Abstract
The mitochondrial rhomboid protease Parl governs apoptosis, morphology, metabolism and might be implicated in Parkinson's disease, but the structural basis of its activity and complex regulation remain unknown. We report the discovery of γ-cleavage, a proteolytic event on the loop connecting the first transmembrane helix (TMH) of Parl to the 6-TMH catalytic rhomboid domain of the protease. This cleavage disrupts the ‘1+6’ structure that defines every mitochondrial rhomboid and generates a new form of Parl, PROD (Parl-rhomboid-domain). Structure–function analysis of Parl suggests that γ-cleavage could be implicated in eliminating Parl proteolytic activity, and structural modeling of PROD reveals structural conservation with the bacterial rhomboid GlpG. However, unlike bacterial rhomboids, which employ a diad-based mechanism of catalysis, Parl appears to use a conserved mitochondrial rhomboid-specific Asp residue on TMH-5 in a triad-based mechanism of catalysis. This work provides unexpected insights into the structural determinants regulating Parl stability and activity in vivo, and reveals a complex cascade of proteolytic events controlling the function of the protease in the mitochondrion.
Similar content being viewed by others
Main
All active prokaryotic and eukaryotic rhomboids share a catalytic domain composed of six-transmembrane helixes (TMHs). In eukaryotes, two distinct and ancient lateral gene transfers gave rise to two rhomboid protein families, PARL and RHO.1 PARL family members, of which the mitochondrial Parl protease is the prototype, share a ‘1+6’ structure, consisting of a TMH appended at the N terminus of the 6-TMHs that form the catalytic rhomboid domain. RHO family members, of which the Drosophila developmental regulator Rhomboid-1 is the prototype,2, 3 are found in non-mitochondrial membranes and share a ‘6+1’ structure, consisting in a TMH appended at the C terminus of the rhomboid domain.1 However, the structural, functional and regulatory contributions of the 7th TMH in PARL and RHO rhomboids remains unknown.
In vitro studies have shown that bacterial rhomboids do not require cofactors to process their substrates, and that the 6-TMH rhomboid domain alone is necessary and sufficient to coordinate their enzymatic activity.4 Within the past 4 years, eight bacterial rhomboid structures have been solved. Seven of them are different conformers or mutants of the GlpG protein from E. coli, and one is from H. influenzae.5, 6, 7, 8 These pioneering studies have shown that bacterial rhomboids are nearly entirely immersed in the detergent micelle, with a compact asymmetrical shape composed of the expected 6-TMH bundle. The universally conserved catalytic Ser residue, on TMH-4, lies submerged ∼10 Å from the presumed plane of the membrane; molecular dynamics studies indicate that water, which is necessary for catalysis, can nonetheless readily access this active site.9 The catalytic Ser is hydrogen bonded to the catalytic His residue situated on TMH-6, both residues are strictly conserved in all active rhomboids. These structures revealed that GlpG lacks a third catalytic residue, which earlier mutagenesis studies suggested to be required for rhomboid's mechanism of catalysis.10 Thus, current biochemical and structural data on prokaryotic rhomboids compellingly support a catalytic mechanism that is based on a Ser and His residue acting as a catalytic dyad.11, 12
The rhomboid domain of mammalian Parl shares <19% of sequence identity with GlpG. Bacterial rhomboid activity is highly sensitive to the phospholipid composition of the membrane.4 Thus, the peculiar composition of the inner mitochondrial membrane (IMM) along with the large electrochemical and pH gradient present inside the organelle might have applied a selective evolutionary pressure that ultimately generated a form of the rhomboid protease capable of efficient catalysis within the unique biophysical environment of the mitochondrion. This possibility is supported by the fact that Parl and its orthologs share highly conserved residues that are not found in bacterial and RHO family members.1 The structural and functional contribution of PARL-specific residues to the activity of mitochondrial rhomboids is unknown.
