis that mTORC1 signaling activity could be altered in patients harboring loss-of-function variants in CB, we sought to generate iPSC-derived neural progenitor cells (hereinafter referred to as iNPCs) from control individuals and from K401 patient. Primary fibroblasts from K401 and from two sex- and age-matched controls were reprogrammed using standard retroviral reprogramming technology.24 We obtained at least two clones from each K401-iPSC and control iPSC, which were subsequently differentiated into iNPCs and iNeurons (for the primary purpose of confirming the differentiation potential of iNPCs into mature neurons). Expression of pluripotency-, NPC-, and neuron-specific markers was determined by immunocytochemistry (Figure 2A). Expression of CB was analyzed by western blotting (Figure 2B) (our commercially available anti-CB antibodies were not suitable for immunocytochemistry studies of the endogenous protein). Interestingly, we observed that CB expression, which is not detected in normal fibroblasts, was reactivated in control iPSCs, iNPCs, and mature iNeurons (Figure 2B). As expected, none of the patient’s cell lineages expressed CB. In addition, similar gephyrin expression levels were observed between patient and control cells. Importantly, we did not detect any major differences between patient and control cells with respect to the reprogramming and differentiation capacities.
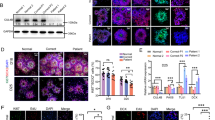
Representative images of iPSCs, iNPCs, and iNeurons, and the expression of lineage-specific markers and CB in these lineages. (A) (a–d) Patient iPSC colonies showing the expression of pluripotent markers: (a) Lin28, (b) Nanog, (c) Sox2, and (d) Oct4. The scale bar represents 100 μm. (e and f) Patient iNPCs showing the expression of neural progenitors markers: (e) Musashi (an RNA-binding protein involved in the regulation of neural stem cell self-renewal) and (f) Nestin (an intermediate filament protein). The scale bar represents 30μm. (g and h) Patient iNeurons showing the expression of neuron-specific cytoskeleton proteins: (g) MAP2 and (h) βIII-tubulin. The scale bar represents 100 μm. Notably, western blot analysis of patient- and control-derived iNeuron lysates showed the expression of PSD95 (a major scaffolding protein localized at the postsynaptic density of excitatory glutamatergic synapses) and GAD65 (an enzyme responsible for GABA synthesis in GABAergic neurons), suggesting the presence of both excitatory and inhibitory synaptic connections (data not shown). (B) Immunoblot analysis of CB and gephyrin expression in K401 patient and control-derived cell lysates. The expression of β-actin was also analyzed as loading control. Cell lysates were prepared from mouse brain tissue (lane 1), control human fibroblasts (lane 2), K401 fibroblasts (lane 3), control iPSCs (lane 4), K401 iPSCs (lane 5), control iNPCs (lane 6), K401 iNPCs (lane 7), control iNeurons (lane 8), and K401 iNeurons (lane 9). CB expression was observed in control iPSCs, iNPCs, and iNeurons (note that CB is expressed at a slightly higher level in iPSCs than in iNPCs and iNeurons). No expression of CB was found in control and K401 fibroblasts, as well as in K401-derived cell lineages (note that one smaller nonspecific band is observed in lysates from control and patient cells). Similar amounts of gephyrin expression were detected between patient and control cells.
CB associates with mTOR in iPSC-derived neural progenitor cells and seem to inhibit the activation of mTORC1 pathway
To examined whether CB and mTOR associate in control iNPCs, we immunoprecipitated endogenous mTOR and assayed for binding of endogenous CB by immunobloting. As depicted in Figure 3a, we found that CB was detected in the mTOR immunoprecipitate. We also observed that, as expected, endogenous eIF3H immunoprecipitates with mTOR (positive control). In addition, although a faint gephyrin band was seen, endogenous gephyrin–mTOR interaction in these cells was also reproduced (Figure 3a).
