Abstract
We have established a systematic high-throughput screen for genes that cause cell death specifically in transformed tumor cells. In a first round of screening, cDNAs that induce apoptosis in a transformed human cell line are detected. Positive genes are subsequently tested in a synthetic lethal screen in normal cells versus their isogenic counterparts that have been transformed by a particular oncogene. In this way, the organic cation transporter-like 3 (ORCTL3) gene was found to be inactive in normal rat kidney (NRK) cells, but to induce apoptosis in NRK cells transformed by oncogenic H-ras. ORCTL3 also causes cell death in v-src-transformed cells and in various human tumor cell lines but not in normal cells or untransformed cell lines. Although ORCTL3 is a member of the organic cation transporter gene family, our data indicate that this gene induces apoptosis independently of its putative transporter activity. Rather, various lines of evidence suggest that ORCTL3 brings about apoptosis by an endoplasmic reticulum stress-mediated mechanism. Finally, we detected ORCTL3 to be downregulated in human kidney tumors.
Similar content being viewed by others
Main
The finding that various cytostatic drugs elicit an apoptotic response in cancer cells1 has generated great interest in the application of this cellular phenomenon against tumor cells. Underlying these efforts is the hope that the manifold changes inflicted on cancer cells during the transformation process and the resultant aberrant signaling circuits render them sensitive to specific pro-apoptotic stimuli that spare normal cells. Several genes have been found to induce the cellular suicide program specifically in transformed tumor cells. Among them are the viral genes E4orf42 and apoptin,3 and also, importantly, cellular genes such as mda-7/IL-24,4 Hamlet,5 par-4,6 and TRAIL.7 For none of these genes is it known how exactly they exert their tumor-specific effect. Nevertheless, these discoveries established that the genome contains sequences that act against malignant transformation by inducing apoptosis. So far, tumor-specific apoptosis genes have been discovered only fortuitously. Moreover, research conducted to date has not addressed the genetic changes of the transformation process that activate these genes as apoptosis inducers. We speculated that additional tumor-specific apoptosis genes exist in the genome that target cellular changes caused by specific oncogenes. Hence, we set up a genome-wide screen to systematically discover pro-apoptotic genes that are activated by the H-ras oncogene. We present here ORCTL3 as the first tumor-specific pro-apoptotic gene from this screen.
Results
Screen for tumor-specific apoptosis-inducing genes
We have used our recently described RISCI screen8 to search for pro-apoptotic cDNA plasmids in the kidney cell line HEK293T. These cells display characteristic features of tumor cells such as growth in soft agar and tumor formation in nude mice.9 Our screen10 has so far led to 112 apoptosis genes out of a total of 150 000 clones or ∼60% of one normalized cDNA library.11, 12, 13, 14, 15 All genes that scored positive in this assay were subsequently transfected in parallel in H-ras-transfected normal rat kidney (NRK) cells and in their untransformed, parental cells. This synthetic lethal screen allowed us to conduct the experiments in a defined isogenic background (Figure 1a). By using NRK cells, we remained in a renal cell system with the cDNAs taken from the kidney and with the first indicator cell line (HEK293T) also from this organ. We chose H-ras as this gene is activated in about 20% of human tumors16 and imposes resistance to apoptosis,17 a hallmark of tumor cells.18 Moreover, H-ras elicits the same malignant changes in NRK cells as observed with HEK293 cells,19 suggesting that it causes common signaling modifications. Phase contrast microscopy revealed the typical irregular growth pattern of transformed cells in contrast to wild-type (WT) NRK cells and an immunoblot produced a prominent signal generated by the transfected H-ras gene (Supplementary Figure 1A and B). This established these cells as a bona fide transformation system. A total of 44 genes from the primary screen remained active for apoptosis induction in H-ras-transformed NRK cells. In accordance with the cell death inhibitory effect of H-ras, several genes were more potent apoptosis inducers in WT NRK cells (not shown). However, one clone, later identified as ORCTL3 (see below), was active for apoptosis induction in the H-ras-transformed cells but did not cause detectable apoptosis in WT NRK cells. Visual inspection of cells marked with a co-transfected GFP construct showed that this gene induced cell shrinkage and membrane blebbing, both typical features of apoptotic cells (Figure 1b and c). Other apoptosis genes from the screen, including a number of transporters and plasma membrane-bound proteins, did not score positive in this assay. A quantitative assay specific for apoptotic DNA cleavage and cell death inhibition by the pan-caspase inhibitor zVAD confirmed apoptosis induction (Figure 1d and e). Sequencing identified the gene as full-length ORCTL3/SLC22A13, which was originally determined through large-scale sequencing of genomic DNA.20 On the basis of its sequence homology, it is a member of the organic cation transporter (OCT) family21 that mediates the absorption and secretion of many structurally diverse cations.21 To date, however, ORCTL3 is one of the least studied OCTs and only recently it has been shown to be a urate and high-affinity nicotinate transporter.22