During metazoan evolution the activity of mitochondrial rhomboids has been recruited to coordinate diverse mitochondrial activities, such as membrane fusion, apoptosis and metabolism.13, 14, 15 This expanded role in the biology of the organelle appears to have required the emergence of novel domains, which were appended at the N terminus of the 6-TMH rhomboid core. Recent studies have shown that a complex mechanism of proteolytic elimination of these domains is required to coordinate Parl activity in vivo. For instance, the vertebrate-specific Parl N terminus, termed Pβ-domain, undergoes two consecutive cleavage events, termed α- and β-cleavage (Figure 1a): the first processing is constitutive and removes the mitochondria-targeting sequence of Parl, to produce MAMP (mature mitochondrial Parl protein); in contrast, β-cleavage is highly regulated and controlled by a phosphorylation switch mechanism that requires Parl rhomboid activity supplied in trans, possibly directly.16 β-Cleavage produces PACT (Parl C-terminal protein), which causes the fragmentation of the mitochondrial reticulum and liberates the Pβ-peptide, a 25 amino-acid-long peptide that is exported to the nucleus to activate nuclear responses linked to mitochondrial biogenesis.14, 15, 16 Thus, while the role of mitochondrial rhomboids as gatekeepers of essential functions of the organelle has emerged,17 the challenge ahead is to provide the structural basis of their activity and regulation. This is all the more essential for mammalian mitochondrial rhomboids because Parl regulates cristae structure13 and mitochondrial biogenesis15 and, being implicated in the cleavage of Pink118, 19, could also be associated to the etiology of Parkinson's disease.20
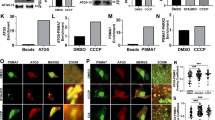
γ-Cleavage generates Parl-rhomboid-domain (PROD). (a) Scheme showing the cleavage of Parl that generates 3 forms of the enzyme. The location of the epitope recognized by our anti-Parl-Cterm is indicated. (b) Expression of the three forms of Parl (MAMP, PACT and PROD) in various rat tissues. (c) Expression of PROD in transfected HeLa cells and epitope mapping experiment showing that γ-cleavage is downstream amino acid 142. A Parl construct with a HA tag at position 142 and a Flag tag at the C-terminus was transfected in HeLa cells. PROD immunoprecipitated with anti-Flag (left lane) but not with anti-HA, indicating that γ-cleavage is in downstream position 142. (d) In vitro activation of β- and γ-cleavage on endogenous Parl. Mouse liver mitochondria preparations were either lysed in RIPA buffer (control) or treated with 1% DDM for 30′ at 4 °C. Right lane shows partial depletion of MAMP and de novo accumulation of PACT and PROD. (e) In vitro activation of β- and γ-cleavage on transfected Parl Flag-CT. Parl cleavage activation assay performed on mitochondria from HeLa cells transfected with a construct expressing Parl Flag-CT. Left lane: control (mitochondria solubilized in RIPA buffer). Right lane: DDM treatment generates de novo PACT and PROD. (f) γ-Cleavage requires the 155INKWW159 sequence on Loop-A. In vitro Parl cleavage activation assay on mutant Parl Δ155–159: deleting this small sequence eliminates PROD but not PACT generation (lane 4). The larger deletion in Parl Δ141–153 does not perturb β- and γ-cleavage (lane 2). (g) Strong conservation of the 155INKWW159 sequence in vertebrate orthologs of Parl. Protein accession numbers are indicated
Results and Discussion
Parl-rhomboid-domain is expressed in vivo
We have shown that β-cleavage of Parl N-terminus causes massive fragmentation of the mitochondrial reticulum.14 To understand the mechanisms implicated in the activation and inactivation of this processing, we investigated the expression of its product, PACT, in vivo. The mechanism(s) underlying β-cleavage activation appears to be under tissue-specific control (Figure 1b). Whereas in rat muscle most of Parl is present as MAMP, in kidney Parl is predominantly present in its β-cleaved form, PACT. Importantly, in tissues where PACT was abundant, such as the spleen and kidney, we also observed an additional smaller form of Parl (Figure 1b). As our anti-Parl antisera detects the C-terminus of Parl and this novel form of the protease has an estimated molecular weight that resembles that of its rhomboid domain (∼20 kDa), it was named PROD (Parl-rhomboid-domain protein). We observed PROD expression also in Parl-transfected HeLa and HEK293 cells (Figure 1c; data not shown).
PROD is generated by γ-cleavage of Parl on the Loop-A
To address whether PROD is the product of a proteolytic processing linked to Parl β-cleavage, we developed an in vitro Parl-cleavage assay. This is based on previous studies using recombinant bacterial rhomboid enzymes4, 21 and consists in partially solubilizing mouse liver or HeLa mitochondria with n-dodecyl-β-D-maltoside (DDM), a mild detergent that is also used to isolate by immunoprecipitation intact mitochondrial electron chain complexes and functional ATPase complexes. After permeabilizing mitochondria with DDM, most of the endogenous MAMP was converted to PACT, and stoichiometric levels of PROD were generated (Figure 1d). Similar results were observed from mitochondria isolated from HeLa cells expressing C-terminally tagged Parl (Figure 1e). Together, these data indicate that PROD is generated by a novel proteolytic processing of Parl, which here we term γ-cleavage.
Given the molecular weight of PROD, the γ-cleavage site is expected to occur on Loop-A (spanning amino acids 121–167); this domain connects the very first TMH of Parl, TMH-A, to the first TMH of the rhomboid domain, TMH-1 (Figure 1a). To map the γ-site, first we performed epitope mapping experiments on a Parl protein double-tagged with a HA sequence at position 142 and a Flag sequence at its C terminus. Mitochondria preparations from HeLa cells transfected with this construct were then subjected to Parl-cleavage assay, followed by separate anti-Flag and anti-HA immunoprecipitations. PROD immunoprecipitated (IP) with anti-Flag, but not with anti-HA, indicating that γ-cleavage occurs after the HA epitope at position 142 (Figure 1c). To identify the site of cleavage, we tested mutants that, carrying deletions downstream of position 142, would ablate the γ-cleavage site. Data showed that whereas a Parl protein lacking amino acids 141–153 could generate PACT and PROD, deleting 155INKWW159 blocked PROD generation, suggesting that γ-cleavage occurs within or near this sequence (Figure 1f). Further, we observed that the W158G and W159G mutations halved the amount of PROD but had no detrimental effect on PACT expression (data not shown). Collectively, these results suggest that the processing that generates PROD occurs in Loop-A. We conclude that γ-cleavage disrupts the ‘1+6’ structure of Parl, the defining feature of every member of the PARL family of mitochondrial rhomboid proteases, and suggest that this processing is functionally implicated in the elimination of Parl activity from the mitochondrion. Notably, Loop-A is strongly conserved in vertebrates (Figure 1g), which can only be explained by functional constraints,22 one of which is thus to host the γ-cleavage site and participate to Parl activity regulation.
