Abstract
Potassium is one of the major macro-nutrients essential for a number of cellular processes in plants. Low potassium level in the soil represents a limiting factor for crop production. Recent studies have identified potassium transporters that are involved in potassium acquisition, and some of them are critical for potassium nutrition under low potassium conditions. However, little is understood on the molecular components involved in low potassium signaling and responses. We report here the identification of a calcineurin B-like protein-interacting protein kinase (CIPK9) as a critical regulator of low potassium response in Arabidopsis. The CIPK9 gene was responsive to abiotic stress conditions, and its transcript was inducible in both roots and shoots by potassium deprivation. Disruption of CIPK9 function rendered the mutant plants hypersensitive to low potassium media. Further analysis indicated that K+ uptake and content were not affected in the mutant plants, implying CIPK9 in the regulation of potassium utilization or sensing processes.
Similar content being viewed by others
Introduction
Plant growth requires a number of mineral nutrients, and potassium (K+) is the most abundant inorganic cation in plants constituting up to 5-10% of a plant's dry weight 1. Potassium performs vital functions in metabolism, growth, and stress adaptation. On the one hand, potassium functions in the cell by directly interacting with proteins resulting in enzyme activation, stabilization of protein synthesis, and neutralization of negative charges on proteins 2, 3, 4. In the second category of functions, K+ is a major driving force for osmotic regulation, for example, in stomatal movement, light-driven and seismonastic movements of organs or phloem transport 3, 5. In cell expansion and growth, accumulation of K+ (together with anions) in plant vacuoles creates the necessary osmotic potential for rapid cell enlargement. K+ movement also provides a charge separation across the membrane that is essential for the movement of other ions. In chloroplasts, energy production through H+-ATPases depends on overall H+/K+ exchange 6, 7.
K+ deficiency is of great agricultural importance 8. This fact was recognized early in plant physiological research that led to an extensive description of K+ starvation symptoms at the physiological level 3. It is well established that K+ starvation leads to growth arrest owing to the lack of the major osmoticum, impaired nitrogen balance owing to inhibition of protein synthesis, reduced levels of sugars owing to inhibition of photosynthesis, and impaired long-distance transport 3. Low potassium content in the soil therefore severely limits crop yield, and farming practice often requires a large amount of organic and/or chemical fertilizers to overcome this condition. Use of fertilizers imposes undesirable economical and environmental problems that can be solved by genetic engineering of low-K+ tolerant crops based on the molecular understanding of K+ nutrition in plants. One important aspect of plant adaptation to low K+ stress is cellular and tissue homeostasis of K+, which involves transport of K+ across various membranes in various tissues 9. K+ transport mechanisms have been studied extensively at the molecular level and many of the transporters have been cloned and functionally analyzed 10, 11, 12, 13. There are typically two mechanisms for K+ acquisition, one is high-affinity and the other is low-affinity uptake from the roots 2, 14, 15, 16. Some of the transporters such as the voltage-gated channels are generally considered as low-affinity transporters although studies also show that these channels can perform high-affinity uptake 17, 18, 19, 20, 21. In addition, a single transporter can mediate both high- and low-affinity transport 20, 22, 23. Most of the soil solutions contain less than 1 mM K+, thus the high-affinity uptake plays a critical role in potassium nutrition in plants 13, 24, 25. Under potassium deficiency, high-affinity uptake is even more important for plant survival. Studies have shown that such high-affinity uptake may be inducible and tightly regulated under low potassium conditions 18, 20, 21. However, it is unknown how plants detect or sense external K+ concentrations and what are the signaling mechanisms that integrate physiological, biochemical, and molecular responses into a concerted adaptive response.
A recent study 26 identifies reactive oxygen species (ROS) as a critical signaling molecule for plant response to potassium-deficiency conditions. Interestingly, ROS in root cells has been shown to elicit changes in cellular calcium and is required for root hair growth and mineral uptake 27. Together, these studies suggest that ROS signal is produced under low potassium conditions and ROS-induced calcium changes may be a crucial messenger for downstream responses. However, it is not known how low-K+-induced calcium signal is perceived by the cell. In addition to low-K+ response, calcium plays multiple roles in many other cellular pathways, including signaling processes in response to pathogen, abiotic stresses, and developmental signals 28, 29, 30, 31, 32, 33. Calcium is also involved in the regulation of K+/Na+ homeostasis and ionic selectivity under saline conditions and exogenous Ca2+ can improve salt tolerance 34, 35, 36, 37. As a second messenger, Ca2+ transmits the primary signal into cellular responses through Ca2+-regulated proteins that include Ca2+sensors and their targets. The calcineurin B-like proteins (CBLs) are a family of recently identified calcium sensors unique to plants 38, 39, 40. CBLs specifically target a family of protein kinases referred to as calcineurin B-like protein-interacting protein kinases (CIPKs) (CBL-interacting protein kinases) 38, 39, 40. In Arabidopsis, both CBLs and CIPKs are encoded by large multi-gene families; 10 members in CBL and 25 members in CIPK gene family provide a high level of diversity and complexity in the function of the CBL-CIPK network. The CBL-CIPK network has been shown to play an essential role in several physiological processes, including abiotic stress responses 37, 41, 42, 43, 44, 45 and response to phytohormone abscisic acid (ABA) 43, 45. Recently, one of the members of CIPK gene family, CIPK23, was shown to be involved in regulation of K-nutrition by regulating the voltage-gated channel AKT1 in roots of Arabidopsis 46, 47. In this study we have identified another member of the CIPK gene family, CIPK9, as a critical regulator of low potassium response in Arabidopsis by genetic analysis, providing a component that functions in between the low-K+-induced calcium signal and downstream responses.
