Abstract
Apoptosis has a vital role in maintaining tissue homeostasis, and dysregulation of the apoptotic pathway is now widely recognized as a key step in tumourigenesis. Increasingly, evidence has demonstrated that microRNA (miRNA) can exert various biological functions in tumours by targeting oncogenes or tumour suppressors. Nevertheless, the role of miRNA in apoptosis remains unclear. Here we show that ectopical expression of miR-148a can induce apoptosis in colorectal cancer cells. In addition, MYB can inhibit miR-148a by directly acting on the transcription factor binding site in miR-148a gene and miR-148a can posttranscriptionally silence Bcl-2. Subsequently, the intrinsic apoptosis pathway is activated by releasing cytochrome c, cleaving caspase 9, caspase 3 and PARP, which eventually induce cancer-cell apoptosis. These findings are part of a hitherto undocumented apoptotic regulatory pathway in which a pleiotropic transcription factor controls the expression of a miRNA and the miRNA inhibits the target, leading to activation of an intrinsic mitochondrial pathway and tumour apoptosis.
Similar content being viewed by others
Main
Apoptosis is a form of programmed cell death that maintains the balance between the cell-generating effect of mitosis and the cell-loss consequence of cell death in multicellular organisms. Once the defects in the regulation of apoptosis interfere with this delicate balance, the disruption may lead to cancer.1 Over the past decades, many oncogenes and tumour suppressor genes have been identified that regulate apoptosis, such as Bcl-2, p53 and MYC.2, 3, 4, 5, 6 These studies have improved our understanding of apoptotic signalling pathways and their dysregulation in cancer progression and therapy.
Nevertheless, a class of small non-coding RNAs, termed microRNAs (miRNAs), which have been identified recently, have revealed the apoptotic signalling pathways to be more complex and enigmatic than previously appreciated. MiRNAs can recognize target genes by pairing to the mRNAs of protein-coding genes and direct their posttranscriptional repression via the degradation of mRNAs, the inhibition of protein translation or a combination of the two.7, 8, 9 Increasingly, data have clearly shown that miRNAs have vital roles in a wide range of pivotal biological processes in both plants and animals, such as cell growth, development, proliferation, differentiation and death.10, 11 Moreover, miRNAs can regulate the apoptosis of tumour cells. For instance, miR-34a mediates the p53 regulation of apoptosis,12 and miR-15 and miR-16 can induce apoptosis and suppress cell growth and the cell cycle by inhibiting the anti-apoptotic factor Bcl-2.13 However, miR-21 can enable tumour cells to evade apoptosis by downregulating several tumour suppressor genes, including PDCD4, PTEN, RECK, TPM1 and MARCKS.14, 15, 16 The miRNA, miR-148a may repress tumour-cell proliferation and metastasis by targeting DNMT1 and TGIF2.17, 18 However, it remains unknown whether miR-148a functions as a regulator of apoptosis in tumours. Therefore, we have investigated and demonstrated a new pathway in which miR-148a mediates apoptosis in colorectal cancer, and provide insights into the mechanisms of tumour-cell apoptosis.
MiR-148a positively regulates cell apoptosis in vitro
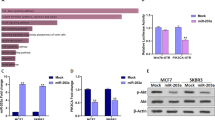
To determine whether miR-148a can regulate apoptosis, we selected three colorectal cancer cell lines, RKO, SW480 and Lovo. First, we investigated the expression of miR-148a in the cell lines by real-time reverse transcription polymerase chain reaction (RT-PCR), which revealed a higher expression in SW480 cells than in RKO and Lovo cells (Figure 1a). We then constructed a miR-148a expression vector that included the genomic sequence of the human miR-148a gene, and ectopically expressed miR-148a in RKO and Lovo cells (Figure 1b). Compared with the control vector, the miR-148a expression vector induced significantly more apoptosis in both RKO and Lovo cells (Figure 1c). To further confirm that miR-148a promoted apoptosis, we used antisense oligonucleotides to silence miR-148a in RKO and Lovo cells transfected with the miR-148a expression vector, and analysed the silencing level by real-time RT-PCR: this showed that the antisense miR-148a inhibitor caused a more than threefold reduction in miR-148a (Figure 1d). We then found that the antisense miR-148a inhibitor could partly reverse the increase in apoptosis induced by miR-148a (Figure 1e). These data suggest that miR-148a may function as a pro-apoptotic activator and trigger early apoptosis in colorectal cancer.