PROD generation is mechanistically coupled to β-cleavage activation
To investigate the mechanisms of PROD generation, we asked whether MAMP and/or PACT can be a substrate for γ-cleavage. To this goal, first we assayed Parl mutant proteins carrying either a mutation (L79E) or a deletion (Δ75–79) of the β-cleavage site. These mutations abolish the generation of PACT Parl is, therefore, expressed only in the MAMP form.14 On these mutants, our in vitro Parl-cleavage assay generated only minimal amounts of PROD (Figure 2a), indicating that MAMP-to-PROD conversion is possible but inefficient in the absence of de novo PACT generation. Next, we tested the Parl Δ84–87 mutant; here, the deletion of four residues near the β-cleavage site ablates the complex mechanisms regulating this processing; therefore, in mitochondria expressing this mutant, Parl is present only in the PACT form.14 Again, PROD was generated in very little amounts (Figure 2b), indicating that PACT-to-PROD conversion is also possible but that, in our assay, efficient γ-cleavage depends on de novo β-cleavage activation and PACT generation. This possibility is further supported by data showing that serine protease inhibitors that impair the self-regulated β-cleavage also blocks γ-cleavage (Figure 2c).
PROD generation is mechanistically coupled to β-cleavage activation but is not required for PACT-induced mitochondria fragmentation. (a) Mutations abolishing β-cleavage (L79E and Δ75–79) severely impair de novo PROD generation in vitro. (b) γ-Cleavage requires de novo PACT generation. A Parl mutant expressed only in the PACT form (Δ84–87) is resistant to de novo PROD generation in vitro. (c) Effect of bacterial rhomboid protease inhibitors DCI and TPCK on in vitro generation of PACT and PROD. (d) γ-Cleavage is not functionally required to induce β-cleavage-mediated mitochondria fragmentation. Cells transfected with mutant Parl Δ155–159, which is resistant to γ-cleavage and expresses only MAMP and PACT, have fragmented mitochondria similar to control wild-type Parl which express MAMP, PACT and PROD. Untransfected cells and cells transfected with the β-cleavage-resistant Parl mutant L79E (express only MAMP) have a mix of elongated and rod-like mitochondria14
PROD generation is not required for PACT-induced mitochondria fragmentation
As β-cleavage induces mitochondria fragmentation14 and γ-cleavage appears to be mechanistically coupled to it, we asked whether PACT-to-PROD conversion is required for PACT-mediated fragmentation of the organelle. To this goal, we transfected the γ-cleavage-resistant Parl Δ155–159 mutant and, as positive and negative controls, wild-type Parl and the β-cleavage-resistant L79E mutant. In cells transfected with Parl Δ155–159, mitochondria were fragmented similar to cells transfected with wild-type Parl, which generates PACT and PROD; conversely, in cells transfected with Parl L79E, which does not generate PACT, mitochondria were elongated similar to untransfected cells (Figure 2d). These data indicate that the generation of PROD is not required for PACT-induced mitochondria fragmentation and support a model where γ-cleavage eliminates PACT activity in mitochondrial fusion arrest.17
The rhomboid domain of Parl is structurally conserved
As γ-cleavage disrupts the ‘1+6’ structure that defines every member of the PARL family of mitochondrial rhomboid proteases,1 a key question is to address whether the function of PROD generation is to eliminate MAMP and/or PACT activity from the organelle. However, to our knowledge it is not feasible to address this hypothesis in vivo. This is due to the difficulty to target PROD to the IMM in a way that would also ensure its proper folding and topology. Therefore, to gain functional insights into PROD we built homology models of the rhomboid domain of Parl, looking for structural evidence that would argue in favor, or against, a potential GlpG-like proteolytic activity of PROD.
Our modeling of PROD was conducted using GlpG structures as templates,5, 6, 7, 8 and a manually curated sequence alignment (Figure 3a and Supplementary Figure 1). Our PROD models share the general topology of the bacterial rhomboid domain, with a 6-TMH core capped by similar L1- and L5-loop regions found in the parent bacterial structures (Figure 3b). The models differ primarily in the loop regions with a backbone RMSD between the 6-TMHs of our model and the GlpG structure 3B45 of 1.55 Å. In all rhomboids, the protease activity is attributed to the invariant Ser and His residues that, akin to many soluble serine proteases, form a catalytic dyad. In our PROD models, these residues are S277 in TMH-4 and H335 in TMH-6 and are poised for catalysis similar to GlpG and other soluble serine proteases that contain a catalytic dyad.23
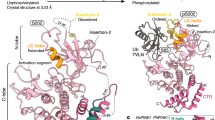
Homology modeling of PROD shows a fold that is similar to that of the bacterial rhomboid GlpG. (a) Sequence alignment of GlpG and Parl used to build homology models. GlpG is a bacterial rhomboid of known structure that served as a template for homology modeling of PROD. The shading indicates the TMHs of GlpG structure 3B45.pdb based on Ramachandran analysis using Insight II. (b) Overall topology of the PROD model with TMHs depicted as cylinders. The catalytic S277 and H335 residues lie on TMH-4 and -6, respectively (depicted as sticks in this and subsequent figures). (c) In structures of GlpG, TMH-1 is stabilized via an N-capping interaction provided by E166 that lies on TMH-2. In our PROD model, a similar stabilizing interaction is organized, however, in a different manner, with Q242 that lies on TMH-3, providing the N-cap (see Supplementary Figure 3 for other examples). Statistical significance: *P<0.05, **P<0.01, ***P<0.001
To assess the quality of our homology model, we looked at whether structural features that appear important for GlpG stability are preserved in PROD, and determined whether there may be any functional requirement for the seventh TMH in the activity of the catalytic core. Analysis of GlpG shows an N-capping interaction that appears to nucleate TMH-1 in an inter-helical manner different than canonical N-capping motifs; here, the interaction arises from the side chain carboxylate of E166 at the end of TMH-2, which reaches across the TMH-2/TMH-1 interface to hydrogen bond with the exposed backbone amides of TMH-1 (Figure 3c, left panel). In our sequence alignment, this E166 of GlpG is V237 in Parl, an amino acid that is unable to form good N-capping interactions. However, in our model residue Q242 at the start of TMH-3 reaches across to form a stabilizing N-capping interaction with TMH-1 (Figure 3c, right panel). Thus, an important structural feature observed in GlpG is conserved in our PROD models, but is performed by a different residue in a different position. This finding was unexpected as our sequence alignment was not guided by such tertiary interactions. Importantly, throughout our models we found other examples of similar conserved structural features, which in some cases are mediated by poorly conserved amino acids and in others by residues located at different positions (Supplementary Figure 3). Thus, despite GlpG and Parl sequences sharing <19% sequence identity, these findings provide confidence that our homology models may be useful for structure–function analysis in vivo.