Materials and Methods
Plant materials, stress treatments, and RNA analysis
Arabidopsis thaliana plants (ecotype Columbia-0) were grown in the greenhouse for seed production as described previously 48. For RNA analysis, 2-week-old seedlings grown on MS medium 49 were treated under different stress conditions. Seeds were surface sterilized as described previously 48. For NaCl and mannitol treatment, solution was overlayed onto the 2-week-old seedlings grown on MS plates to ensure total coverage of the foliage area. Seedlings were incubated at room temperature under white light. In parallel experiments, water was overlayed as control. To perform wounding treatments, 2-week-old seedlings grown in MS medium were mechanically injured as described previously 43. For cold treatment 2-week-old seedlings were exposed to 4 °C, and whole seedlings were harvested after a defined period of time. Total RNA was isolated with Tripure isolation reagent (Roche Diagnostics, Indianapolis, IN, USA). All experiments were repeated at least three times, and results from one representative experiment are shown.
For low-K+-treatment, 10-day-old Arabidopsis seedlings grown on ½ MS medium were transferred to modified MS (10 mM K+) or the K+-deficient (0 mM K+) liquid medium for different lengths of times as indicated in Figure 2C and 2D.
Induction of CIPK9 transcripts by stress and low-K+ conditions. (A) RT-PCR analysis of CIPK9 mRNA levels under various conditions. 14-day-old seedlings grown on MS medium were treated with cold (4 °C), NaCl (300 mM), mannitol (400 mM), and wounding (mechanical injury). Semi-quantitative RT-PCR analyses (25 cycles) were performed with CIPK9-specific primers. (B) Differential expression of CIPK9 transcript at different developmental stages. Wild-type Arabidopsis seedlings were grown on MS-agar plate at standard long-day growth conditions. Total RNA was isolated from roots and shoots harvested at different developmental stages (7, 14 and 60 days). Semi-quantitative RT-PCR (35 cycles) was conducted by using CIPK9-specific primers. (C) Induction of CIPK9 mRNA in roots and shoots under potassium-deficient conditions. Wild-type Arabidopsis seedlings were grown on ½ MS-agar media for 10 days before transfer to modified MS with K+(10 mM K+) or without K+ (0 mM K+) for 48 h. Total RNA was isolated from roots and shoots and semi-quantitative RT-PCR (28 cycles) was performed. (D) Time course of CIPK9 expression under potassium-deficient condition. 10-day-old Arabidopsis seedlings grown on ½ MS medium were transferred to liquid-modified MS media containing 10 or 0 mM K+. Samples were collected at 0, 24, 48, 96, and 120 h after transfer and total RNA was extracted from whole seedlings for semi-quantitative RT-PCR analysis using CIPK9-specific primers. Actin-2 was used as a loading control in (A-D).
Reverse transcriptase-PCR analysis of gene expression
To examine the expression of CIPK9, reverse transcriptase-mediated PCR (RT-PCR) was performed as described previously 48 using full open reading frame of CIPK9 (At1g01140) forward (5′-AAA TCTAGA ATG AGT GGA AGC AGA AGG AAG GCG ACG CC-3′) and reverse (5′-AA AGGTCC TTA TTG CTT TTG TTC TTC AGC GGC TGC ATC-3′) primers to amplify a PCR product of 1.344 kb. Expression levels of Actin-2 was monitored with forward (5′-GGA AAG GAT CTG TAC GGT AAC-3′) and reverse (5′-TGT GAA CGA TTC CTG GAC-3′) primers to serve as a quantifying control. Aliquots of individual PCR products were resolved by agarose gel electrophoresis and visualized with ethidium bromide by Gel Doc 1000 (Bio-Rad, Hercules, CA, USA).
Isolation of T-DNA insertion mutants
The cipk9 mutants for At1g01140 were isolated from the T-DNA insertion collections of Torrey Mesa Research Institute (cipk9-1 is SAIL_252_F06) and SALK (http://signal.salk.edu/; cipk9-2 is SALK_058629). The T-DNA borders of cipk9-1 and cipk9-2 alleles were defined by sequencing PCR products obtained using a T-DNA border primer and a gene-specific primer. SAILLB3 (5′-TAG CAT CTG AAT TTC ATA ACC AAT CTC GAT ACA C-3′) and SALKLB1 (5′-GCA AAC CAG CGT GGA CCG CTT GCT GCA ACT-3′) were used as T-DNA border primers for the mutant obtained from the Torrey Mesa Research Institute and SALK, respectively. The T-DNA insertion in the mutants (cipk9-1 and cipk9-2) was confirmed by PCR and DNA gel blot analysis, and its exact position was determined by sequencing. After selfing of heterozygous plants, homozygous cipk9 mutants were identified by genomic PCR to confirm disruption of the endogenous gene and by RT-PCR to confirm disruption of gene expression. The expression of Actin-2 was monitored with forward (5′-GGA AAG GAT CTG TAC GGT AAC-3′) and reverse (5′-TGT GAA CGA TTC CTG GAC-3′) primers to serve as a quantifying control.