miR-148a positively regulates cell apoptosis. (a) Expression of miR-148a in SW480, RKO and Lovo cells by real-time RT-PCR. (b) Real-time RT-PCR of miR-148a in RKO and Lovo cells with miR-148a expression vector. (c) Apoptosis assay for RKO and Lovo cells with overexpressed miR-148a; the right column, quantitative analysis of apoptosis, with statistics. (d) Reduction of miR-148a in cells (upper panel for RKO cell, lower panel for Lovo cells) with overexpressed miR-148a by antisense miR-148a inhibitor (anti-miR-148a). (e) Partial recovery of miR-148a induced apoptosis by anti-miR-148a; the right column, quantitative analysis of apoptosis, with statistics
The transcription factor MYB directly represses miR-148a
Having demonstrated that miR-148a promoted apoptosis in colorectal cancer cells, we then wanted to investigate how miR-148a itself is regulated. As we thought that miR-148a might be the target of certain transcription factors, we therefore searched TRANSFAC public release 7.019 for the transcription factor binding profile of miR-148a. Among the transcription factors found, MYB was of interest, as it had been identified as an oncogene involved in enhancing proliferation and survival, and in inhibiting apoptosis in human malignancies.20 We then scanned a 5-kb genomic sequence (4.5 kb upstream and 0.5 kb downstream) of the human miR-148a stem loop using pwmatch software (http://www.bioinf.uni-leipzig.de/Software/pwmatch/), and screened two TFBSs, at −3706 bp (TFBS-1) and −2998 bp (TFBS-2), that MYB can recognize (Figure 2a). So, chromatin immunoprecipitation (ChIP) assays were performed to determine whether MYB regulates the expression of miR-148a by binding directly to the miR-148a gene. Two pairs of primers were designed to amplify the two portions spanning the two putative TFBS (TFBS-1 primers and TFBS-2 primers), respectively. Following ChIP of RKO cells with antibody for MYB, PCR was performed and the products were found with only the TFBS-2, but not the TFBS-1 primers (Figure 2b). Similar results were obtained in SW480 cells. This experiment made it clear that MYB was bound to TFBS-2, but not to TFBS-1 in the miR-148a gene. To further determine whether MYB can directly bind the TFBS-2 site in miR-148a gene, electrophoresis mobility shift assay (EMSA) was carried out to analyse the activity for MYB binding to TFBS-2. As shown in Figure 2c, the TFBS-2 oligonucleotide binding activity of nuclear extracts from MYB knockdown cells was significantly reduced relative to that of control cells, which suggested that MYB may act as the modulator of miR-148a by binding to the TFBS-2 site.
miR-148a is repressed by MYB. (a) Human miR-148a gene. The purple lines represent the two TFBSs (TFBS-1 and TFBS-2) and the transcription start site (TSS). (b) ChIP assay in RKO cells. PCR was performed with primers specific for miR-148a TFBS-1 and TFBS-2, respectively. (c) EMSA for detection of the activity for MYB binding to TFBS-2 in the miR-148a gene, the arrows representing the specific formation of complexes with nuclear extract and TFBS-2 probes. (d) Upper panel, expression of miR-148a in SW480, RKO and Lovo cells with siRNA for MYB (siMYB); lower panel, immunoblotting for MYB in SW480, RKO and Lovo cells with siMYB. (e and f) The luciferase activities of TFBS-1 and TFBS-2 reporter plasmid in SW480 (e) and RKO (f) cells with siRNA for MYB, only the TFBS-2 reporter plasmid caused a significant increase in luciferase reporter activity in MYB knockdown cells. (g and h) Apoptosis of RKO (g) and Lovo (h) cells transfected with siMYB and co-transfected with siMYB and anti-miR-148a. The color reproduction of this figure is available on the html full text version of the manuscript
Subsequently, we assayed the expression of miR-148a after knockdown of MYB using siRNA; the results revealed that downregulation of MYB could increase miR-148a expression in RKO, Lovo and SW480 cells (Figure 2d). To provide stronger evidence that miR-148a is suppressed by MYB, we constructed two reporter plasmids comprising the two portions spanning the two putative TFBSs, respectively. When they were transfected to MYB knockdown, control RKO and SW480 cells, the TFBS-2 reporter plasmid caused a significant increase in luciferase reporter activity in the MYB knockdown cells; however, there was no significant change for TFBS-1 reporter plasmid in luciferase reporter activity between MYB knockdown and control cells (Figures 2e and f).
As the knockdown of MYB promoted miR-148a expression, we wanted to know whether MYB affected cell apoptosis through miR-148a. Therefore, we analyzed apoptosis in RKO and Lovo cells after transfection of siRNA for MYB, which showed that knockdown of MYB could also partly induce apoptosis in both cell lines (Figures 2g and h). More interestingly, the antisense miR-148a inhibitor could restore the downregulated MYB induction of apoptosis (Figures 2g and h). Taken together, MYB seems to be a key regulator for miR-148a for promoting apoptosis in colorectal cancer.