Parl expression and activity analysis in vivo
In order to experimentally confirm the predictions arising from our structural analysis, we generated a series of point mutants to assay PARL enzymatic activity in transfected HeLa cells. For this, we examined the conversion of MAMP to PACT, which is mediated by Parl in trans.14, 16 Accordingly, mitochondria of HeLa cells transfected with wild-type Parl express MAMP and PACT (Figures 1c, 2a–c, Supplementary Figure 4A), whereas those from cells transfected with the catalytically dead mutant S277G express MAMP, with only minor amounts of PACT being produced by the endogenous Parl activity (Supplementary Figure 4A). Therefore, the proteolytic activity of any Parl mutant can be estimated by comparing, and statistically analyzing, its PACT/MAMP ratio (normalized for the actual level of expression of the mutant protein and for endogenous Parl activity levels) with that of wild-type and S277G Parl (see Materials and Methods). To our knowledge, no other method exists that allow to measure Parl activity in vitro or in a cell-based system; hence, we adopted this approach to address the major structural features of our PROD models.
The packing of TMH-4 and -6 governs Parl expression and activity, and is mediated by small residues
Enzymatic activity of Parl requires proper positioning of S277 and H335 on TMH-4 and TMH-6, which can only be achieved by proper spatial ordering and orientation of these TMHs. In our PROD structures, the positioning of these TMHs appears to depend on Gly residues, consistent with the known role of these residues in mediating helix–helix interactions, often through GxxxG-like motifs.24 In TMH-4, G278 sits next to S277 and forms a GxxxG-like motif that is positioned directly opposite of the catalytic H335 on TMH-6, allowing a close approach of these two TMHs and of their catalytic residues (Figure 4a). Thus, a G278A substitution would be expected to affect catalysis by distancing TMH-4 from TMH-6. However, when we replaced G278 with Ala, Leu or Arg, we found instead that the expression of the Parl protein, but not of its mRNA, was completely lost (Figure 4b, Supplementary Figure 4A), suggesting that the close approach of these TMHs is important also for Parl folding and/or stability in vivo. This result is noteworthy because this residue is nearly universally conserved in all rhomboids, yet in GlpG the corresponding G202C mutation does not affect activity or expression of the bacterial protease.25
Catalytic activity requires close apposition of TMH-4 and -6. (a) A cutaway view of TMH-4/-6 shows close packing that is mediated by small Gly and Ala residues, which allow proper positioning of the catalytic S277 and H335 residues. TMH-4/-6 are depicted as ribbons, interfacial residues are depicted with CPK space-filling models. (b) Replacement of small residues with bulkier amino acids on TMH-4 and -6 disrupts Parl expression and activity, suggesting an essential role of the TMH-4/-6 interface for Parl folding and/or stability in vivo. The bars graph reports the expression level and catalytic activity of the tested Parl mutants. The black column indicates the level of MAMP expression of the mutant relative to wild-type Parl, which is given an arbitrary 100% expression level. The gray column indicates catalytic activity, which is measured by the PACT/MAMP ratio because PACT is generated by the rhomboid activity of MAMP;16 wild-type Parl is considered 100% active and the S277G mutant 0% active. Note that to normalize the catalytic activity of a given Parl mutant by its level of expression, the mutant's PACT/MAMP ratio (Rmut) was divided by the wild-type's PACT/MAMP ratio (Rwt): [(Rmut/Rwt) × 100]. Only mutants having an expression level higher than 30% of wild-type Parl were subjected to this analysis. Expression levels and catalytic activities are reported as mean values (numbers of experiments in squared brackets); error bars represent S.D. Also, note that for mutant Parl proteins with no or low expression levels, proper mRNA expression was verified by semiquantitative RT-PCR (Supplementary Figure 4B). Statistical significance: **P<0.01
Given these results, replacing small residues on TMH-6 with bulkier amino acids would also be expected to distance TMH-4/-6 and disrupt their interface. On TMH-6, G338 and G342 lie just downstream from the catalytic H335 and form a bona fide GxxxG motif that is conserved in all rhomboids. In the case of G338, this residue lies opposite of a ‘hole’ created by G278 and T282 (Figure 4a). When we mutated G338L, Parl expression was lost again. The G338A substitution was more tolerated as it only partially reduced MAMP expression (63±18%) but, as expected, it dramatically reduced proteolytic activity (22±13%; Figure 4b). Importantly, in GlpG structures the corresponding residue, G257, lies opposite to 202GxxxA206, on TMH-4; when mutated in a Val residue, the activity of the recombinant bacterial protease was also severely compromised.21
To further confirm the importance of the TMH-4/TMH-6 interface for Parl expression and catalysis, we also tested G342, which is part of the 338GxxxG342 motif on TMH-6 (these Gly residues are conserved in GlpG, but mutations at these positions have not been tested for expression or activity). In our models, G342 is positioned one turn of helix away from G338 and helps mediate a close approach of TMH-6 and -4. When we introduced the G342L substitution, Parl expression was lost (Figure 4b). As G342 lies opposite to A285 and A286 on TMH-4, we extended our analysis also to these residues. Whereas replacing A285 with the bulkier Leu dramatically decreased Parl expression (11±11%), replacing this amino acid with the similarly sized Cys residue was less perturbing on Parl expression (89±6%) and normalized catalytic activity (79±3%). As a control, we mutated a residue on TMH-4 that does not lie at the TMH-4/TMH-6 interface, C288, and found that the C288A substitution had only minimal perturbing effect on Parl expression or activity (Figure 4b). Thus, replacing small residues at the TMH-4/TMH-6 interface with bulky residues disrupts Parl expression and activity, presumably by preventing a close approach of these helices. Importantly, a major role of TMH-3 appears to be that of positioning and orient the catalytic TMH-4 and -6 (Supplementary Figure 5).