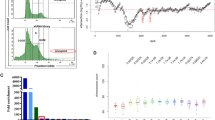
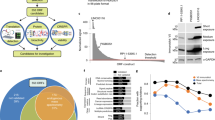
ORCTL3 specifically induces apoptosis in H-ras-transformed cells compared to wild-type (WT) NRK cells. (a) Outline of the screen for tumor-specific apoptosis genes. Aliquots containing single bacteria clones were inoculated in wells of 96-well blocks in LB medium, and the plasmid DNA was isolated using silica oxide. After calcium phosphate transfections in 96-well plates, an enzymatic readout for apoptosis detection was employed. The plasmids of those genes that scored positive in this assay were then examined in parallel transfections in WT NRK cells and NRK cells transformed by the H-ras oncogene. To this end the transfection mixes were equally split and added to the cells. (b) Specific generation of the apoptosis phenotype by ORCTL3 in H-ras-transformed cells. Transfection mixes with ORCTL3 (or luciferase) and GFP expression plasmids were equally distributed between WT and H-ras-transformed NRK cells. Fluorescence pictures were taken after 24 h. (c) Quantification of the apoptosis induction by ORCTL3 in NRK WT and NRK H-ras cells. At 24 h after co-transfecting an expression vector for GFP and ORCTL3 (or luciferase), apoptosis was assessed by phenotype quantification. Shown are the means and the standard deviation of three independent experiments (P<0.01 in H-ras NRK cells, nonsignificant in WT NRK cells). (d) Quantification of apoptosis by ORCTL3 with an ELISA against DNA fragments. At 24 h after the transfection of expression constructs for ORCTL-3 or luciferase, an ELISA was performed that detects DNA fragments generated during apoptosis (P<0.01 in H-ras NRK cells, P-value nonsignificant in WT NRK cells, n=3). (e) Reduction of apoptosis by the pan-caspase inhibitor zVAD. NRK H-ras cells were transfected with the indicated plasmids together with an expression construct for GFP. After the transfection, 25 μ M zVAD was added to the cells. Apoptosis was quantified after 24 h by phenotype inspection (P<0.01, n=3)
Characterization of cell death by ORCTL3
We performed a structure–function analysis by testing progressive deletions of ORCTL3 for apoptosis induction. Figure 2a reveals that the C-terminal half of the protein can be deleted without substantially disrupting the apoptosis potential of ORCTL3. This indicated that ORCTL3 does not require its putative transporter activity for apoptosis induction.
Characterization of cell death by ORCTL3. (a) Structure–function analysis of ORCTL3. The indicated C-terminal deletion mutants of ORCTL3 were transfected into HeLa cells and the extent of apoptosis recorded by phenotype quantification. The shaded areas in the schematic representation of ORCTL3 indicate the putative transmembrane domains (P<0.01 for all ORCTL3 constructs in comparison to Luc samples, n=3). (b) Comparison of ORCTL3-induced apoptosis with the effects of two other transporter molecules. Equal amounts of expression vectors for ORCTL3 and two organic cation transporters, SLC22A1 and SLC22A12, were transfected into HeLa cells and apoptosis determined by phenotype quantification (P-value for both nonsignificant relative to Luc samples, n=3). (c) Transient co-transfection of H-ras together with ORCTL3 does not induce apoptosis. Equal amounts of expression plasmids for H-ras and ORCTL3 or luciferase were transfected into NRK WT cells. After 24 h apoptosis was quantified (P-value nonsignificant, n=3). (d) Apoptosis induction in v-src-transformed NRK cells by ORCTL3. Photomicrographs show the morphological appearance of NRK v-src cells transfected with a GFP expression plasmid together with the indicated expression plasmids for luciferase or ORCTL3. (e) Quantification of apoptosis in v-src-transformed NRK cells. At 24 h after the transfection of the respective plasmids, apoptosis was quantified by phenotype inspection (P<0.01, n=3)
To determine the specificity of the signal elicited by ORCTL3, we transfected SLC22A1 and SLC22A12, two related gene family members of the OCT family, but have not observed any effect on apoptosis (Figure 2b). Even when we doubled the amount of plasmid, we did not detect an apoptotic response (not shown). Transient transfection and activation of H-ras, which also occurs in normal cells, is not sufficient for ORCTL3 to induce apoptosis (Figure 2c). As our sequencing of the endogenous H-ras gene (variants 1, 2, 3) from HEK293T cells did not reveal activating mutations at codons 12, 13 or 61 (data not shown), we speculated that apoptosis induction by ORCTL3 is not limited to H-ras-transformed cells. Hence, we assessed cell death by ORCTL3 in NRK cells transformed by v-src (Supplementary Figure 1C and D) and likewise observed apoptosis in these cells (Figure 2d and e). We then compared the ability of ORCTL3 to induce apoptosis in a number of transformed tumor cells and untransformed cells. Figure 3a shows that ORCTL3 can induce apoptosis in the human tumorigenic kidney cell line HEK293T, the human cervical HeLa cell line and in human LnCap and PC3 prostate cancer cells. ORCTL3 is also able to cause apoptosis in transformed baby hamster kidney (BHK) cells, but not in untransformed Madin–Darby canine kidney (MDCK) cells (Figure 3b). Moreover, ORCTL3 was likewise inactive for apoptosis induction in human umbilical vein endothelial cells (HUVEC) and primary human renal proximal tubule (RPT) cells.