Analysis of CIPK9 promoter-GUS expression in transgenic plants
To generate the CIPK9 promoter-GUS construct, the 5′ flanking DNA of the CIPK9 coding region was amplified with forward (5′-AAA GTCGAC ATA TAT TCA CAA TCA TCG AAT ACA AGA ACA CC-3′) and reverse (5′-AAA GAATTC TTT CTT TTT CCG ATT AAG AAA ATC AAC G-3′) primers. The 2.58 kb of PCR fragment was cloned into SalI and EcoRI sites (underlined in the primer sequences) in the pBI101.1 vector (Clontech, Palo Alto, CA, USA). The construct was transformed into Arabidopsis plants 50 and transformants were selected on 50 μg/ml kanamycin. Several independent T1 transgenic lines were stained with 5-bromo-4-chloro-3-indolyl-D-glucuronide for 12 h followed by incubation in 80% ethanol to remove chlorophyll 51.
Growth measurement on media with variable potassium levels and different nutrient strengths
For growth measurements of wild-type and cipk9 mutants on low potassium nutrition, seeds were plated to a modified MS medium containing various concentrations of KCl 36. Modified MS medium contains potassium-free 1/20 strength MS major salts and 1× MS minor salts. Potassium-free medium was prepared by replacing MS salts with the following (1× MS): 1650 mg/l NH4NO3, 440 mg/l CaCl2·2H2O, 370 mg/l MgSO4·7H2O, 165 mg/l (NH4)2HPO4, 27.8 mg/l FeSO4·7 H2O, 37.3 mg/l di-sodium EDTA, 0.7495 mg/l NaI, 6.3 mg/l H3BO3, 16.9 mg/l MnSO4·H2O, 8.6 mg/l ZnSO4·7H2O, 0.25 mg/l Na2MO4·2H2O, 0.016 mg/l CuSO4·5H2O, and 0.0267 mg/l CoSO4·6H2O. For solidification of MS media, 1% ultra-pure agarose (Invitrogen, San Diego, CA, USA) was used instead of agar. Varying levels of potassium in the media were achieved by adding appropriate amounts of KCl to the K+-free medium. We have measured the K+ content in K+-free buffer and in agarose and found negligible levels in all buffers; however, 1% agarose media contain approximately 0.003 mM K+. For different nutrient strengths of MS media, 1× MS media was diluted to 1/20 to 1/10 000 times with or without addition of KCl.
86Rb+ uptake assay
For measurement of potassium uptake using 86Rb+ as a tracer, we performed experiments as described 52 with some modifications. Seven-day-old seedlings from MS liquid cultures were transferred to K+-free medium for 2 days. Seedlings were collected, rinsed briefly in K+-free medium, and then placed into a 10-ml uptake solution containing K+-free medium supplemented with indicated levels of KCl and 0.5 μCi/ml of 86Rb+ (Amersham). The uptake was performed at room temperature under white fluorescent light. Uptake at different K+ concentrations was performed for 1 h. The time course of uptake at 0.02 or 20 mM K+ was performed for 30, 60, 90, and 120 min. At the completion of uptake, the seedlings were rinsed three times with 50 ml of ice-cold K+-free media. The seedlings were blotted dry on filter paper and weighed, and the radioactivity was measured using a Beckman LS6500 scintillation counter (Beckman Instruments, Fullerton, CA, USA). Values of K+ uptake were obtained based on 86Rb+ uptake and the external [K+].
Determination of K+ content
For the measurements of K+ content in plant tissues, the 7-day-old seedlings of various mutant and wild-type Arabidopsis were transferred from MS medium to the low potassium medium and treated for 4 days. The seedlings were then collected, rinsed 4-5 times with distilled water, and dried at 70 °C for 24 h and weighed. The samples were digested with 0.1 N HNO3, and the K+ concentrations were determined with an atomic absorption spectrophotometer (model 560; Perkin-Elmer, Norwalk, CT, USA).
Emergence of cotyledons and root elongation assays
Approximately 100 seeds each from the wild type (Col-0), cipk9 mutant alleles (cipk9-1 and cipk9-2) and 35S::CIPK9 transgenic lines were planted in triplicate on modified MS-agarose medium with or without different concentrations of K+or different nutrient strengths of MS media, and incubated at 4 °C for 6 days before being placed at 23 °C under long-day conditions. Emergence of cotyledons was scored daily for 15 days. The vertical root growth assays shown in Figures 4 and 5 were performed in a similar manner except that the plates were placed vertically on a rack and the root length was measured and recorded. Relative root growth and relative emergence of cotyledons were determined as the ratio of root growth or emergence of cotyledons for respective mutant allele to wild type at a particular concentration.
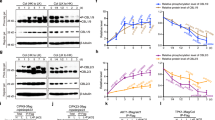
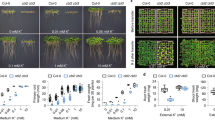
Hypersensitive growth of cipk9 mutants under low potassium media. (A) Seedlings were plated on vertical MS-agarose plates (½ MS; 10 mM K+), or modified MS (see Materials and Methods) with 0, 0.001, 0.005, 0.01, 0.02, 0.05, 0.100, or 10 mM K+ for 10 days before taking the photographs. (B) Measurement of root growth under various K+ levels. Root length of 7-day-old seedlings was measured (n = 95 for each condition). Three independent experiments were performed and values are means ± SD. (C) Cotyledon emergence under various K+ levels. Emergence of cotyledons was scored after 9 days of growth on the modified MS media containing various levels of K+(n = 100 for each condition). Three independent experiments were performed and values are means ± SD. Student's t test was performed to determine the significance, in (B) and (C) *p < 0.0001, **p = 0.07, and ***p> 0.5.