Bcl-2 is a direct target for miR-148a silencing
Bcl-2 is a critical molecule for regulating the apoptotic pathway.21 We wanted to know whether miR-148a can silence apoptosis posttranscriptionally by specifically recognizing the 3′-UTR of Bcl-2. The target sites of miR-148a were searched in Bcl-2 mRNA (NM_000633) through RNA22 (http://cbcsrv.watson.ibm.com/rna22.html), and a seed sequence was found in the 3′-UTR of Bcl-2 mRNA (Figure 3a). To validate the predicted target, we cloned the wild-type 3′-UTR of Bcl-2 with the seed sequence to the reporter plasmid, and in parallel we constructed another reporter plasmid carrying a mutant Bcl-2 3′-UTR, especially mutated in the seed sequence (Figure 3a). Transient transfection of the wild reporter plasmid to miR-148a-overexpressing RKO cells produced a significant reduction (P<0.001) in luciferase reporter gene activity compared with the control (Figure 3b); however, the activity of the mutant luciferase reporter gene was not reduced in miR-148a-overexpressing RKO cells. Hence, the 3′-UTR of Bcl-2 is a functional target site for the miR-148a silencing of Bcl-2. In order to further support these results, we investigated whether miR-148a affected the protein or mRNA expression of Bcl-2. The mRNA levels of Bcl-2 were evaluated by real-time RT-PCR: no significant difference was observed between miR-148a-overexpressing RKO cells and control cells (Figure 3c). The levels of Bcl-2 protein were clearly reduced in miR-148a-overexpressing RKO and Lovo cells compared with control cells (Figure 3d). Furthermore, introduction of the antisense miR-148a inhibitor to miR-148a-overexpressing RKO and Lovo cells re-established the Bcl-2 expression (Figure 3e). We also detected the expression of Bcl-2 protein in SW480, RKO and Lovo cell lines; the results showed that the Bcl-2 protein was inversely correlated with the expression of miR-148a (Figure 3f). Then we ascertained that siRNA for MYB could also repress the expression of Bcl-2 (Figure 3g). More intriguingly, the reduced level of Bcl-2 was increased again when the antisense miR-148a inhibitor was transfected into the MYB-downregulated RKO and Lovo cells (Figure 3g).
Bcl-2 is the direct target for miR-148a. (a) The putative target site of miR-148a in the 3′-UTR of Bcl-2, the red letters representing the mutant luciferase reporter gene sequence. (b) Luciferase activity of wild-type (BCL2-UTR-WT) or mutant (BCL2-UTR-mut) BCL2-UTR reporter gene in RKO cells transfected with the miR-148a or control vector. (c) Expression of Bcl-2 in RKO-miR-148a by real-time RT-PCR. (d) Expression of Bcl-2 in RKO and Lovo with overexpressed miR-148a by immunoblotting. (e) Immunoblotting for Bcl-2 in RKO and Lovo cells co-transfected with miR-148a and anti-miR-148a. (f) Expression of Bcl-2 protein in RKO, Lovo and SW480 cells. (g) Immunoblotting for Bcl-2 in RKO and Lovo cells transfected with siMYB and co-transfected with siMYB and anti-miR-148a. (h) Immunoblotting for cleaved caspase 3 and cleaved PARP (left panel), cytochrome c and cleaved caspase 9 (right panel) in RKO cells with miR-148
Bcl-2 can potently inhibit apoptosis in response to most cytotoxic damage, which prevents the release of cytochrome c from the mitochondria and blocks the activation of caspase 9 by the cytoplasmic scaffolding protein Apaf-1.22, 23 To clarify the mechanism of the miR-148a induction of apoptosis through Bcl-2, we evaluated the changes in cytochrome c, cleaved caspase 9, and cleaved caspase 3 and cleaved PARP levels involved in the intrinsic apoptosis pathway: these proteins were upregulated in miR-148a-overexpressing RKO cells (Figure 3h). Hence, these data indicate that miR-148a is controlled by the transcription factor MYB, and activates the cytochrome c–caspase 9–caspase 3–PARP intrinsic apoptosis pathway by silencing Bcl-2 posttranscriptionally in colorectal cancer cells.
MiR-148a inhibits Bcl-2 expression in vivo
The above data have shown that miR-148a inhibited Bcl-2 expression in vitro; can miR-148a also alter Bcl-2 in vivo? To explore this question, miR-148a-transfected or control-transduced RKO cells were implanted into nude mice. Visible xenograft tumours were observed within 3 weeks, and the experiment was terminated at week 8. Real-time RT-PCR revealed that miR-148a was successfully ectopically expressed in the xenograft tumours (Figure 4a). However, there was no significant difference in tumour volume between miR-148a-transfected and control-transduced groups (Figure 4b). However, by immunochemistry and immunoblotting techniques, we found similar results corresponding to those observed in vitro, which showed that Bcl-2 was expressed less in miR-148a ectopic-expressed xenograft tumours than in controls (Figures 4c and d). To elucidate the phenomenon that downregulated Bcl-2 did not change the tumour volume, we investigated another pro-survival member, Bcl-XL expression in miR-148a ectopic-expressed xenograft tumours. Compared with controls, there was no significant difference for the expression of Bcl-XL in two miR148a tumours (Figure 4e, lanes 4 and 5). But the expression of Bcl-XL was reduced in another miR148a tumour (Figure 4e, lane 6), in which (Figure 4d, lane 6) Bcl-2 expression was just relatively higher than that in other miR148a tumours (Figure 4d, lanes 4 and 5). Altogether, these data indicated that the functions of Bcl-2 and Bcl-XL may be compensated for maintaining the tumour survival.