Effect of ORCTL3 on apoptosis in transformed versus normal cells. (a) ORCTL3 induces apoptosis in various transformed tumor cells. Expression plasmids for ORCTL3, luciferase as a negative control and caspase-2 as a positive control, were transfected into the respective cell lines and apoptosis was quantified. The data for the human tumor cell lines HEK293T and PC3 are based on flow cytometry analysis and normalized to the transfection efficiency as determined by the percentage of transfected (GFP-positive) cells. The experiments were repeated three times independently. The data for the tumor cell lines HeLa, LnCap and BHK represent the ratio of morphologically apoptotic, GFP-positive (transfected) cells to all GFP-positive (transfected) cells. Cells were co-transfected at a ratio of 1 : 1 with a vector for GFP and an expression vector for ORCTL3, caspase-2 or luciferase. Each data point represents the mean of triplicate counts±standard deviation. Background apoptosis levels observed in luciferase-transfected cells were subtracted (P<0.01 for ORCTL3 relative to Luc-transfected cells in BHK and HeLa, P<0.05 in all other cell types). (b) ORCTL3 fails to provoke apoptosis in primary and untransformed cells. Primary HUVEC were transfected with an efficient electroporation method (Amaxa). Normal canine kidney cells (MDCK) were co-transfected with GFP and expression vectors (ORCTL3, caspase-2 or luciferase). Apoptosis in these cells was determined by phenotype quantification as in a (P-value nonsignificant in both cells types, n=3). Background apoptosis level of luciferase-transfected cells was subtracted. Apoptosis in primary renal cells (RPT) was quantified by the Vybrant Apotosis Assay Kit (P>0.05, n=3). Data are presented relative to the transfection efficiency determined by the percentage of GFP-transfected cells. As the pro-apoptotic Bax gene was a weak cell death inducer in these cells, they were treated with 1.5 mM H2O2 for 6 h as a positive control (Co is negative control)
To more closely relate the expression of ORCTL3 with the apoptotic response of the cells, we generated a C-terminal ECFP fusion protein of ORCTL3 and tested its apoptosis induction in HeLa cells, HEK293T cells, HUVEC and in WT and ras-transformed NRK cells. Figure 4a shows that only untransformed HUVEC and WT NRK cells did not succumb to apoptosis when this construct was transfected (see also Supplementary Figure 2). We also studied the expression level of the ORCTL3 fusion protein in the various cell types by measuring its fluorescence signal and normalizing this to the respective transfection efficiency. As Figure 4b reveals, the level of the ORCTL3-ECFP signal was comparable in all the cells. We also assayed the expression level of the endogenous ORCTL3 and compared this to the transfected ORCTL3 fusion protein in WT and ras-transformed NRK cells. Figure 4c shows the increase of the signal as detected in a semiquantitative RT-PCR.
Apoptosis assessment by an ORCTL3-ECFP fluorescent fusion protein. (a) Quantification of cell death in various cell lines induced 24 h after the transfection of the ORCTL3-ECFP plasmid. Cell death in 293T and HeLa was measured by FACS after PI staining, The percentage of apoptosis was normalized to the transfection efficiency as assessed by the CFP signal (P<0.01 for both cell types, n=4). HUVEC apoptosis was determined by counting fluorescent (transfected) cells with apoptotic morphology of the nucleus upon Hoechst staining (P-value nonsignificant, n=4). The cell death in NRK cells was quantified by cell morphology of fluorescent (transfected) cells (P-value nonsignificant in WT cells, P<0.01 in H-ras NRK cells, n=3). (b) Quantification of ORCTL3 expression in various cell types. The plasmid for ORCTL3-ECFP was transfected into WT and ras-transformed NRK cells (with JetPI Polyplus), 293T (with Superfect), HeLa (with Effectene) and HUVEC (with Amaxa kit). The fluorescence intensity of the ORCTL3-ECFP was measured 24 h after transfection using a plate reader. This intensity was normalized to the transfection efficiency measured by FACS analysis. The histograms are the result of two independent experiments (2 wells of a 24 well plate per experiment) (all crosswise comparisons P>0.05). (c) Upregulation of ORCTL3 upon transfection. Semiquantitative RT-PCR analysis in WT and ras-transformed NRK cell shows the gene expression level of ORCTL3 before transfection (NT for nontransfected) or after transfection with the pCDNA3-GFP (control) or the pCDNA3-ORCTL3-ECFP plasmid
Role of endoplasmic reticulum stress for ORCTL3 apoptosis induction
We studied the spatial distribution of the ECFP fusion protein of ORCTL3 with specific staining of organelles in HeLa cells. This fusion protein was found at the endoplasmic reticulum (ER), the Golgi and the plasma membrane (Figure 5), indicating that the ORCTL3 protein transport conforms to the known internal route of plasma membrane proteins. Importantly, we have not found an overlap with the signal from the mitochondria-specific probe, MitoTracker Red CMXRos. Because another tumor-specific apoptosis inducer the viral apoptin protein is differentially localized in normal versus tumor cells,3 we determined the localization of the ORCTL3-ECFP fusion protein in WT and ras-transformed NRK cells but found no difference in its cellular locale between these cell types (not shown). As ORCTL3 could cause cell death at intracellular sites during transit, we applied the pan-caspase inhibitor zVAD to block apoptosis and expected that those cells that accrue contain ORCTL3 at the relevant locale