Phenotypic analysis of cipk9 mutants grown in diluted MS media. (A) Seedlings of Col-0, cipk9-1, and cipk9-2 were grown vertically in different strengths of MS-agarose media (1/20, 1/100, 1/1000, 1/5 000 and 1/10 000 dilutions) with or without supplementing K+. Photographs were recorded after 12 days of growth at standard growth conditions. (B) Measurement of root growth under different strengths of MS media. Root length of 12-day-old seedlings was measured (n = 25 for each condition). Three independent experiments were performed and values are means ± SD. (C) Measurement of cotyledon emergence under different strengths of MS media. Emergence of cotyledons was scored after 12 days of growth on the different strengths of MS media (n = 100 for each condition). Three independent experiments were performed and values are means ± SD. Student's t test was performed to determine the significance, in (B) and (C), *p < 0.0003; **p > 0.15; and ***denotes p > 0.60.
Results
CIPK9 gene is inducible by abiotic stresses and potassium deficiency
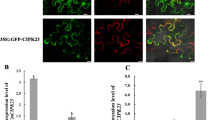
As a step towards dissecting the function of each CIPK in calcium signaling in plants, the expression patterns of these genes were surveyed by RNA blot analyses. We noticed that the expression levels of one CIPK family member, CIPK9, were hardly detectable in adult plants by RNA gel blot analysis (data not shown). Therefore, we determined CIPK9 mRNA levels using RT-mediated PCR. CIPK9 mRNA was detected in all the tissues of the Arabidopsis plant by RT-PCR (Figure 1A). To analyze the expression of the CIPK9 gene in more detail, we fused the putative promoter region of the CIPK9 gene to the β-glucuronidase (GUS) reporter and analyzed GUS activity in transgenic plants by a histochemical procedure. As shown in Figure 1B, CIPK9 promoter was highly active in the germinating seeds (1 day after sowing, mostly in the upper part of the radicle). A strong GUS activity was detected in the mature region of the roots, including the root hairs. However, no GUS activity was detected in the root tip and much of the elongation zone. GUS activity was also detected in most of the tissues of young rosette plants, consistent with the RT-PCR results shown in Figure 1A. A typical rosette leaf exhibited low detectable GUS activity whereas anthers and stigma in the flower were stained strongly. Significant GUS activity was also detected in the petals, sepals, and siliques (Figure 1B).
Expression patterns of CIPK9 gene. (A) RT-PCR analysis of CIPK9 transcripts in different organs of Arabidopsis plants and during seed germination. Total RNA was isolated from various tissues (root, rosette leaf, cauline leaf, stem, flower bud, opened flower, and silique) of wild-type plants (ecotype Col-0) grown under long-day conditions or from germinating seeds (2 days after sowing). Thirty-five cycles of RT-PCR were performed with CIPK9-specific primers (top gel) or Actin-2-specific primers (bottom gel). (B) Histochemical GUS analysis of CIPK9 promoter-GUS transgenic plants. The top panel depicts from left to right a germinating seed (1), a seedling of 2 (2) and 5 (3) days after germination, and a 2-week-old plant (4). The bottom row shows root elongation and mature zone (5), root tip (6), a leaf of rossette stage plant (7), opened flower (8), and a silique (9).
As adult plants expressed CIPK9 at low levels, we suspected that CIPK9 expression may be regulated by environmental factors. Indeed, stress signals (including cold, hyper-osmotic stress, and wounding) strongly induced the expression of the CIPK9 gene (Figure 2A). Induction by osmotic stress (mannitol) was the strongest, followed by high salt, cold, and wounding. The induction of CIPK9 transcript by abiotic stress conditions was transient as shown for the induction of many other signaling components such as CBL1, CBL9, and CIPK3 in the CBL-CIPK network 42, 43, 45. For example, in the cold treatment, the CIPK9 transcript increased after 3 h and reached a maximum at 6 h followed by a gradual decline at 12 and 24 h. In NaCl treatment, CIPK9 mRNA levels rapidly increased after 1 h and reached a maximum at 3 h followed by a decline. Under mannitol treatment, CIPK9 transcript levels also increased rapidly at 1 h and retained a maximum at 6 and 12 h and were reduced at the 24 h time point. CIPK9 was also expressed at different levels in the roots and shoots at different developmental stages of plant growth (Figure 2B).
Interestingly, CIPK9 mRNA levels were upregulated under potassium-deficient growth conditions (Figure 2C and 2D). Such induction was observed in both roots and shoots although a stronger induction was observed in the shoots (Figure 2C). The mRNA levels of CIPK9 increased significantly, as the low potassium treatment continued for more than 2 days and further increased within the 5-day period (Figure 2D). This result was consistent with the data in the TAIR (The Arabidopsis Information Resource) micro-array database 53. Although induction by abiotic stress conditions is a common property of several genes in the CBL and CIPK families, strong induction by low-potassium condition is rather unique to CIPK9 and may indicate a unique function of CIPK9 in low potassium responses.
The cipk9 mutant was specifically hypersensitive to low potassium conditions
To examine the function of CIPK9 gene, we have isolated two T-DNA insertion alleles of this gene, cipk9-1 and cipk9-2, from the collections of T-DNA-transformed Arabidopsis lines (SAIL252_F06 and SALK_058629, ABRC). Homozygous mutant lines were established after selfing. Sequence analysis indicated that the insertion site in the cipk9-1 allele is located in the 5′ UTR (un-translated region) approximately at 100 bp upstream of ATG and the T-DNA in the cipk9-2 allele is inserted in the fifth exon following the 1211th bp after ATG (Figure 3A). RT-PCR analysis showed that both insertions abolished the expression of CIPK9 in the mutant plants (Figure 3B).