miR-148a inhibits Bcl-2 expression in vivo. (a) Expression of miR-148a in the xenograft. (b) The volume of xenograft tumours with miR-148a. (c) Expression of Bcl-2 in the xenograft by immunochemistry (upper panel for Bcl-2, lower panel for control). (d) Expression of Bcl-2 in the xenograft by immunoblotting. (e) Expression of Bcl-XL in the xenograft by immunoblotting
Expression of miR-148a correlates inversely with Bcl-2 in colorectal tumours
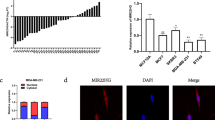
To explore the significance of miR-148a in clinical material, we detected its expression in tumour tissues from 42 patients with colorectal adenocarcinoma; as shown in Figure 5a, there was no significant difference in miR-148a expression between tumour and matched tissue adjacent to the tumour. Furthermore, no significant differences were observed between miR-148a expression and clinicopathological characteristics, including sex, age, tumour invasion, clinical stage or metastasis. However, we found significant differences in miR-148 expression among three groups with different Bcl-2 expressions as defined by immunochemistry (Figure 5b). Moreover, a strong inverse correlation between miR-148a expression levels and Bcl-2 was observed in colorectal cancer tissues evaluated by Spearman's rank correlation (r=−0.604; P<0.001).
Discussion
Apoptosis is crucial for regulating tissue homeostasis, and it is increasingly recognized that impaired apoptosis is a key step in tumorigenesis. Our present work provides a new measure of understanding in the apoptotic pathway. In this study, the transcription factor MYB had been identified as an upstream regulator inhibiting miR-148a expression, and in turn miR-148a could posttranscriptionally silence Bcl-2. Subsequently, the intrinsic apoptosis pathway was activated by releasing cytochrome c, cleaving caspase 9, caspase 3 and PARP, which eventually induced cancer-cell apoptosis (Figure 6). More significantly, not only had Bcl-2 been confirmed as a target of miR-148a in vitro and in vivo, but also the expression of miR-148a was inversely correlated with Bcl-2 in colorectal cancer tissues, although there were no significant differences between miR-148a and clinicopathological characteristics. Otherwise, the data from in vivo experiments showed no significant difference in tumour volume between miR-148a and control xenograft tumours; one cause might be that the function of Bcl-2 that was silenced by miR-148a could be compensated by Bcl-XL or other pro-survival protein; another cause might be that the microenvironment maintained the tumour growth. These data probably could interpret why there were no significant differences between miR-148a and clinicopathological characteristics. However, recently Chen et al. found that miR-148a expression was lower in tumour tissues compared with tissues adjacent to the tumour tissue, and low miR-148 expression correlated with increased tumour size and advanced colorectal adenocarcinoma stage.24 This previous study showed that miR-148a was involved in colorectal cancer metastasis, but they focused on the correlation between miR-148a hypermethylation and metastasis.18 The main cause of the disparity between these and our results was that we only analysed 42 primary tumour samples obtained from the same region at the time of surgical excision, and clinical correlative studies of molecular markers in colorectal cancer and other diseases will require a larger number of primary and metastatic tumours with long-term follow-up.
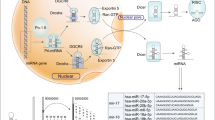
Pathway of miR-148a induction of apoptosis. In the nucleus, pre-miR-148a is transcribed and processed from the miRNA gene in chromosome, which can also be repressed by transcription factor MYB by binding the TFBS-2 upstream of the miR-148a gene. The pre-miR-148a is exported to the cytoplasm, where it is further processed into the mature miR-148a, which is incorporated into the RISC with the argonaute family and other proteins. The RISCs silence Bcl-2 posttranscriptionally by directly binding the 3′-UTRs of Bcl-2 mRNAs. The downregulation of Bcl-2 subsequently activates the intrinsic apoptosis pathway by releasing cytochrome c from mitochondria, cleaving caspase 9, caspase 3 and PARP