CB interacts with mTOR in control iNPCs and its absence increases mTORC1 signaling activation and translation initiation in K401 iNPCs. (a) Control iNPCs extracts were incubated with either rabbit anti-mTOR antibody or normal rabbit IgG as a negative control. Gephyrin, CB, and eIF3H (positive control) were detected in the mTOR immunoprecipitate by immunoblotting. The inverse co-immunoprecipitation experiment was not carried out because our commercially available anti-CB antibodies are not suitable for immunoprecipitation. Endogenous expression of t-mTOR, gephyrin, CB, and eIF3H are shown (lysate lane). IP, antibodies used for immunoprecipitation; Blot, antibodies used for immunoblot analysis. (b) K401 iNPCs and control iNPCs (C1 and C2) were starved for 24 h and cell extracts were analyzed for the phosphorylation status of p-mTORS2448, p-S6K1T389, and p-rpS6S240/244 by immunoblotting. The graph shows quantification of phosphorylated proteins relative to their corresponding loading control (t-mTOR, t-S6K1, or β-actin). Error bar represents means ±SD (n=2 control iNPCs). We observed that K401 iNPCs show greater levels of basal p-mTORS2448, p-S6K1T389, and p-rpS6S240/244 compared with controls. For each individual, iNPCs derived from a second iPSC clone were also analyzed for p-mTORS2448, p-S6K1T389, and p-rpS6S240/244 levels with similar results (data not shown). (c) Representative images show double immunofluorescence labeling for p-rpS6S240/244 (red) and Nestin (green) in control and K401 iNPCs, indicating that p-rpS6S240/244 expression came from nestin-positive cells. The scale bar represents 30 μm. (d) Effect of rapamycin on translation in K401 iNPCs and control iNPCs (C1 and C2) measured by SUnSET. The detection of puromycin-labeled peptides was carried out by immunoblotting using anti-puromycin antibody. The graph shows quantification of puromycin signals relative to the loading control β-actin. Error bar represent means±SD (n=2 controls). The results suggest that K401 iNPCs exhibit increased cap-dependent translation. Results in (a–d) are representative of at least three independent experiments.
Because endogenous CB form a complex with mTOR, we next examined whether mTORC1 pathway could be altered in K401-derived iNPCs. To this end, we analyzed by immunoblotting the phosphorylation status of p-mTORS2448 (activated mTORC1 predominantly contains mTOR phosphorylated on Ser2448 site) and its downstream targets p-S6K1T389 and p-rpS6S240/244, in control (n=2) and K401 iNPCs, at similar passage numbers. To reduce individual variations due to cell cycle desynchronization and make potential differences in signaling activation even more evident, cells were starved for 24 h before protein extraction. Under this condition, we observed hyperactivation of mTORC1 signaling pathway in K401 iNPCs (Figure 3b). Double immunofluorescent staining for p-rpS6S240/244 and Nestin (a specific NPC marker) revealed that p-rpS6S240/244 expression came from a homogeneous population of nestin-positive cells (Figure 3c). It is important to mention that when the iNPCs from the K401 patient and from control subjects were maintained in standard supplemented medium without a prior step of growth factors starvation, we did not observe any differences in the phosphorylation levels of mTORC1 cascade components between patient and control cells (data not shown). Therefore, the overactivation of mTORC1 signaling in cultured K401-derived iNPCs is only observed under nutrient-starved (basal) conditions.
Because mTORC1 signaling positively controls protein synthesis, we used SUnSET28 to investigate whether protein synthesis was altered in K401 iNPCs. Patient and control cells (n=2) were starved for 24 h, then incubated with rapamycin for 2 h, and subsequently monitored for mRNA translation by immunoblotting (Figure 3d). The results suggest increased de novo cap-dependent translation in patient cells.
CB also interacts with mTOR in a heterologous expression system and inhibits mTORC1 signaling
To provide additional support and further strengthen our results, we next examined whether CB when overexpressed in HEK293T cells would interact with mTOR and affect endogenous mTORC1 signaling. To this end, 293T cells were transfected with a plasmid expressing full-length human CB, lysed, and mTOR immunoprecipitated complexes were assayed for binding of overexpressed CB by immunoblotting. As depicted in Figure 4a, we found that overexpressed CB interacts with endogenous mTOR in 293T cells. As expected, we also observed that endogenous eIF3H is detected in mTOR immunoprecipitates (positive control).