Isolation of the CIPK9 T-DNA insertion mutants. (A) Intron-exon organization of the Arabidopsis CIPK9 gene and T-DNA locations. Solid black boxes indicate un-translated regions (UTRs), and gray shaded boxes and lines indicate coding regions and introns, respectively. The positions of the T-DNA insertion in two alleles, cipk9-1 and cipk9-2, are indicated by arrowhead (not to scale). (B) Thirty-five cycles of RT-PCR analysis of CIPK9 transcripts in wild-type (Col-0) and mutant alleles (cipk9-1 and cipk9-2). Expression of Actin-2 was analyzed as a loading control.
We did not observe any significant phenotypic changes in cipk9 mutant plants compared with wild-type plants under normal growth conditions in the growth chamber or greenhouse. As CIPK9 is highly inducible under some abiotic stress conditions and was responsive to potassium deficiency, we speculated that it may function under these stress conditions. We performed phenotypic analysis of both seedlings and adult plants under different abiotic stress conditions, such as cold, salinity, osmotic stress, and in the presence of several plant hormones including ABA, auxin, cytokinin, and GA. No significant changes were scored between the mutants and the wild-type plants under these conditions (data not shown).
When we grew the seedlings on the modified MS media with low potassium levels (in the micro-molar range), the growth of cipk9 mutant plants was significantly more inhibited as compared to the wild type. As shown in Figure 4, both roots and shoots in the mutants were more sensitive to the low potassium condition than the wild type. Such hypersensitivity was consistently observed in both the cipk9-1 and cipk9-2 alleles. The growth of wild-type plants (Col-0) was also reduced under low potassium media, but the growth of the mutant alleles was clearly hypersensitive to low potassium conditions. In 10 mM KCl (equivalent to the K+ level of half-strength MS salts), cipk9 mutant seedlings were indistinguishable from the wild type. Results of more detailed analyses of root growth under different concentrations of K+ are depicted in Figure 4B. The root growth in low potassium media (from 0 to 0.05 mM K+) is strongly inhibited in both wild type and mutant. However, the cipk9 mutant seedlings were significantly more inhibited by the low potassium level. When the K+ concentration in the MS media is equal to or higher than 0.1 mM, root growth of wild type and cipk9 mutants was not significantly different. At 0.02 mM K+, the root length of both cipk9 mutant alleles was about 50% of wild type seedlings. Similarly, cotyledon emergence and expansion were also strongly inhibited in the low potassium media and cipk9 mutants again displayed hypersensitivity (Figure 4C). At 0.01 mM K+, 82 % wild-type seedlings expanded cotyledons whereas only 34% and 31% of mutant cipk9 alleles (cipk9-1 and cipk9-2, respectively) did so after 9 days of growth at 23 °C (p<0.0001). These results demonstrated that CIPK9 plays a critical role in plant growth under low potassium concentrations. Because [K+] in most of the soils is in the micro-molar range 24, 25, CIPK9 may function to regulate potassium nutrition under physiological conditions.
Hypersensitivity to low potassium in the cipk9 mutant may be specific to potassium or cipk9 mutant may also be hypersensitive to deficiency of other nutrients or simply hypersensitive to overall low nutrition status in the media. To test whether cipk9 mutants are hypersensitive to deficiency of other nutrients, we grew wild type and cipk9 mutant plants in modified MS media depleted in each of the following nutrients (calcium, magnesium, iron, phosphorus, nitrate and ammonia and some micronutrients) and found no significant difference between the wild type and mutant plants (data not shown). To test whether cipk9 mutant is hypersensitive to a general low nutrition status, we grew the wild type and cipk9 mutants in different dilutions of the MS media (including 1/20, 1/100, 1/1 000, 1/5 000 and 1/10 000 dilutions). The growth of both wild type and cipk9 mutant seedlings was drastically inhibited as the MS media was diluted progressively from 1/20 to 1/10 000. Again, the cipk9 mutant plants were more inhibited than wild type in root growth and emergence of cotyledons (Figure 5A). A detailed analysis of the root growth and emergence of cotyledons (Figure 5B and 5C) clearly showed that both cipk9 mutant alleles were strongly more sensitive than wild-type seedlings. At 1/1 000 dilution of MS media where the K+concentration is calculated to be 0.02 mM, the root growth of both the cipk9 mutant alleles was approximately 50% of wild type. The emergence of cotyledons in wild type was 83% compared to 54% and 58% for cipk9-1 and cipk9-2, respectively (p<0.0001). Such hypersensitive response of cipk9 mutant alleles to the dilution of MS medium was eliminated by addition of 0.1 mM K+ in the diluted MS media (Figure 5A-C), confirming that CIPK9 specifically regulates potassium nutrition in Arabidopsis. We did not observe a significant difference in growth improvement for over-expression lines of CIPK9 under low low-K+ growth conditions when compared to wild type (data not shown).