for apoptosis induction. Figure 6a shows that upon cell death inhibition the percentage of cells with a prominent intracellular accumulation of the ORCTL3-ECFP protein increased whereas the percentage of apoptotic cells decreased proportionally (see also Supplementary Figure 3). Hence, we speculated that ORCTL3 exerts its apoptosis activity at the ER or the Golgi. A fusion construct with an ER retention signal at the C terminus of ORCTL3 led to higher apoptosis induction than WT ORCTL3, whereas a fusion construct with a Golgi retention signal caused less apoptosis (Figure 6b). Consequently, we speculated that ORCTL3 is inducing the signal for apoptosis at the ER, an organelle with a well-recognized function in apoptosis. In support of this hypothesis, we found that co-transfection of the ER stress inhibitor BiP/GRP78 caused a reduction in apoptosis (Figure 6c). ER stress is a complex cellular response that leads to the upregulation of both pro- and anti-apoptotic genes. We determined the protein levels of the ER stress factors ATF4/CREB2 as a pro-apoptotic protein, and BiP as an anti-apoptotic protein in ORCTL3-transfected cells. In these experiments, we used the ER stressors tunicamycin or ionomycin as controls whose apoptotic effects are influenced by the expression level of BiP.23, 24 Our results show that in cells that undergo apoptosis upon ORCTL3 expression (HeLa and HEK293T cells), ATF4 was upregulated to an extent comparable with control-treated cells. BiP, on the other hand, did not increase when compared to tunicamycin- or ionomycin-treated cells (Figure 6d and e). In contrast, HUVECs that do not succumb to apoptosis upon ORCTL3 transfection failed to show any signs of ER stress, with neither BiP nor ATF4 being upregulated (Figure 6f). Transfection efficiency in these experiments, as determined by GFP-transfected cells, was 40% in HeLa cells, 60% in HEK293T cells and 65% in HUVEC (not shown). Treatment with the ER stress inducer tunicamycin could also increase BiP in HUVEC (Figure 6f) and did not indicate that HeLa cells are more susceptible to ER stress than HUVEC (Supplementary Figure 4).
ORCTL3 localizes to the cell membrane, ER and Golgi, but not to mitochondria. HeLa cells were plated on coverslips, transfected with an ORCTL3-ECFP expression vector and incubated with zVAD. Cells were processed as described under Materials and Methods section. The overlay of the fluorescence microscopic image of CFP and the histochemical staining indicated the localization of the fusion protein
ORCTL3 exerts an ER stress signal. (a) Effect of apoptosis inhibition on the percentage of cells with apoptosis morphology and with different degrees of internal accumulation of ORCTL3. HeLa cells were transfected with ORCTL3-ECFP expression vector using the Effectene transfection reagent (Qiagen) and grown either in the presence (100 μ M) or absence of the pan-caspase inhibitor zVAD. Cellular localization of the fusion protein in the presence or absence of apoptosis inhibition was ascertained 24 h after transfection using a fluorescent microscope (FITC filter). The data in the graph represent the percentage of HeLa cells with a prominent internal accumulation of the ORCTL3 fusion protein (P<0.01), a balanced distribution of the ORCTL3 fusion protein on internal and membrane sites (P-value nonsignificant) or apoptotic cells (P<0.05). The inhibitory effect of zVAD on apoptosis is shown in percentage of apoptotic cells, ±zVAD. The data represent the mean of triplicate counts (300 cells per count)±standard deviation. (b) An ORCTL3 construct with an ER retention signal (ORCTL3-ER) is more efficient for apoptosis induction than wild-type ORCTL3 or a fusion construct with a Golgi retention signal (ORCTL3-G). ORCTL3 was fused to either an ER or a Golgi retention signal and its apoptosis induction was determined by phenotype quantification after co-transfection with GFP in HeLa cells (P<0.01 for ORCTL3-ER in relation to ORCTL3 and ORCTL3-G, n=3). (c) BiP co-transfection reduces apoptosis by ORCTL3. HeLa cells were transfected with ORCTL3 and expression constructs for Luc or BiP at a ratio of 1 : 2.7 and scored after 24 h (P<0.05, n=3). (d–f) Western immunoblotting showing the expression levels of the ER stress transcription factor ATF4 and of the chaperone protein BiP in HeLa cells, HEK293T cells or HUVEC. (d) Western immunoblots reveal the protein levels of ATF4 (left panel) and BiP (middle panel) in HeLa cells at various time points after transfection of ORCTL3 or luciferase (Luc). Ionomycin (Iono, 5 μ M)-treated cell lysates were used as a control for BiP. The protein levels of ATF4 and BiP were also detected in control HeLa cell extracts treated with the indicated concentrations of tunicamycin for given time points (right panel). Similar experiments were conducted with HEK293T (e) with 2.5 μ M tunicamycin and in HUVEC (f). These also included lysates of ORCTL3-ER-transfected cells. β-Actin was used as a loading control in all experiments
ORCTL3 is downregulated in human kidney tumors
A broader function of ORCTL3 during tumorigenesis was suggested as deletions in its chromosomal locus 3p21.3 have been linked to carcinomas of various organs.25, 26 We found microarray data with significant P-values from the Oncomine database indicating that this gene is repressed in tumors of the adrenal gland and the bladder (not shown). Importantly, using the Cancer Profiling Array, a blot that compares tumor versus normal tissue of the same patients, we detected that ORCTL3 is significantly downregulated in human renal carcinomas (Figure 7).