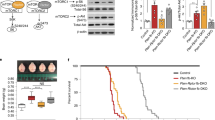
CB interacts with mTOR in 293T cells and inhibits mTORC1 signaling activation. HEK293T cells were transfected with a plasmid expressing full-length human CB (pcDNA-CB) or empty pcDNA vector (pcDNA-Φ) as control. (a) Extracts from cells overexpressing CB were incubated with either anti-mTOR antibody or normal rabbit IgG as a negative control and overexpressed CB was detected in the immunoprecipitated sample by immunoblotting using a CB-specific antibody. The presence of eIF3H in the mTOR immunoprecipitate was analyzed using an anti-eIF3H antibody (positive control). Overexpression of exogenous CB, and endogenous t-mTOR and eIF3H expression in transfected cells is also shown (lysate lane). Results are representative of at least three independent experiments. IP, antibody used for immunoprecipitation; Blot, antibody used for immunoblot analysis. (b) The 293T cells overexpressing CB or empty vector were serum starved for 24 h and cell extracts were analyzed for the phosphorylation status of p-mTORS2448, p-S6K1T389, and p-rpS6S240/244 by immunoblotting. Overexpression of exogenous CB in cells transfected with pcDNA-CB was confirmed by immunoblotting. Lane CB, extracts from 293T cells transfected with pcDNA-CB; Lane Φ, extracts of 293T cells transfected with pcDNA-Φ. The graph shows quantification of phosphorylated proteins relative to corresponding loading control (t-mTOR, t-S6K1, or β-actin). Error bars represent means±SD (n=3 independent experiments); *P<0.05. Cells overexpressing CB show diminished levels of basal p-mTORS2448, p-S6K1T389, and p-rpS6S240/244 compared with control.
Next, 293T cells overexpressing either CB or control empty vector were starved for 24 h and total cell lysates were analyzed for p-mTORS2448, p-S6K1T389, and p-rpS6S240/244 levels by immunoblotting. As illustrated in Figure 4b, we observed that the phosphorylation status of these proteins were significantly reduced in cells overexpressing CB compared with control cells. This result corroborates our previous findings suggesting that CB seems to inhibit basal mTORC1 signaling.
Discussion
The previously established CB/gephyrin–eIF3 and gephyrin–mTOR interactions led us to hypothesize that CB also associates with mTOR and is involved in mTORC1-mediated protein translation, and that dysfunctional signaling by mTORC1 may also contribute to the neurological abnormalities observed in patients with CB variants affecting gene expression/function. To test these hypotheses, we used two model systems: iPSC-derived NPCs from a patient with a CB gene deletion (patient K401) and from normal subjects, and heterologous expression in 293T cells.
Herein, we provide evidence that endogenous CB and mTOR form a complex in control iNPCs. We also show that in K401-derived iNPCs, the CB absence seems to increase basal mTORC1 pathway function in these neuronal cells compared with control iNPCs, suggesting that CB may inhibit mTORC1 signaling activity. Consistent with the enhanced mTORC1 signaling in K401-derived iNPCs, our results suggest exaggerated translation in these cells. Finally, these findings were strengthened by overexpression studies in 293T cells, where ectopically expressed CB interacts with endogenous mTOR and inhibits basal mTORC1 signaling.
On the basis of our results, we suggest a model whereby CB forms a complex with mTOR and eIF3 and by sequestering these proteins downregulates mTORC1 signaling and protein synthesis. The molecular details of these interactions have yet to be defined. As CB is known to promote plasma membrane attachment of gephyrin complexes by binding membrane lipids,14, 15, 16 it would be interesting to analyze whether CB–mTOR complexes are also found in the plasma membrane of neuronal cells. In addition, it is possible that CB–gephyrin–mTOR can form functional ternary complexes and that these complexes are involved in mTORC1-mediated protein synthesis regulation.
Loss-of-function variants in negative regulators of mTORC1 signaling are the cause of several monogenic syndromes associated with intellectual disability and autism spectrum phenotypes, such as Fragile X syndrome,9, 29 tuberous sclerosis,30 and PTEN-related syndromes.31 In addition, overactivated mTORC1 signaling has also been linked to the pathophysiology of non-syndromic ASD.10, 11, 32, 33 This hyperactivation of the mTORC1 pathway has been shown to stimulate excessive protein synthesis in neuronal cells, leading to disturbances in neuronal differentiation and morphology, synaptic connectivity, and plasticity. Loss-of-function variants in CB, which are known to reduce GABAergic transmission and alter synaptic plasticity, have been associated with overlapping phenotypes, that is, intellectual disability, epilepsy, anxiety,15, 22, 23, 24, 25 and now autism, present in the patient here described. Therefore, the data presented here point to the possibility that mTORC1 pathway overactivation in CB-deficient neuronal cells might also contribute to the cognitive and behavioral abnormalities in patients with loss-of-function variants in CB. Although further experiments and larger samples of patients and controls are required to fully verify this hypothesis, our findings are encouraging and should stimulate more research along similar lines.