K+-uptake and content are not affected in the cipk9 mutant
From earlier studies, it is conceivable that potassium nutrition can be regulated at multiple levels including acquisition and utilization 11, 54. To identify possible mechanisms for the low-potassium hypersensitivity in cipk9 mutants, we examined the potassium acquisition in the mutants and wild-type plants by measuring K+ uptake and content. Using 86Rb+ tracer experiments described in the Materials and methods, we measured the uptake in the wild type and cipk9 mutant seedlings under low (0.02 mM) and high (20 mM) K+ conditions, and did not find significant differences (Figure 6A). We also measured K+ contents in both wild type and mutant plants (seedlings and adult plants) by the atomic absorption spectrophotometry (see Materials and Methods). Again there was no difference in K+ content between wild type and cipk9 mutants under low and optimal levels of K+(Figure 6B). These results indicate that CIPK9 may regulate processes other than K+ acquisition.
Potassium uptake and content in cipk9-1, cipk9-2, and the wild-type (Col-0) plants. (A) 86Rb+-uptake at 0.02 and 20 mM external K+. The uptake time was set at 1 h. (B) Potassium contents in the wild-type (Col-0), cipk9-1, and cipk9-2 seedlings treated at 0.02 and 20 mM external K+. Three independent experiments were performed and values are means ± SD. Student's t test was performed to determine the significance, in (A) and (B), ***p > 0.60.
Discussion
Potassium is a key element for plant growth and low-K+ levels in the soil present a serious problem for agricultural practices. Understanding the molecular mechanism underlying low-K+ response and adaptation in plants will provide a platform for improving crop tolerance to low-K+conditions. We show here that CIPK9, a calcium sensor-interacting protein kinase, functions as an essential component for plant growth under low-K+ conditions. Disruption of CIPK9 gene function in the cipk9 mutants rendered the plants hypersensitive to low-K+ media.
It is well established that the capability of high-affinity K+-uptake in plants plays a pivotal role in low-K+ adaptation 10, 11, 12, 13. As a result, disruption of AKT1, a gene coding for a K+ channel involved in the high-affinity uptake of K+ from the soil or growth medium, resulted in poor growth of Arabidopsis seedlings under low-K+conditions 18, 20. Another high-affinity K+ transporter, HAK5, is also important for potassium nutrition 21. Studies have also demonstrated that the activity and/or expression of a number of K+-transporters including AKT1 and HAK5 are often highly induced by the low-K+ conditions. This finding identifies a natural mechanism for plants to adapt to a low-K+ environment and establishes a “signaling pathway” starting with the low-K+ as a “signal” and increased transport as a “response”. What components are in between the signal and response? In a recent study, Shin and Schachtman 26 have shown that K+-deficient condition leads to generation of H2O2 and changes in gene expression, with the ROS production being the early root response that might modulate the overall physiology of the seedling development. Such a finding is consistent with an earlier study that demonstrated a link between H2O2 production, calcium changes, and root hair development 27. It is shown that a mutation in the NADPH oxidase reduced H2O2 production leading to stunted root growth and shorter root hairs in the mutant plants. The reduced H2O2 production resulted in low calcium channel activity in the root cells. It is believed that H2O2 is required for the activation of calcium channels that in turn trigger the cellular calcium changes essential for cell growth in root hairs and possibly also in other root cells. Under low-K+ conditions, production of H2O2 may also trigger calcium changes required for further signaling steps leading to expression of a number of low-K+-induced genes. Therefore, our finding of a calcium-regulated protein kinase in this study provides a possible molecular link between calcium signals and further cellular regulatory steps. The kinase CIPK9 is a member of the CIPK family, a subfamily of the extended SNF1-like protein kinases (or SnRKs). The activity of CIPKs is regulated by interactions with calcium sensors called CBLs 38, 39, 40. We hypothesize that calcium changes in the cytoplasm alter the conformation of CBLs that interact with and regulate the activity of CIPKs. In this case, CIPK9, after being activated by a CBL, may regulate the activity of other protein components involved in low-K+ responses. These could be transcriptional factors required for low-K+-induced gene expression, metabolic enzymes involved in osmotic regulation, or K+ transporters that are activated under low-K+ condition. It is noteworthy that K+ content and uptake capability were not altered in the cipk9 mutants, indicating that the low-K+ hypersensitivity phenotype may not be caused by less K+ acquisition from the medium. In other words, the downstream targets for CIPK9 may not be directly related to K+ uptake. In addition to participation of CIPK9 in low-K+ tolerance, other members of the CIPK family have been shown to function in other cellular processes including abiotic stress responses and responses to plant hormones such as ABA and GA during seed germination 38, 39, 40 55. One member of CIPKs, SOS2/CIPK24, was shown to be essential for salt tolerance and K+ nutrition by excluding Na+ from the cell 37. Unlike sos2/cipk24 mutant, cipk9 mutant did not show any phenotypic changes under salt or osmotic stress (data not shown), suggesting that CIPK9 functions specifically in K+ nutrition and is not involved in salt tolerance. Taking together, the members in the CBL-CIPK network may function as major components for the regulation of ionic homeostasis including salt tolerance and potassium nutrition in plants. Recently, one of the CIPK members, CIPK23, has been shown to form complexes with CBL1 and CBL9 and regulate potassium nutrition 46, 56. A detailed mechanistic analysis has revealed that CIPK23 physically interacts with and phosphorylates AKT1. The AKT1 channel activity is regulated by CIPK23-CBL1 or CIPK23-CBL9 complexes, thereby regulating potassium nutrition 46, 47. In contrast, CIPK9 was not found to interact with any potassium transporter in the yeast two-hybrid system and did not regulate AKT1 function (data not shown). Future analysis of downstream targets of CIPK9 will add more information to the molecular mechanisms underlying low-K+ response and adaptation.
References
Leigh RA, Jones RGW . A hypothesis relating critical potassium concentrations for growth to the distribution and functions of this ion in the plant-cell. New Phytol 1984; 97:1–13.