ORCTL3 is downregulated in human renal tumors. A commercial filter blot (Cancer Profiling Array; Clontech) with matched RNA of tumor and normal tissue from each patient was probed with ORCTL3. The signal intensities were measured with densitometry and the percentage values of the tumor signals relative to the normal tissues were calculated for each patient and plotted (P<0.01, n=20). No signals were detected in other tissues, most likely due to reduced expression levels
Discussion
In this study, we have presented an assay to systematically search the genome for genes that induce apoptosis specifically in tumor cells. This is achieved by two consecutive assays that culminate in a synthetic lethal screen, which directly compares the activities of genes in cells transformed by specific oncogenes versus their normal counterparts (Figure 1a). The concept of synthetic lethality has recently gained wide recognition as a useful tool in cancer research.27 Its integration into our assay has the potential to reveal pro-apoptotic signaling pathways specific for certain oncogenes, such as H-ras. We observed that ORCTL3, our first isolate from this screen, is likewise active in H-ras- and v-src-transformed cells, and in several other cell lines of different genetic background (Figures 2 and 3). This suggests that at least one aspect of the transformation by H-ras is shared with other tumor cells and that this exposes cells to particular pro-apoptotic signaling pathways, one of which is targeted by ORCTL3.
With ORCTL3 being a member of the transporter gene superfamily, we expected its putative transporter activity to be involved in its ability to induce apoptosis. On the basis of our deletion studies (Figure 2a), however, we do not believe that it is working directly as a transporter for apoptosis induction. Our data indicate that ORCTL3 causes ER stress in cells undergoing apoptosis. ER stress is induced by the accumulation of unfolded proteins within the ER, but also by numerous additional signals that lead to the disruption of ER function.28 It is believed that ER stress first triggers an anti-apoptotic response in the cells with upregulation of chaperones such as BiP. However, if the stimulus persists, this is then followed by a pro-apoptotic reaction. The signal induced by ORCTL3 seems to lead to an unconventional ER stress response. Although BiP is normally strongly upregulated as part of the anti-apoptotic response during ER stress, under the conditions used we have not observed an increase of the BiP protein upon ORCTL3 transfection comparable to tunicamycin-treatment. In contrast, the pro-apoptotic ATF4 transcription factor is upregulated upon ORCTL3 overexpression (Figure 6). A reduced BiP response despite ER stress induction is not without precedent. It was found in WEHI7.1 cells upon thapsigargin treatment, which led to efficient apoptosis induction.29 As specific upregulation of BiP by co-transfection can inhibit ORCTL3-induced apoptosis (Figure 6c), the reduced increase of BiP could be an important aspect of ORCTL3's activity in long-established tumor cell lines, which can be expected to have accumulated numerous mutations that thwart cell death and support the anti-apoptotic response during ER stress. This per se, however, is unlikely to explain the tumor-specific signal of ORCTL3 as BiP knockout has been shown to cause apoptosis in untransformed cells.30 Our data indicate that apoptosis upon ER stress by tunicamycin is not more efficiently induced in HeLa cells compared to HUVEC (Supplementary Figure 4), which argues against ORCTL3 exploiting a general difference in the sensitivity to ER stress in normal and transformed cells. Hence, we believe that ORCTL3 induces a signal that is interpreted as pro-apoptotic ER stress in tumor cells but not in normal cells. Interestingly, another tumor-specific gene, mda-7/IL-24, has recently been implicated in ER stress as it localizes to the ER and interacts with BiP.31 Future studies will need to address the precise differences in signaling pathways between normal and tumor cells, which allow ORCTL3 to induce tumor-specific ER stress.
ORCTL3 seems to be involved in tumorigenesis as indicated by its downregulation in renal tumors (Figure 7). It may also be downregulated in additional cancers as it is expressed at low levels also in tissues other than the kidney,20 but our commercial filter blot was not sensitive enough to detect this. We assume that even at its endogenous expression level this gene can be activated for cell destruction during tumorigenesis and that its downregulation allows for continued survival of cancer cells. This qualifies ORCTL3 as a candidate for a tumor suppressor gene and demonstrates that our screen can also serve as a novel tool to determine such activities.