Elucidation of the signaling network regulating protein synthesis in neuronal cells is essential for understanding the pathological process of several neurological disorders and to develop targeted therapies, and further studies on the involvement of CB in translation control will likely prove highly fruitful.
References
Laplante M, Sabatini DM : mTOR signaling at a glance. J Cell Sci 2009; 122: 3589–3594.
Han J, Wang B, Xiao Z et al: Mammalian target of rapamycin (mTOR) is involved in the neuronal differentiation of neural progenitors induced by insulin. Mol Cell Neurosci 2008; 39: 118–124.
Han J, Xiao Z, Chen L et al: Maintenance of the self-renewal properties of neural progenitor cells cultured in three-dimensional collagen scaffolds by the REDD1-mTOR signal pathway. Biomaterials 2013; 34: 1921–1928.
Magri L, Cambiaghi M, Cominelli M et al: Sustained activation of mTOR pathway in embryonic neural stem cells leads to development of tuberous sclerosis complex-associated lesions. Cell Stem Cell 2011; 9: 447–462.
Jaworski J, Sheng M : The growing role of mTOR in neuronal development and plasticity. Mol Neurobiol 2006; 34: 205–219.
Costa-Mattioli M, Monteggia LM : mTOR complexes in neurodevelopmental and neuropsychiatric disorders. Nat Neurosci 2013; 16: 1537–1543.
Kelleher 3rd RJ, Bear MF : The autistic neuron: troubled translation? Cell 2008; 135: 401–406.
Auerbach BD, Osterweil EK, Bear MF : Mutations causing syndromic autism define an axis of synaptic pathophysiology. Nature 2011; 480: 63–68.
Hoeffer CA, Sanchez E, Hagerman RJ et al: Altered mTOR signaling and enhanced CYFIP2 expression levels in subjects with fragile X syndrome. Genes Brain Behav 2012; 11: 332–341.
Tang G, Gudsnuk K, Kuo SH et al: Loss of mTOR-dependent macroautophagy causes autistic-like synaptic pruning deficits. Neuron 2014; 83: 1131–1143.
Oguro-Ando A, Rosensweig C, Herman E et al: Increased CYFIP1 dosage alters cellular and dendritic morphology and dysregulates mTOR. Mol Psychiatry 2015; 20: 1069–1078.
Holz MK, Ballif BA, Gygi SP, Blenis J : mTOR and S6K1 mediate assembly of the translation preinitiation complex through dynamic protein interchange and ordered phosphorylation events. Cell 2005; 123: 569–580.
Sertie AL, de Alencastro G, De Paula VJ, Passos-Bueno MR : Collybistin and gephyrin are novel components of the eukaryotic translation initiation factor 3 complex. BMC Res Notes 2010; 3: 242.
Kins S, Betz H, Kirsch J : Collybistin, a newly identified brain-specific GEF, induces submembrane clustering of gephyrin. Nat Neurosci 2000; 3: 22–29.
Harvey K, Duguid IC, Alldred MJ et al: The GDP-GTP exchange factor collybistin: an essential determinant of neuronal gephyrin clustering. J Neurosci 2004; 24: 5816–5826.
Poulopoulos A, Aramuni G, Meyer G et al: Neuroligin 2 drives postsynaptic assembly at perisomatic inhibitory synapses through gephyrin and collybistin. Neuron 2009; 63: 628–642.
Saiepour L, Fuchs C, Patrizi A, Sassoè-Pognetto M, Harvey RJ, Harvey K : Complex role of collybistin and gephyrin in GABAA receptor clustering. J Biol Chem 2010; 285: 29623–29631.
Tyagarajan SK, Ghosh H, Harvey K, Fritschy JM : Collybistin splice variants differentially interact with gephyrin and Cdc42 to regulate gephyrin clustering at GABAergic synapses. J Cell Sci 2011; 124: 2786–2796.
Hoon M, Soykan T, Falkenburger B et al: Neuroligin-4 is localized to glycinergic postsynapses and regulates inhibition in the retina. Proc Natl Acad Sci USA 2013; 108: 3053–3058.