Addressing the role of the L1-loop and L5-cap in Parl expression and activity
Molecular dynamics simulations in GlpG suggest that K132 and other L1-loop residues help orient the rhomboid domain in the lipid bilayer.9 Our data support such role for Parl L1-loop (Figure 5). Furthermore, the L5-cap, which in GlpG is thought to mediate access to the catalytic diad, seems to have a similar function also in mitochondrial rhomboids (Supplementary Figure 6).
The L1-loop residues are important for expression and activity. (a) In structures of GlpG the L1-loop is an array of short amphiphilic helices and unstructured regions that connects TMH-1 and -2. Its functional role is unclear, but in silico studies show cross-talk between the L1-loop and TMH-5 (despite lying on opposite faces), suggesting that these regions are coupled in a manner important for catalysis.9 In our PROD models, an interesting structural feature of the L1-loop is a tripartite hydrogen bonding cluster involving Y195, S198 and S202 (these residues in GlpG are A124, F127 and L131 and would be unable to form such a cluster). In our model, S187 appears involved in an N-capping interaction of the L1a helix. K203 appears to be pointing directly into the hydrophobic membrane bilayer, whereas Y195, T197 and S202 appear to make a hydrogen-bond cluster. (b) Mutation of these residues has mixed effects on expression and activity with profound differences for Y195A and K203A, suggesting an important yet undefined role for the L1-loop. (c and d) The L1-loop is strongly conserved among metazoan orthologs of Parl, but not between Parl and RHO rhomboids. Statistical significance: **P<0.01, ***P<0.001
The PARL-specific Asp319 is critical for Parl expression and activity, and might serve as catalytic residue
Data shown in Supplementary Figure 7 show a critical function of the TMH-2/-5 interface for Parl expression and activity and introduce the possibility that transverse shift of TMH-5 might be important for substrate recognition and activity. Another important feature of TMH-5 is to have an Asp residue (D319) that is strictly conserved in the PARL but not the RHO subfamily of eukaryotic rhomboid proteases (Figures 6a and c and data not shown). When we mutated D319 to Ala/Leu, Parl expression dramatically decreased, which prevented catalytic activity measurements. When we replaced D319 with Glu, which has only an additional CH2 group, we detected no expression of Parl whatsoever. A substitution with Asn, which is nearly isosteric with Asp, was somewhat more tolerated as it reduced expression to 61±9%; however, compared with wild-type Parl, the D319N mutation severely reduced proteolytic activity to 19±9% (Figure 6b).
The PARL family-specific Asp319 is central for Parl expression and activity. (a and b) In our PROD models, D319 is pointed into the membrane bilayer. Note that a rotation of TMH-5 would bring this residue adjacent to H335, where it could serve as a third catalytic residue. Mutations in TMH-5 designed to allow (A314I and G309A) or prevent (G322L, M318I) rotation of TMH-5 have effects on Parl activity that are consistent with this possibility. (c) Strict conservation of D319 on the TMH-5 of eukaryotic orthologs of Parl. Note that this residue is conserved only in PARL family members and not in RHO family members or in bacterial rhomboids. Statistical significance: *P<0.05, **P<0.01
These results are noteworthy because aspartate has an important role as a third catalytic residue in the classical serine protease mechanism. We therefore wondered if D319 may act in this manner. In our homology models, D319 on TMH-5 lies in the same plane of the catalytic Ser and His residues, but points away from the active site and appears directed into the lipid bilayer. However, if TMH-5 is rotated, then D319 could be poised to stabilize the orientation of H335 in the same manner observed in the serine protease chymotrypsin.26 In GlpG, TMH-5 appears dynamic and adopts different conformations.7, 21 Further, the GlpG crystallographic data show a lack of electron density for the L5-cap, and the molecular dynamics simulations of GlpG show large root-mean-squared fluctuations for the TMH-5 and the adjacent loop,9 which would be a prerequisite for its rotation.