The new role for miR-148 has been demonstrated in the present work, namely that it is a pro-apoptotic miRNA, which operates by targeting Bcl-2. Nonetheless, there are some other targets for miR-148a to silence posttranscriptionally. Among them, pregnane X receptor (PXR) is an important target of miRNA-148a,25 which has an anti-apoptotic function in cancer cells.26, 27 More interestingly, the anti-apoptotic capacity of PXR may involve induction of Bcl-2 and protection of cells against Fas-induced apoptosis.28 So, miR-148a can not only directly silence Bcl-2 by binding its 3′-UTR, but it also indirectly represses Bcl-2 by suppressing PXR. It is well known that Bcl-2 acts as the vital regulator against the intrinsic apoptotic pathway. So far, several microRNAs (miRNAs) have been identified as upstream inhibitors of Bcl-2. MiR-15 and miR-16 negatively regulate Bcl-2 expression by the incomplete complementarity with the mRNA of Bcl-2 and promote apoptosis.13 Recently, miR-181b was also found to target Bcl-2 and to induce apoptosis, and then to modulate multidrug resistance in cancer cells.29 These data suggest that miR-148a, together with miR-15, miR-16, miR-181b and other miRNAs, may have important roles in the intrinsic apoptotic pathway. Apart from miRNAs, the transcription factors are also important upstream regulators for their target genes. Every coin has two sides, and the transcription factor MYB, also known as a proto-oncogene, operates as a transcriptional activator as well as transcriptional repressor.30, 31, 32, 33 Hitherto, some evidence has strengthened the case that MYB can transactivate Bcl-2 by recognizing the transcription factor binding site (TFBS).34, 35 More excitingly, the data from our study revealed that MYB functions as a transcriptional repressor by inhibiting miR-148a, which could also silence Bcl-2. So MYB can directly activate Bcl-2: likewise, MYB could positively but indirectly regulate Bcl-2 by transcriptionally suppressing miR-148a. Zhao et al. found that MYB could directly recognize and bind the upstream promoter region of the miR-15/miR-16-1 cluster to promote expression of miR-15; in return, miR-15a could also repress expression of MYB. Consequently, MYB and miR-15 interacted to make an auto-regulatory feedback loop, which modulated both erythroid and myeloid differentiation.36 Taken together, the reciprocal regulation among the miRNAs and their targets comprises an intricate network to control apoptosis.
In conclusion, our studies clarify some important aspects of a new apoptotic pathway in vitro, in which miR-148a suppresses the expression of Bcl-2 at the posttranscriptional level and induces apoptosis by activating a caspase cascade colorectal cancer; moreover, the transcription factor MYB represses the transcription of miR-148a, so miR-148a mediates MYB to regulate Bcl-2. Our data strongly support the idea that miRNAs have important roles in regulating apoptosis. Further research into the targets and upstream regulator of miR-148a and elucidating their interactive mechanism involved in apoptosis may highlight promising targets of therapeutic value for colorectal cancer in the future.
Materials and Methods
Cell culture
The human colorectal cancer cell lines SW480, RKO and Lovo were purchased from the American Type Culture Collection (Manassas, VA, USA). The cells were maintained in RPMI 1640 medium supplemented with 10% (v/v) heat-inactivated bovine serum (Hyclone, Tauranga, New Zealand) and grown at 37 °C in an atmosphere of 95% air and 5% CO2.
DNA constructs
MiR-148a expression plasmids were constructed using BLOCK-iT Pol II miR RNAi Expression Vector Kits (Invitrogen, Carlsbad, CA, USA), and the genomic sequence of pre-miR-148a was synthesized and cloned to the pcDNA6.2-GW/EmGFP-miR vector. The luciferase reporter plasmid PGL3 (Promega, Madison, WI, USA) was applied to clone the reporter gene. Two portions spanning the two putative MYB TFBS of miR-148a respectively were amplified using the TFBS-1 primers (sense: 5′-CGAGCTCACGCCAGGTCAAGTTA-3′; antisense: 5′-CCGCTCGAGGGGGCTAGCCTCATGA-3′) and the TFBS-2 primers (sense: 5′-CGAGCTCTGGTTCGTCTGCTACCC A-3′; antisense: 5′-CCGCTCGAGCGGGGTCCAC CCTCA-3′), and inserted into the upstream of PGL3. The wild and mutated 3′-UTRs of the Bcl-2 gene were synthesized and cloned into the XbaI site downstream of PGL3.
RNA extraction and real-time PCR
Total RNA was obtained from cell lines and tissue samples using TRIzol (Invitrogen). Mature miR-148a and U6 were detected by stem-loop real-time RT-PCR analysis using the TaqMan Human MicroRNA Assay kits (Applied Biosystems, Foster City, CA, USA). The Bcl-2 mRNA was assayed using the primers sense: 5′-CCGTTGGCCCCCGTTGCTTT-3′; antisense: 5′-CTGGCGGAGGGTCAGGTGGA-3′, and the primers sense: 5′-ACCACAGTCCATGCCATCAC-3′; antisense: 5′-TCCACCACCCTGTTGCTGTA-3′ were applied to detect GAPDH. Real-time RT-PCR was performed with SYBR PrimeScript RT-PCR Kit (Takara, Dalian, China).
Transfection assay and oligonucleotides
All transfection experiments were performed using Lipofectamine 2000 according to the manufacturer's instructions. The antisense miR-148a inhibitor (5′-ACAAAGUUCUGUAGUGCACUGA-3′) and siRNA for MYB (sense: 5′-CCACAUUUCUGAAGCACAATT-3′; antisense: 5′-UUGUGCUUCAGAAAUGUGGTA-3′) were synthesized and modified chemically (Invitrogen). Cells were harvested for luciferase and apoptosis assays 72 h after transfection.