Kochian LV, Lucas WJ . Potassium transport in roots. Adv Bot Res 1988; 15:93–178.
Marschner H . Mineral Nutrition of Higher Plants. 2nd Edition, London: Academic Press, 1995.
Maathuis FJM, Sanders D . Mechanisms of potassium absorption by higher plant roots. Physiol Plant 1996; 96:158–168.
Perrenoud S . Potassium and Plant Health. Basel: International Potash Institute, 1990.
Tester M, Blatt MR . Direct measurement of K+ channels in thylakoid membranes by incorporation of vesicles into planar lipid bilayers. Plant Physiol 1989; 91:249–252.
Wu W, Peters J, Berkowitz GA . Surface charge-mediated effects of Mg2+ and K+ flux across the chloroplast envelope are associated with regulation of stroma pH and photosynthesis. Plant Physiol 1991; 97:580–587.
Laegreid M, Bockman OC, Kaarstad O . Agriculture, Fertilizers and the Environment. Oxon: CABI, 1999.
Amtmann A, Armengaud P, Volkov V . Potassium nutrition and salt stress. In Blatt MR, ed. Membrane Transport in Plants. Oxford: Blackwell Publishing, 2004.
Maser P, Thomine S, Schroeder JI, et al. Phylogenetic relationships within cation transporter families of Arabidopsis . Plant Physiol 2001; 126:1646–1667.
Very AA, Sentenac H . Molecular mechanisms and regulation of K+ transport in higher plants. Annu Rev Plant Biol 203; 54:575–603.
Cherel I . Regulation of K+ channel activities in plants: from physiological to molecular aspects. J Exp Bot 2004; 55:337–351.
Ashley MK, Grant M Grabov A . Plant responses to potassium deficiencies: a role for potassium transport proteins. J Exp Bot 2006; 57:425–436.
Epstein E, Rains DW Elzam OE . Resolution of dual mechanisms of potassium absorption by barley roots. Proc Natl Acad Sci USA 1963; 49:684–692.
Epstein E . Dual pattern of ion absorption by plant cells and by plants. Nature 1966; 212:1324–1327.
Siddiqi MY, Glass ADM . Studies of the growth and mineral nutrition of barley varieties. 2. Potassium uptake and its regulation. Can J Bot 1983; 61:1551–1558.
Lagarde D, Basset M, Lepetit M, et al. Tissue specific expression of Arabidopsis AKT1 gene is consistent with a role in K+ nutrition. Plant J 1996; 9:195–203.
Hirsch RE, Lewis BD, Spalding EP, et al. A role for the AKT1 potassium channel in plant nutrition. Science 1998; 280:918–921.
Spalding EP, Hirsch RE, Lewis DR, et al. Potassium uptake supporting plant growth in the absence of AKT1 channel activity: inhibition by ammonium and stimulation by sodium. J Gen Physiol 1999; 113:909–918.
Dennison KL, Robertson WR, Lewis BD, et al. Functions of AKT1 and AKT2 potassium channels determined by studies of single and double mutants of Arabidopsis . Plant Physiol 2001; 127:1012–1019.
Gierth M, Maser P, Schroeder JI . The potassium transporter AtHAK5 functions in K+-deprivation-induced high-affinity K+-uptake and AKT1 K+channel contribution to K+ uptake kinetics in Arabidopsis roots. Plant Physiol 2005; 137:1105–1114.
Fu HH, Luan S . AtKuP1: a dual-affinity K+ transporter from Arabidopsis . Plant Cell 1998; 10:63–73.
Bruggemann L, Dietrich P, Becker D, et al. Channel-mediated high-affinity K+ uptake into guard cells from Arabidopsis . Proc Natl Acad Sci USA 1999; 96:3298–3302.
Adams F . Soil solution. In: Carson EW, ed. The Plant Root and its Environment. Charlottesville, VA: University Press of Virginia, 1971; 441–481.
Glass ADM . Plant Nutrition: An Introduction to Current Concepts. Boston, MA: Jones and Bartlett, 1989.
Shin R, Schachtman DP . Hydrogen peroxide mediates plant root cell response to nutrient deprivation. Proc Natl Acad Sci USA 2004; 101:8827–8832.
Foreman J, Demidchik V, Bothwell JH, et al. Reactive oxygen species produced by NADPH oxidase regulate plant cell growth. Nature 2004; 422:442–446.
Felle HH, Hepler PK . The cytosolic Ca2+ concentration gradient of Sinapis alba root hairs as revealed by Ca2+-selective microelectrode tests and fura-dextran ratio imaging. Plant Physiol 1997; 114:39–45.
Holdaway-Clarke TL, Feijo JA, Hackett GR, et al. Pollen tube growth and the intracellular cytosolic calcium gradient oscillate in phase while extracellular calcium influx is delayed. Plant Cell 1997; 9:1999–2010.
Wymer CL, Bibikova TN, Gilroy S . Cytoplasmic free calcium distributions during the development of root hairs of Arabidopsis thaliana . Plant J 1997; 12:427–439.
Sanders D, Brownlee C, Harper JF . Communicating with calcium. Plant Cell 1999; 11:691–706.
Rudd JJ, Franklin-Tong VE . Unraveling response-specificity in Ca2+ signaling pathways in plant cells. New Phytol 2001; 151:7–33.
Reddy ASN . Calcium: silver bullet in signaling. Plant Sci 2001; 160:381–404.