In Parl, a similar dynamic nature of TMH-5 might allow D319 to act as a third catalytic residue. Given the difficulty of testing this possibility in the absence of a high-resolution structure, we asked whether mutations designed to prevent TMH-5 rotation or dynamics would affect Parl activity. We reasoned that such dynamic movements would most likely arise from rotation of the entire helix as in our model, and not from local unfolding. Therefore, we engineered a number of single point mutations designed to test whether TMH-5 rotation was plausible.
One helical turn away from D319 is G322, which points into the membrane bilayer. In our models, the G322L substitution would inhibit TMH-5 rotation by creating steric clashes with the L5-cap. Consistent with this prediction, the G322L mutation had little effect on Parl expression or stability, but reduced its normalized activity to less than half of wild type (Figure 6b).
If TMH-5 is indeed dynamic, it could also be expected that replacing a small residue that mediates the TMH-4/-5 interface with a bulky residue would not disrupt activity in the same manner we observed for other TMH-4 interfaces. To test this, we replaced A314 with Ile. A314 lies on TMH-5 at the point of closest approach with TMH-4; it is also conserved in our sequence alignment with GlpG (Figure 3a). The A314I substitution was very well tolerated (Figure 6b), consistent with a model where the TMH-4/-5 interface is governed by a rigid TMH-4 and a dynamic TMH-5.
Toward the start of TMH-5 lies G309 (G228 in GlpG), which is positioned slightly away from the TMH-2/-5 interface. This residue would be expected to remain pointing into the membrane bilayer upon rotation of TMH-5 and, therefore, tolerate substitutions with aliphatic amino acids. Consistent with this prediction, we found that the G309A/L substitutions had only modest effect on Parl activity.
Adjacent to D319 lies M318, which in our models is at the TMH-5/-6 interface. In order for D319 to rotate into the catalytic site, M318 must also rotate in toward the rhomboid core and ultimately end up at the TMH-5/-2 interface. Met is well suited for such motion as it is well known for its structural plasticity.27, 28, 29 When we replaced M318 with Ile, which is bulkier and populates only one of the three low-energy rotameric states in an α-helix,30 the expression of the M318I mutant protein was modestly decreased but its normalized activity fell below 50%. Interestingly, the M318A substitution would be expected to allow rotation of TMH-5; however, this mutation dramatically suppressed protein expression, warning that positions 318/319 of Parl might be essential also for governing protein folding and stability in vivo.
These results support the possibility that γ-cleavage, by disrupting the ‘1+6’ structure of Parl, triggers a rotation of TMH-5 that would bring D319 away from the catalytic site and destabilize the protein, supporting the concept that this novel processing of Parl presides to the negative regulation of the activity of the rhomboid protease in the mitochondrion. Ultimately, whether D319 participates in catalysis should be resolved with high-resolution structures of PROD which, being similar to those of GlpG, we anticipate to be possible to obtain.
Materials and Methods
Parl β- and γ-cleavage in vitro activation assay
HeLa cells were transfected using Fugene6 (Roche Diagnostics, Indianapolis, IN, USA) for 36 h. Crude mitochondria preparations were obtained as described31 from three 175 cm2 plates of HeLa cells grown to confluence; for each assay, 700 μg of proteins (estimated by Bradford analysis) were added to ice-cold PBS buffer containing 1% dodecyl maltoside (5 mg/ml final). After 30 min of incubation in ice, RIPA buffer (65 mM Tris base, 150 mM NaCl, 1% NP40, 0.25% Na deoxycholate, 0.1% SDS, 1 mM EDTA, pH 7.4) containing a protease inhibitors cocktail (Roche) was added (500 μl, final volume). Lysed membranes were incubated at 4 °C for 30′, and cleared by centrifugation; the various forms of Parl were IP and immunoblotted (WB) as indicated.
Parl protein expression and proteolytic activity analysis
HeLa cells were seeded in six-well plates at 2 × 105 cells/well and transfected the following day with a pcDNA3 construct expressing a wild-type or mutant Parl tagged at the C-terminus with the Flag sequence. After 72 h, cells were lysed in 800 μl of RIPA buffer containing protease inhibitors. After high-speed centrifugation, lysates were incubated overnight at 4 °C with anti-Flag M2 beads (Sigma-Aldrich, St. Louis, MO, USA). After extensive washes with STEN buffer, immunoprecipitates were run on a 4–12% gradient polyacrylamide gels, blotted, and immunolabeled with anti-Flag-HRP (Sigma-Aldrich). Detection of the various forms of Parl was carried out by chemiluminescence (Pierce, Thermo Fisher Scientific, Rockford, IL, USA) using the VersaDoc2000 imaging system. Densitometric analysis of MAMP and PACT was performed using the Quantity-One analysis software (Bio-Rad, Mississauga, Ontario, Canada).
Typically, the protein expression and activities of each mutant was measured in three independent experiments. Constructs expressing wild-type and catalytically dead Parl (S277G) were included in each experiment, as they served as positive and negative control of Parl activity as well as positive controls of Parl protein expression. The expression level of wild-type Parl was arbitrarily assigned to be 100%. The expression level of a mutant Parl protein was calculated by [(MAMPmut/MAMPwt) × 100].