How many genes with a tumor-specific activity can be expected? The analysis so far revealed one cDNA library as tumor specific. Although our current screen is still limited to cellular genes, viral DNAs, cDNA fragments encoding protein domains or other genetic means such as shRNAs could potentially expand the positive hits in this screen. Additional screens could target oncogenes that, in contrast to H-ras, do not exert an anti-apoptotic effect or genetic changes specific for HEK293 cells such as the viral genes E1A and E1B.32 Moreover, because tumorigenesis is a multistep process, one could introduce additional genetic lesions in already transformed cells against which the synthetic lethal screen is directed. Hence, our approach could be of wider use to uncover signals that specifically induce the demise of transformed cells.
Materials and Methods
Materials
The pan-caspase inhibitor zVAD-fmk was purchased from Enzyme Systems Products. Cross-linked polyethylenimine was a kind gift from Dr. Klibanov, Cambridge, USA. The ER-Tracker Blue-White DPX, BODIPY TR C5 Ceramide Golgi-tracker dye, MitoTracker Red CMXRos, and Vybrant Apotosis Assay Kit#4 were from Molecular Probes, Invitrogen. All fine chemicals were from Sigma Inc. unless specified otherwise. The clones for SLC22A and SLC22A12 were obtained from the IMAGE clone collection through the GeneService (Cambridge, UK).
Statistics
The t-test was used to assess the statistical significance of the data. If P>0.1 we state that the P-value was nonsignificant. Experiments with n=3 were repeated at least once and one representative result is shown.
Cell culture
HEK293T,33 HeLa (ATCC no.: CCL-2) and NRK cells (CRL-6509) were cultured in DMEM (Sigma) supplemented with 5% (HEK293T, NRK) or 10% (HeLa) FCS (Sigma). HUVEC and RPT cells (Lonza, Basel, Switzerland) were grown in Endothelial Growth Medium Bulletkit (Lonza) and Renal Epithelial Growth Medium Bulletkit (Lonza), respectively. HUVEC were transfected using HUVEC Nucleofector kit (Amaxa), whereas RPT cells were transfected using basic nucleofector kit for Primary Mammalian Epithelial Cells (Amaxa). HEK293T, HeLa cells and NRK cells were transfected with Superfect (Qiagen), Effectene (Qiagen) and Fugene (Roche, Mannheim, Germany), respectively, according to manufacturer's instructions. For stable transfections, NRK cells were selected with 500 μg/ml G418 (Calbiochem). PC3 (CRL-1435) were grown in RPMI 10% FCS and transfected with IBAfect/MA Lipofection Enhancer (Promokine, Heidelberg, Germany). BHK cells (CCL-10) were grown in DMEM, 5% FCS and transfected using calcium phosphate. LnCap (CRL-1740) were grown in RPMI, 10% FCS and transfected with Lipofectamine 2000 (Invitrogen). MDCK (CCL-34) were maintained in DMEM, 10% FCS and transfected using cross-linked polyethylenimine.34
Fluorescent imaging
HeLa cells were grown on coverslips for 24 h, transfected with an ORCTL3-ECFP expression vector and incubated for another 24 h with zVAD to avoid secondary effects due to apoptosis induction. For nuclear staining, cells were washed twice in PBS and incubated in 4% formaldehyde for 20 min at room temperature. Cells were then incubated in 4′,6-diamidino-2-phenylindole (DAPI) solution for 1 h in the dark at room temperature. Before the other organelle stainings, cells were washed in 1 × PBS and then incubated in 600 nM ER-Tracker Blue-White DPX, 5 μ M BODIPY TR C5 Ceramide Golgi-tracker dye or 40 nM MitoTracker Red CMXRos. Cells were rinsed three times in ice-cold growth medium and incubated in fresh full growth medium at 37°C for a further 30 min. Cells were then washed twice in 1 × PBS and incubated for 20 min at room temperature with 4% formaldehyde. Coverslips were mounted on microscope slides. Fluorescence was detected with FITC, DAPI and TRITC filters of a fluorescent microscope (Zeiss) at magnification × 63, using a Zeiss Axiovert camera. WT and ras-transformed NRK cells were stained using the same protocol as for HeLa cells.
DNA constructs
The H-ras and the v-src construct that were used to generate stably transfected NRK cells have been described.35 For the deletion mutants suitable primers were used with the Expand Long Template PCR System (Roche Diagnostics), which has proofreading activity. Expression plasmids were constructed by inserting the cDNAs into the vector pcDNA3 (Invitrogen). All PCR products were sequenced to verify the correct sequence. BiP was amplified using suitable primers with the Phusion high-fidelity DNA polymerase (Finnzymes) and subcloned into the pcDNA3 vector. Fusion constructs of ORCTL3 in the mammalian expression vector pcDNA3 were generated with eight-amino-acid ER retention sequence36 and Golgi retention signal37 derived from the G1 protein of UUK virus, which was a kind gift from Dr. Pettersson (Karolinska Institute, Sweden). ORCTL3-ECFP was generated as a fusion construct with a pcDNA3 vector backbone. Plasmid DNA was isolated using Qiagen maxi-prep kit according to the manufacturer's instructions. The ORCTL3 sequence was identified with the IMAGE 4609915 cDNA clone.