Luciferase reporter assay
Various luciferase reporter plasmids were transfected into the cells with Renilla luciferase plasmid. Briefly, for the assay involving MYB repressing miR-148a, 800 ng of PGL3 reporter plasmids for TFBS-1, TFBS-2 or control, 25 ng of Renilla luciferase plasmids and 20 pmol of siRNA for MYB or control were transfected into RKO or SW480 cells of 90% confluence in 24-well plates; otherwise, for the assay involving 148a repressing Bcl-2, 800 ng of PGL3 reporter plasmids for wild or mutated 3′-UTR of the Bcl-2, 25 ng of Renilla luciferase plasmids and 800 ng of pre-miR-148a or control were transfected into RKO cells of 90% confluence in 24-well plates; 24 h after transfection, cells were lysed and luciferase activities were assayed using the Dual-Luciferase Reporter Assay System (Promega).
Apoptosis assay
5 × 105 cells were collected and washed with phosphate-buffered saline (PBS). Cells were then stained with Annexin V-R-PE and 7-AAD, or AnnexinV-FITC (SouthernBiotech, Birmingham, AL, USA) and propidium iodide, and stained cells were detected by flow cytometry. The experiments for the apoptosis assay were performed at least three times.
Chromatin immunoprecipitation
Chromatin immunoprecipitation for MYB interacting with miR-148a was performed using EZ-ChIP kit (Upstate, Waltham, MA, USA). Then, 4 μg of antibody for MYB (Millipore (Temecula, CA, USA) and Santa Cruz (Santa Cruz, CA, USA)) was applied to each immunoprecipitation. The pure DNA was amplified with primers specific for MYB TFBS-1 (sense: 5′-TGGTTCGTCTGCTACCCAACAGT-3′; antisense: 5′-CGGGGTCCACCCTCAGCGAT-3′) and TFBS-2 (sense: 5′-ACGCCAGGTCAAGTTACCCCT-3′; antisense: 5′-GGGGCTAGCCTCATGACATTTGCAG-3′) of miR-148a.
Electrophoretic mobility shift assay
Nuclear extracts were prepared from control and MYB knockdown cells, and EMSA was performed with DIG-labeled oligonucleotides (TFBS-2 probe: 5′-ATGTACACAACGGTATGCAAA-3′) to analyse the binding activity of MYB and Mir-148a using EMSA kit (Roche, Mannheim, Germany).
Immunoblotting
Cells and tissues were lysed on ice for 1 h. Proteins from total cell lysates were resolved by SDS-PAGE, transferred to nitrocellulose membrane, blocked in 5% non-fat milk in PBS/Tween-20, and blotted with the antibodies for Bcl-2 (1 : 1000, Cell Signaling), Bcl-XL (1 : 1000, Cell Signaling), MYB (1 : 1000, Millipore; 1 : 500, Santa Cruz; which was the same as that used for ChIP), β-actin (1 : 5000, Santa Cruz), cytochrome c (1 : 1000, Millipore), cleaved caspase 9 (1 : 1000, Cell Signaling), cleaved caspase 3 (1 : 1000, Cell Signaling) and cleaved PARP (1 : 1000, Cell Signaling). The experiments for the immunoblotting were performed at least three times.
Immunohistochemistry
Paraffin-embedded tissue sections (4 μm) were stained with antibodies for Bcl-2 (1 : 400, Cell Signaling; which was the same as that used for immunoblotting). The positive cells were scored as 0 for <5%, 1 for 6–25%, 2 for 26–50%, 3 for 51–75%, and 4 for 76–100%. Staining intensities were scored as 0 for no staining, 1 for weak, 2 for moderate, and 3 for strong. Immunoreactive scores for Bcl-2 expression were calculated by multiplying these two grading scores, which ranged from 0 to 12.37 Finally, these samples were divided into three groups: group low for 0–4, group middle for 5–8, group high for 9–12.
Tumour cell xenograft study
Eight-week-old male BALB/c-nu/nu mice were used for xenograft experiments. Cells with miR-148a overexpression and control cells (1 × 106 cells) were suspended in 0.2 ml of PBS, and injected subcutaneously into the nape. Xenograft growth and mouse weight were monitored every 3 days from the formation of visible tumour. At the eighth week after inoculation, the xenografts were excised for the analysis of Bcl-2 by immunoblotting and immunohistochemistry.
Clinical data
In total, 42 fresh patient tissue samples of sporadic colorectal adenocarcinoma were obtained from the Quzhou Hospital of Zhejiang, China. Diagnosis and staging followed the 2000 WHO tumor classification and the 2002 UICC TNM staging system. The study was approved by the Ethics Board of Biomedicine, Zhejiang University, China, and all samples were collected with informed consent.