The catalytic activity of Parl was calculated by the PACT/MAMP ratio. The value for wild-type Parl (Rwt) was assigned to be 100%, and that of the catalytically dead S277G mutant (Rdead) to be 0%. Note that as a small amount of Parl S277G mutant is cleaved in trans by endogenous Parl, the actual Rdead, typically negligible, was nonetheless subtracted from Rwt as well as from every calculated Rmut. To normalize the catalytic activity of a given Parl mutant by its level of expression, the mutant's PACT/MAMP ratio (Rmut) was divided by Rwt: [(Rmut/Rwt) × 100]. Note that only mutants having an expression level higher than 30% of wild-type Parl were subjected to this analysis. Catalytic activities are reported as mean values; error bars represent S.D.
Homology modeling and validation of PARL core structure
Homology models of the PROD domain of PARL with the bacterial rhomboid protease GlpG were built using structures5, 6, 7, 8, 32, 33 (pdb accession codes 2IC8, 2NRF – chain A and B, 2NR9, 2O7L, 3B44, 3B45, 2IRV – chain A and B) and a manually curated sequence alignment. In generating our alignment, we kept insertions to loop regions (the Pfam alignment creates an insertion in TMH-5), maximized sequence conservation in TMH between Parl and GlpG, took into account helical propensity principles using AGADIR,34 and applied other considerations that are typically not incorporated into sequence alignment algorithms. This curation produced an alignment on which structure models of PROD were generated using manual and automated approaches with INSIGHT II (Accelrys Inc., San Diego, CA, USA), the Swiss Modeler server35 and the MODELLER software.36 The figures displayed are from a single model generated from Swiss-Modeller (similar structural features were found in other tested models). In this model, we found by Molprobity analysis37 that 92.9% of all residues were in favored (98%) regions and 6.7% (176/182) of all residues were in allowed (>99.8%) regions. There were six outliers (see Ramachandran plot in Supplementary Figure 2): V185, P186, L274, G275, G294 and T305. These residues lie in a turn directly after TMH-1 (V185 and P186), the loop between TMH-3/-4 (L275 and G275) and the loop between TMH-4/-5 (T305). The figures were made using PyMOL.38
Abbreviations
- MAMP:
-
mature mitochondrial Parl protein
- PACT:
-
Parl C-terminal protein
- PROD:
-
Parl rhomboid domain protein
- TMH:
-
transmembrane helix
- IMM:
-
inner mitochondrial membrane
- DDM:
-
n-dodecyl-β-D-maltoside
References
Koonin EV, Makarova KS, Rogozin IB, Davidovic L, Letellier MC, Pellegrini L . The rhomboids: a nearly ubiquitous family of intramembrane serine proteases that probably evolved by multiple ancient horizontal gene transfers. Genome Biol 2003; 4: R19.
Urban S, Lee JR, Freeman M . Drosophila rhomboid-1 defines a family of putative intramembrane serine proteases. Cell 2001; 107: 173–182.
Urban S . Rhomboid proteins: conserved membrane proteases with divergent biological functions. Genes Dev 2006; 20: 3054–3068.
Urban S, Wolfe MS . Reconstitution of intramembrane proteolysis in vitro reveals that pure rhomboid is sufficient for catalysis and specificity. Proc Natl Acad Sci USA 2005; 102: 1883–1888.
Wang Y, Zhang Y, Ha Y . Crystal structure of a rhomboid family intramembrane protease. Nature 2006; 444: 179–180.
Wu Z, Yan N, Feng L, Oberstein A, Yan H, Baker RP et al. Structural analysis of a rhomboid family intramembrane protease reveals a gating mechanism for substrate entry. Nat Struct Mol Biol 2006; 13: 1084–1091.
Ben-Shem A, Fass D, Bibi E . Structural basis for intramembrane proteolysis by rhomboid serine proteases. Proc Natl Acad Sci USA 2007; 104: 462–466.
Lemieux MJ, Fischer SJ, Cherney MM, Bateman KS, James MN . The crystal structure of the rhomboid peptidase from Haemophilus influenzae provides insight into intramembrane proteolysis. Proc Natl Acad Sci USA 2007; 104: 750–754.
Bondar AN, del Val C, White SH . Rhomboid protease dynamics and lipid interactions. Structure 2009; 17: 395–405.
Clemmer KM, Sturgill GM, Veenstra A, Rather PN . Functional characterization of Escherichia coli GlpG and additional rhomboid proteins using an aarA mutant of Providencia stuartii. J Bacteriol 2006; 188: 3415–3419.
Erez E, Fass D, Bibi E . How intramembrane proteases bury hydrolytic reactions in the membrane. Nature 2009; 459: 371–378.
Ha Y . Structure and mechanism of intramembrane protease. Semin Cell Dev Biol 2009; 20: 240–250.
Cipolat S, Rudka T, Hartmann D, Costa V, Serneels L, Craessaerts K et al. Mitochondrial rhomboid PARL regulates cytochrome c release during apoptosis via OPA1-dependent cristae remodeling. Cell 2006; 126: 163–175.
Jeyaraju DV, Xu L, Letellier MC, Bandaru S, Zunino R, Berg EA et al. Phosphorylation and cleavage of presenilin-associated rhomboid-like protein (PARL) promotes changes in mitochondrial morphology. Proc Natl Acad Sci USA 2006; 103: 18562–18567.
Civitarese AE, MacLean PS, Carling S, Kerr-Bayles L, McMillan RP, Pierce A et al. Regulation of skeletal muscle oxidative capacity and insulin signaling by the mitochondrial rhomboid protease PARL. Cell Metab 2010; 11: 412–426.
Sík A, Passer BJ, Koonin EV, Pellegrini L . Self-regulated cleavage of the mitochondrial intramembrane-cleaving protease PARL yields Pbeta, a nuclear-targeted peptide. J Biol Chem 2004; 279: 15323–15329.