Apoptosis detection
Apoptosis was quantified by four different methods. The first involved determining the ratio of transfected (GFP positive) and phenotypically apoptotic cells (displaying volume loss and/or membrane blebbing) in relation to all GFP-positive and therefore transfected cells. At least 300 cells were counted in each independent experiment. The second method involved measuring nucleosomal DNA fragments by an ELISA kit (Roche Diagnostics). The third method utilized sub-G1 DNA content determined by flow cytometry in transfected cells as described.11 Finally, apoptosis in human renal proximal tubular epithelial cells (RPT) was quantified using the Vybrant Apotosis Assay Kit#4 (Molecular Probes, Invitrogen) and analyzed by flow cytometry according to the manufacturer's instructions.
Immunoblotting
Whole cell lysates were isolated using RIPA buffer supplemented with protease inhibitors (Complete Mini; Roche). Each sample (40 μg) was separated in a 10 or 12% SDS-PAGE gel and electrophoretically transferred onto polyvinylidene or nitrocellulose membranes (Amersham Pharmacia). The membranes were blocked with 5% nonfat dry milk powder in 0.1% Tween-20 (Sigma) and incubated with antibodies against H-ras (Santa Cruz), src (Oncogene), ATF4/CREB2 (C-20; Santa Cruz), BiP/GRP78 (N-20; Santa Cruz) or β-actin (Sigma). The membranes were then washed three times in 0.1% Tween/1 × PBS and incubated in appropriate secondary antibodies. After additional washes, the reactions were developed by enhanced chemiluminescence reagents (Pierce or Amersham Biosciences).
Isolation of H-ras from HEK293T cells
RNA was isolated from HEK293T cells using 1 ml Trizol (Invitrogen) per 10 cm2 dish of cells and resuspended in 20 μl ultrapure H2O. The Titan One Tube RT-PCR kit (Roche) was used to carry out RT-PCR using 2 μg total RNA. Specific H-ras primers (fwd primer for variant 1, 2 and 3: ACTGTAAGCTTCCGCCATGACGGAATATAAGCTGG), (rev primer for variant 1 and 3: ACTGTGATATCTCAGGAGAGCACACACTTG), (rev primer for variant 2: ACTGTGATATCTCACATGGGTCCCGGGGGG) were used for PCR reactions. Twenty-nine PCR cycles were used (10 s at 94°C, 30 s at 54°C and 50 s at 68°C; followed by 7 min at 68°C). PCR products were separated by gel electrophoresis and purified using the Wizard SV Gel and PCR Clean-Up System (Promega). Purified PCR products and pCDNA3 plasmid were then cut using EcoRV and HindIII, analyzed with agarose gel electrophoresis and again purified using the Wizard SV Gel and PCR Clean-Up System. T4 DNA Ligase (Promega) was used to ligate the cut PCR product and the pCDNA3 plasmid DNA together and the constructs were transformed into competent DH5α cells. Colonies were picked for bacterial culture using LB with ampicillin (50 μg/ml). Mini-prep sequencing was performed using T7 (forward) and BGH (reverse) sequencing primers at the MRC CSC Genomics Core Laboratory at Imperial College London, Hammersmith Campus. Two clones per ras variant were analyzed in this way. Sequences were analyzed using BLAST on the NCBI Web site.
Quantification of ORCTL3 expression level by fluorescence
The plasmid pCDNA3-ORCTL3-ECFP was transfected in duplicates in 24-well plates with the various cells lines. Each experiment was performed twice. At 24 h after the transfection, the FLUOstar OPTIMA plate reader (BMG Labtech) measured the fluorescence intensity emitted by each well at 520 nm. Then the cells from the same well were analyzed by flow cytometry. The FL1 channel was used to detect the number of cells expressing ORCTL3-ECFP. The fluorescence intensity measured by the plate reader was then normalized to the percentage of transfected cells determined by flow cytometry.
Quantification of ORCTL3 expression levels by reverse transcription PCR
WT and ras-transformed NRK cells were transfected (JetPI Polyplus) or not by the following vectors: pCDNA3-ORCTL3-ECFP or pLantern-GFP, 24 h before the extraction of the RNA using TRIzol reagent (Invitrogen). The RNA was applied directly to the Titan One Tube RT-PCR kits (Roche Applied Sciences) using AMV reverse transcriptase for first strand cDNA synthesis and the Expand High-Fidelity enzyme consisting of Taq DNA plymerase and Tgo DNA polymerase for amplification of the cDNAs by PCR. All reactions were carried out following the procedures described by the manufacturers. ORCTL3 expression was analyzed in a 1% agarose gel, after amplification of a 477-nt fragment with the following oligonucleotides:
5′-CCAGCTTTGAGCTCTA-3′ and 5′-CTGTCCACAAACCAGACA-3′.