Statistical analysis
The Student's t-test was performed to compare paired data (miR-148a and Bcl-2 mRNA real-time RT-PCR, luciferase reporter assay, apoptosis assay). The relation between miR-148a and the clinicopathological characteristics was analysed by one-way ANOVA. The correlation between the levels of miR-148a and Bcl-2 was determined using Spearman's test. The statistical significance was considered for P<0.05.
Abbreviations
- miR-NTC:
-
control vector
- RT-PCR:
-
reverse transcription polymerase chain reaction
- kb:
-
kilobase pairs
- TFBS:
-
transcription factor binding site
- TSS:
-
transcription start site
- ChIP:
-
chromatin immunoprecipitation
- EMSA:
-
electrophoresis mobility shift
- UTR:
-
untranslated region
- anti-148a:
-
anti-miR-148a or antisense oligonucleotides for miR-148a
- BLK:
-
blank
- siMYB:
-
MYB-siRNA or siRNA for MYB
- si-oligos:
-
control oligonucleotides
- Ago2:
-
Argonaute2
- RISC:
-
RNA-induced silencing complex
- ORF:
-
open reading frame
References
Cotter TG . Apoptosis and cancer: the genesis of a research field. Nat Rev Cancer 2009; 9: 501–507.
Erster S, Mihara M, Kim RH, Petrenko O, Moll UM . In vivo mitochondrial p53 translocation triggers a rapid first wave of cell death in response to DNA damage that can precede p53 target gene activation. Mol Cell Biol 2004; 24: 6728–6741.
Green DR, Kroemer G . Cytoplasmic functions of the tumour suppressor p53. Nature 2009; 458: 1127–1130.
Hoffman B, Liebermann DA . Apoptotic signaling by c-MYC. Oncogene 2008; 27: 6462–6472.
Klefstrom J, Verschuren EW, Evan G . c-Myc augments the apoptotic activity of cytosolic death receptor signaling proteins by engaging the mitochondrial apoptotic pathway. J Biol Chem 2002; 277: 43224–43232.
Pelengaris S, Khan M, Evan GI . Suppression of Myc-induced apoptosis in beta cells exposes multiple oncogenic properties of Myc and triggers carcinogenic progression. Cell 2002; 109: 321–334.
Lee RC, Feinbaum RL, Ambros V . The C elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993; 75: 843–854.
Bartel DP . MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004; 116: 281–297.
Bartel DP . MicroRNAs: target recognition and regulatory functions. Cell 2009; 136: 215–233.
Carrington JC, Ambros V . Role of microRNAs in plant and animal development. Science 2003; 301: 336–338.
Ambros V . MicroRNA pathways in flies and worms: growth, death, fat, stress, and timing. Cell 2003; 113: 673–676.
Chang TC, Wentzel EA, Kent OA, Ramachandran K, Mullendore M, Lee KH et al. Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol Cell 2007; 26: 745–752.
Cimmino A, Calin GA, Fabbri M, Iorio MV, Ferracin M, Shimizu M et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci USA 2005; 102: 13944–13949.
Li J, Huang H, Sun L, Yang M, Pan C, Chen W et al. MiR-21 indicates poor prognosis in tongue squamous cell carcinomas as an apoptosis inhibitor. Clin Cancer Res 2009; 15: 3998–4008.
Park JK, Lee EJ, Esau C, Schmittgen TD . Antisense inhibition of microRNA-21 or -221 arrests cell cycle, induces apoptosis, and sensitizes the effects of gemcitabine in pancreatic adenocarcinoma. Pancreas 2009; 38: e190–e199.
Li T, Li D, Sha J, Sun P, Huang Y . MicroRNA-21 directly targets MARCKS and promotes apoptosis resistance and invasion in prostate cancer cells. Biochem Biophys Res Commun 2009; 383: 280–285.
Braconi C, Huang N, Patel T . MicroRNA-dependent regulation of DNA methyltransferase-1 and tumor suppressor gene expression by interleukin-6 in human malignant cholangiocytes. Hepatology 2010; 51: 881–890.
Lujambio A, Calin GA, Villanueva A, Ropero S, Sanchez-Cespedes M, Blanco D et al. A microRNA DNA methylation signature for human cancer metastasis. Proc Natl Acad Sci USA 2008; 105: 13556–13561.
Wingender E, Karas H, Knuppel R . TRANSFAC database as a bridge between sequence data libraries and biological function. Pac Symp Biocomput 1997: 477–485.
Ramsay RG, Gonda TJ . MYB function in normal and cancer cells. Nat Rev Cancer 2008; 8: 523–534.
Youle RJ, Strasser A . The BCL-2 protein family: opposing activities that mediate cell death. Nat Rev Mol Cell Biol 2008; 9: 47–59.
Cory S, Adams JM . The Bcl2 family: regulators of the cellular life-or-death switch. Nat Rev Cancer 2002; 2: 647–656.
Cory S, Huang DC, Adams JM . The Bcl-2 family: roles in cell survival and oncogenesis. Oncogene 2003; 22: 8590–8607.