Sabatini DM, Barrow RK, Blackshaw S et al: Interaction of RAFT1 with gephyrin required for rapamycin-sensitive signaling. Science 1999; 284: 1161–1164.
Wuchter J, Beuter S, Treindl F et al: A comprehensive small interfering RNA screen identifies signaling pathways required for gephyrin clustering. J Neurosci 2012; 32: 14821–14834.
Marco EJ, Abidi FE, Bristow J et al: ARHGEF9 disruption in a female patient is associated with X linked mental retardation and sensory hyperarousal. J Med Genet 2008; 45: 100–105.
Kalscheuer VM, Musante L, Fang C et al: A balanced chromosomal translocation disrupting ARHGEF9 is associated with epilepsy, anxiety, aggression, and mental retardation. Hum Mutat 2009; 30: 61–68.
Shimojima K, Sugawara M, Shichiji M et al: Loss-of-function mutation of collybistin is responsible for X-linked mental retardation associated with epilepsy. J Hum Genet 2011; 56: 561–565.
Lesca G, Till M, Labalme A et al: De novo Xq11.11 microdeletion including ARHGEF9 in a boy with mental retardation, epilepsy, macrosomia, and dysmorphic features. Am J Med Genet A 2010; 155A: 1706–1711.
Takahashi K, Tanabe K, Ohnuki M et al: Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 2007; 131: 861–872.
Marchetto MC, Carromeu C, Acab A et al: A model for neural development and treatment of Rett syndrome using human induced pluripotent stem cells. Cell 2010; 143: 527–539.
Schmidt EK, Clavarino G, Ceppi M, Pierre P : SUnSET, a nonradioactive method to monitor protein synthesis. Nat Methods 2009; 6: 275–277.
Sharma A, Hoeffer CA, Takayasu Y et al: Dysregulation of mTOR signaling in fragile X syndrome. J Neurosci 2010; 30: 694–702.
Meikle L, Talos DM, Onda H et al: A mouse model of tuberous sclerosis: neuronal loss of Tsc1 causes dysplastic and ectopic neurons, reduced myelination, seizure activity, and limited survival. J Neurosci 2007; 27: 5546–5558.
Zhou J, Blundell J, Ogawa S et al: Pharmacological inhibition of mTORC1 suppresses anatomical, cellular, and behavioral abnormalities in neural-specific Pten knock-out mice. J Neurosci 2009; 29: 1773–1783.
Ebert DH, Greenberg ME : Activity-dependent neuronal signalling and autism spectrum disorder. Nature 2013; 493: 327–337.
Suzuki AM, Griesi-Oliveira K, de Oliveira Freitas Machado C et al: Altered mTORC1 signaling in multipotent stem cells from nearly 25% of patients with nonsyndromic autism spectrum disorders. Mol Psychiatry 2015; 20: 551–552.
Acknowledgements
We are grateful to all of the subjects who participated in this work. We thank Elaine Cristina Zachi for performing formal psychological evaluation of the patient. We thank Marta Diniz and Andressa Morales for technical assistance. We thank Soely Moura, Aline Moraes, and Constância G Urbani for secretarial assistance. This work was supported by grants from CNPq (National Counsel of Technological and Scientific Development), FAPESP (São Paulo Research Foundation), CAPES (Higher Education Co-Ordination Agency), and Autismo and Realidade.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Machado, C., Griesi-Oliveira, K., Rosenberg, C. et al. Collybistin binds and inhibits mTORC1 signaling: a potential novel mechanism contributing to intellectual disability and autism. Eur J Hum Genet 24, 59–65 (2016). https://doi.org/10.1038/ejhg.2015.69
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2015.69
This article is cited by
-
Human ARHGEF9 intellectual disability syndrome is phenocopied by a mutation that disrupts collybistin binding to the GABAA receptor α2 subunit
Molecular Psychiatry (2022)
-
Sex differences in the effects of prenatal bisphenol A exposure on autism-related genes and their relationships with the hippocampus functions
Scientific Reports (2021)
-
The role of GABAergic signalling in neurodevelopmental disorders
Nature Reviews Neuroscience (2021)
-
De novo ARHGEF9 missense variants associated with neurodevelopmental disorder in females: expanding the genotypic and phenotypic spectrum of ARHGEF9 disease in females
neurogenetics (2021)
-
Copy number variants (CNVs): a powerful tool for iPSC-based modelling of ASD
Molecular Autism (2020)