LaHaye PA, Epstein E . Science 1969; 166:395–396.
Lauchli A . In: Leonard RT, Hepler PK, eds.Calcium in Plant Growth and Development. The American Society of Plant Physiologists Symposium Series. Rockville, MD: American Society Plant Physiology, 1990; 4:26–35.
Liu J, Zhu JK . An Arabidopsis mutant that requires increased calcium for potassium nutrition and salt tolerance. Proc Natl Acad Sci USA 1997; 94:14960–14964.
Zhu JK . Regulation of ion homeostasis under salt stress. Curr Opin Plant Biol 2003; 6:441–445.
Luan S, Kudla J, Rodriguez-Concepcion, et al. Calmodulins and calcineurin B-like proteins: Calcium sensors for specific signal response coupling in plants. Plant Cell 2002; 14:S389–S400.
Kolukisaoglu U, Weinl S, Blazevic D, et al. Calcium sensors and their interacting protein kinases: genomics of the Arabidopsis and rice CBL-CIPK signaling networks. Plant Physiol 2004; 134:43–58.
Batistic O, Kudla J . Integration and channeling of calcium signaling through the CBL calcium sensor/CIPK protein kinase network. Planta 2004; 219:915–924.
Kudla J, Xu Q, Harter K, et al. Genes for calcineurin B-like proteins in Arabidopsis are differentially regulated by stress signals. Proc Natl Acad Sci USA 1999; 96:4718–4723.
Cheong YH, Kim KN, Pandey GK, et al. CBL1, a calcium sensor that differentially regulates salt, drought, and cold responses in Arabidopsis . Plant Cell 2003; 15:1833–1845.
Kim KN, Cheong YH, Grant JJ, et al. CIPK3, a calcium sensor-associated protein kinase that regulates abscisic acid and cold signal transduction in Arabidopsis . Plant Cell 2003; 15:411–423.
Gong D, Guo Y, Schumaker KS, Zhu JK . The SOS3 family of calcium sensors and SOS2 family of protein kinases in Arabidopsis . Plant Physiol 2004; 134:919–926.
Pandey GK, Cheong YH, Kim KN, et al. The calcium sensor calcineurin B-like 9 modulates abscisic acid sensitivity and biosynthesis in Arabidopsis . Plant Cell 2004; 16:1912–1924.
Xu J, Li HD, Chen LQ, et al. A protein kinase, interacting with two calcineurin B-like proteins, regulates K+ transporter AKT1 in Arabidopsis . Cell 2006; 125:1347–1360.
Li L, Kim BG, Cheong YH, et al. A Ca2+ signaling pathway regulates a K+ channel for low-K response in Arabidopsis. Proc Natl Acad Sci USA 2006; 103:12625–12630.
Pandey GK, Grant JJ, Cheong YH, et al. ABR1, an APETALA2-domain transcription factor that functions as a repressor of ABA response in Arabidopsis . Plant Physiol 2005; 139:1185–1193.
Murashige T, Skoog F . A revised medium for rapid growth and bioassays with tobacco tissue culture. Physiol Plant 1 1962; 5:473–497.
Clough SJ, Bent AF . Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana . Plant J 1998; 16:735–743.
Jefferson RA, Kavanagh TA, Bevan MW . GUS fusion:beta-glucurodinase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 1987; 6:3901–3907.
Wu S-J, Ding LT, Zhu JK . SOS1, a genetic locus essential for salt tolerance and potassium acquisition. Plant Cell 1996; 8:617–627.
Armengaud P, Breitling R, Amtmann A . The potassium-dependent transcriptome of Arabidopsis reveals a prominent role of jasmonic acid in nutrient signaling. Plant Physiol 2004; 136:2556–2576.
Horie T, Schroeder JI . Sodium transporters in plants. Diverse genes and physiological functions. Plant Physiol 2004; 136:2457–2462.
Hwang YS, Bethke PC, Cheong YH, et al. A gibberellin-regulated calcineurin B in rice localizes to the tonoplast and is implicated in vacuole function. Plant Physiol 2005; 138:1347–1358.
Cheong YH, Pandey GK, Grant JJ, et al. Two calcium sensors and their interacting kinase regulate transpiration and potassium uptake in Arabidopsis . Plant J 2006, submitted.
Acknowledgements
We are grateful to the ABRC (Ohio State University, Columbus, OH) for Arabidopsis seeds and Syngenta Research and Technology, Torrey Mesa Research Institute (San Diego, CA, USA) for providing the T-DNA insertion lines. This work was supported by a grant from the National Science Foundation (USA) (to SL).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pandey, G., Cheong, Y., Kim, BG. et al. CIPK9: a calcium sensor-interacting protein kinase required for low-potassium tolerance in Arabidopsis. Cell Res 17, 411–421 (2007). https://doi.org/10.1038/cr.2007.39
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2007.39
Keywords
This article is cited by
-
Potassium nutrient status drives posttranslational regulation of a low-K response network in Arabidopsis
Nature Communications (2023)
-
Calcium signaling-mediated transcriptional reprogramming during abiotic stress response in plants
Theoretical and Applied Genetics (2023)
-
Dealing with Environmental Fluctuations: Diversity of Potassium Uptake Systems Across the Three Domains of Life
Journal of Plant Growth Regulation (2023)
-
Molecular and expression analysis indicate the role of CBL interacting protein kinases (CIPKs) in abiotic stress signaling and development in chickpea
Scientific Reports (2022)
-
New functions of CIPK gene family are continue to emerging
Molecular Biology Reports (2022)