Hill RB, Pellegrini L . The PARL family of mitochondrial rhomboid proteases. Semin Cell Dev Biol 2010; 21: 582–592.
Deas E, Plun-Favreau H, Gandhi S, Desmond H, Kjaer S, Loh S et al. PINK1 Cleavage at position A103 by the mitochondrial protease PARL. Hum Mol Genet 2010; 20: 867–879.
Jin SM, Lazarou M, Wang C, Kane LA, Narendra DP, Youle RJ . Mitochondrial membrane potential regulates PINK1 import and proteolytic destabilization by PARL. J Cell Biol 2010; 191: 933–942.
Shi G, Lee JR, Grimes DA, Racacho L, Ye D, Yang H et al. Functional alteration of PARL contributes to mitochondrial dysregulation in Parkinson's disease. Hum Mol Genet 2011 (E-pub ahead of print 25 February 2011).
Baker RP, Young K, Feng L, Shi Y, Urban S . Enzymatic analysis of a rhomboid intramembrane protease implicates transmembrane helix 5 as the lateral substrate gate. Proc Natl Acad Sci USA 2007; 104: 8257–8262.
Koonin EV, Wolf YI . Constraints and plasticity in genome and molecular-phenome evolution. Nat Rev Genetics 2010; 11: 487–498.
Rawlings ND, Barrett AJ . Families of serine peptidases. Methods Enzymol 1994; 244: 19–61.
MacKenzie KR, Fleming KG . Association energetics of membrane spanning alpha-helices. Curr Opin Struct Biol 2008; 18: 412–419.
Maegawa S, Koide K, Ito K, Akiyama Y . The intramembrane active site of GlpG, an E. coli rhomboid protease, is accessible to water and hydrolyses an extramembrane peptide bond of substrates. Mol Microbiol 2007; 64: 435–447.
Tsukada H, Blow DM . Structure of alpha-chymotrypsin refined at 1.68 A resolution. J Mol Biol 1985; 184: 703–711.
Bernstein HD, Poritz MA, Strub K, Hoben PJ, Brenner S, Walter P . Model for signal sequence recognition from amino-acid sequence of 54K subunit of signal recognition particle. Nature 1989; 340: 482–486.
O′Neil KT, Erickson-Viitanen S, DeGrado WF . Photolabeling of calmodulin with basic, amphiphilic alpha-helical peptides containing p-benzoylphenylalanine. J Biol Chem 1989; 264: 14571–14578.
Gellman SH . On the role of methionine residues in the sequence-independent recognition of nonpolar protein surfaces. Biochemistry 1991; 30: 6633–6636.
McGregor MJ, Islam SA, Sternberg MJ . Analysis of the relationship between side-chain conformation and secondary structure in globular proteins. J Mol Biol 1987; 198: 295–310.
Jeyaraju DV, Cisbani G, De Brito OM, Koonin EV, Pellegrini L . Hax1 lacks BH modules and is peripherally associated to heavy membranes: implications for Omi/HtrA2 and PARL activity in the regulation of mitochondrial stress and apoptosis. Cell Death Differ 2009; 16: 1622–1629.
Wang Y, Ha Y . Open-cap conformation of intramembrane protease GlpG. Proc Natl Acad Sci USA 2007; 104: 2098–2102.
Wang Y, Maegawa S, Akiyama Y, Ha Y . The role of L1 loop in the mechanism of rhomboid intramembrane protease GlpG. J Mol Biol 2007; 374: 1104–1113.
Muñoz V, Serrano L . Development of the multiple sequence approximation within the AGADIR model of alpha-helix formation: comparison with Zimm-Bragg and Lifson-Roig formalisms. Biopolymers 1997; 41: 495–509.
Arnold K, Bordoli L, Kopp J, Schwede T . The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 2006; 22: 195–201.
Eswar N, Webb B, Marti-Renom MA, Madhusudhan MS, Eramian D, Shen MY et al. Comparative protein structure modeling using MODELLER. Curr Protoc Protein Sci 2007 Chapter 2: Unit 2.9.
Chen VB, Arendall WB, Headd JJ, Keedy DA, Immormino RM, Kapral GJ et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr D Biol Crystallogr 2010; 66: 12–21.
DeLano WL . The PyMOL molecular graphics system. DeLano Scientific 2002.
Acknowledgements
We thank Li Xu for assistance in mitochondria morphology analysis, Katalin Toth for support with the statistical analysis, and former members of the Pellegrini Laboratory for technical assistance. This study was supported by a NIH grant to RBH (RO1GM067180), by a CIHR grant to LP (MOP-82718), and by a FRSQ senior scholarship to LP. DVJ is a CIHR Canada Graduate Scholar.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by L Scorrano
Supplementary Information accompanies the paper on Cell Death and Differentiation website
Supplementary information
Rights and permissions
About this article
Cite this article
Jeyaraju, D., McBride, H., Hill, R. et al. Structural and mechanistic basis of Parl activity and regulation. Cell Death Differ 18, 1531–1539 (2011). https://doi.org/10.1038/cdd.2011.22
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2011.22
Keywords
This article is cited by
-
The rhomboid protease family: a decade of progress on function and mechanism
Genome Biology (2011)
-
Derlin-1 is a rhomboid pseudoprotease required for the dislocation of mutant α-1 antitrypsin from the endoplasmic reticulum
Nature Structural & Molecular Biology (2011)