Chen Y, Song Y, Wang Z, Yue Z, Xu H, Xing C et al. Altered expression of MiR-148a and MiR-152 in gastrointestinal cancers and its clinical significance. J Gastrointest Surg 2010; 14: 1170–1179.
Takagi S, Nakajima M, Mohri T, Yokoi T . Post-transcriptional regulation of human pregnane X receptor by micro-RNA affects the expression of cytochrome P450 3A4. J Biol Chem 2008; 283: 9674–9680.
Zhou J, Liu M, Zhai Y, Xie W . The antiapoptotic role of pregnane X receptor in human colon cancer cells. Mol Endocrinol 2008; 22: 868–880.
Masuyama H, Nakatsukasa H, Takamoto N, Hiramatsu Y . Down-regulation of pregnane X receptor contributes to cell growth inhibition and apoptosis by anticancer agents in endometrial cancer cells. Mol Pharmacol 2007; 72: 1045–1053.
Zucchini N, de Sousa G, Bailly-Maitre B, Gugenheim J, Bars R, Lemaire G et al. Regulation of Bcl-2 and Bcl-xL anti-apoptotic protein expression by nuclear receptor PXR in primary cultures of human and rat hepatocytes. Biochim Biophys Acta 2005; 1745: 48–58.
Zhu W, Shan X, Wang T, Shu Y, Liu P . miR-181b modulates multidrug resistance by targeting BCL2 in human cancer cell lines. Int J Cancer Feb 16.
Hooper J, Maurice D, Argent-Katwala MJ, Weston K . Myb proteins regulate expression of histone variant H2A Z during thymocyte development. Immunology 2008; 123: 282–289.
Chen J, Kremer CS, Bender TP . The carbonic anhydrase I locus contains a c-Myb target promoter and modulates differentiation of murine erythroleukemia cells. Oncogene 2006; 25: 2758–2772.
Wang DM, Sevcikova S, Wen H, Roberts S, Lipsick JS . v-Myb represses the transcription of Ets-2. Oncogene 2007; 26: 1238–1244.
Afroze T, Husain M . c-Myb-binding sites mediate G(1)/S-associated repression of the plasma membrane Ca(2+)-ATPase-1 promoter. J Biol Chem 2000; 275: 9062–9069.
Lang G, White JR, Argent-Katwala MJ, Allinson CG, Weston K . Myb proteins regulate the expression of diverse target genes. Oncogene 2005; 24: 1375–1384.
Wolff L, Schmidt M, Koller R, Haviernik P, Watson R, Bies J et al. Three genes with different functions in transformation are regulated by c-Myb in myeloid cells. Blood Cells Mol Dis 2001; 27: 483–488.
Zhao H, Kalota A, Jin S, Gewirtz AM . The c-myb proto-oncogene and microRNA-15a comprise an active autoregulatory feedback loop in human hematopoietic cells. Blood 2009; 113: 505–516.
Xue H, Lu B, Zhang J, Wu M, Huang Q, Wu Q et al. Identification of serum biomarkers for colorectal cancer metastasis using a differential secretome approach. J Proteome Res 2010; 9: 545–555.
Acknowledgements
This research is supported by grants from the National Natural Science Foundation of China (30770989, 81090421), Natural Science Foundation of Zhejiang province (D2080011), Science and Technology Department of Zhejiang province (2007C13020), Department of Education of Zhejiang Province (Y200908956) and National Science & Technology Pillar Program during the Eleventh Five-Year Plan Period of China (2006BAI0214 and 2009BAI80B00). We thank Dr Brian Eyden (Manchester, UK) for English language assistance.
Author ContributionsM Lai conceived the project and supervised research. H Zhang designed and performed experiments. Y Li performed DNA constructs. Q Hang collected tumour samples and executed immunohistochemistry. X Ren fulfilled statistical analysis. H Hu conducted apoptosis assay. H Sheng performed tumour cell xenograft study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by D Huang
Rights and permissions
About this article
Cite this article
Zhang, H., Li, Y., Huang, Q. et al. MiR-148a promotes apoptosis by targeting Bcl-2 in colorectal cancer. Cell Death Differ 18, 1702–1710 (2011). https://doi.org/10.1038/cdd.2011.28
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2011.28
Keywords
This article is cited by
-
Comprehensive analysis of miRNA-mRNA regulatory pairs associated with colorectal cancer and the role in tumor immunity
BMC Genomics (2023)
-
Chicken miR-148a-3p regulates immune responses against AIV by targeting the MAPK signalling pathway and IFN-γ
Veterinary Research (2023)
-
Mir-675-5p supports hypoxia-induced drug resistance in colorectal cancer cells
BMC Cancer (2022)
-
MicroRNAs as biomarkers for prostate cancer prognosis: a systematic review and a systematic reanalysis of public data
British Journal of Cancer (2022)
-
MicroRNAs and drug resistance in colorectal cancer with special focus on 5-fluorouracil
Molecular Biology Reports (2022)