The β-actin gene was amplified in parallel with the following oligonucleotides: 5′-GCTCGTCGTCGACAACGGCTC-3′ and 5′-CAAACATGATCTGGGTCATCT-3′ and used as a loading control.
Blot array hybridization
For hybridization of the Matched Tumor/Normal Profiling Array (Clontech), the coding sequence of ORCTL3 was labeled with 5′-[α32P]-dCTP using the RediPrime random prime labeling kit (Amersham) with 200 ng plasmid and 50 μCi [32P]dCTP. The blot was prehybridized for 3 h at 65°C in 40 ml of hybridization buffer (6 × SSC, 5 × Denhardt's reagent, 0.5% SDS, 0.1 mg/ml salmon sperm DNA), then 10 ml of a 50% dextran sulfate solution and finally the labeled probe (approximately 3 × 107 c.p.m.) was added to the hybridization solution and incubated at 62°C for 6 h. The blot was washed with buffer 1 (2 × SSC, 0.5% SDS), buffer 2 (1 × SSC, 0.5% SDS) and buffer 3 (0.5 × SSC, 0.5% SDS), exposed in a BAS 2500 phosphoimager screen (Fujifilms, Heidelberg, Germany) and the signals were quantified using the Image Gauge V3.01 software.
Abbreviations
- NRK:
-
normal rat kidney
- WT:
-
wild type
- OCT:
-
organic cation transporter
- ORCTL3:
-
organic cation transporter-like 3
- BHK:
-
baby hamster kidney
- HUVEC:
-
human umbilical vein endothelial cells
- ER:
-
endoplasmic reticulum
- MDCK:
-
Madin–Darby canine kidney
References
Debatin KM, Poncet D, Kroemer G . Chemotherapy: targeting the mitochondrial cell death pathway. Oncogene 2002; 21: 8786–8803.
Shtrichman R, Sharf R, Barr H, Dobner T, Kleinberger T . Induction of apoptosis by adenovirus E4orf4 protein is specific to transformed cells and requires an interaction with protein phosphatase 2A. Proc Natl Acad Sci USA 1999; 96: 10080–10085.
Danen-Van Oorschot AA, Fischer DF, Grimbergen JM, Klein B, Zhuang S, Falkenburg JH et al. Apoptin induces apoptosis in human transformed and malignant cells but not in normal cells. Proc Natl Acad Sci USA 1997; 94: 5843–5847.
Su ZZ, Madireddi MT, Lin JJ, Young CS, Kitada S, Reed JC et al. The cancer growth suppressor gene mda-7 selectively induces apoptosis in human breast cancer cells and inhibits tumor growth in nude mice. Proc Natl Acad Sci USA 1998; 95: 14400–14405.
Svensson M, Hakansson A, Mossberg AK, Linse S, Svanborg C . Conversion of alpha-lactalbumin to a protein inducing apoptosis. Proc Natl Acad Sci USA 2000; 97: 4221–4226.
Nalca A, Qiu SG, El-Guendy N, Krishnan S, Rangnekar VM . Oncogenic Ras sensitizes cells to apoptosis by Par-4. J Biol Chem 1999; 274: 29976–29983.
Walczak H, Miller RE, Ariail K, Gliniak B, Griffith TS, Kubin M et al. Tumoricidal activity of tumor necrosis factor-related apoptosis-inducing ligand in vivo. Nat Med 1999; 5: 157–163.
Grimm S . The art and design of genetic screens: mammalian culture cells. Nat Rev Genet 2004; 5: 179–189.
Numa F, Hirabayashi K, Tsunaga N, Kato H, O’Rourke K, Shao H et al. Elevated levels of syndecan-1 expression confer potent serum-dependent growth in human 293T cells. Cancer Res 1995; 55: 4676–4680.
Albayrak T, Grimm S . A high-throughput screen for single gene activities: isolation of apoptosis inducers. Biochem Biophys Res Commun 2003; 304: 772–776.
Bauer MKA, Schubert A, Rocks O, Grimm S . Adenine nucleotide translocase-1, a component of the permeability transition pore, can dominantly induce apoptosis. J Cell Biol 1999; 147: 1493–1502.
Schoenfeld N, Bauer MK, Grimm S . The metastasis suppressor gene C33/CD82/KAI1 induces apoptosis through reactive oxygen intermediates. FASEB J 2004; 18: 158–160.
Albayrak T, Scherhammer V, Schoenfeld N, Braziulis E, Mund T, Bauer MK et al. The tumor suppressor cybL, a component of the respiratory chain, mediates apoptosis induction. Mol Biol Cell 2003; 14: 3082–3096.
Gewies A, Grimm S . UBP41 is a proapoptotic ubiquitin-specific protease. Cancer Res 2003; 63: 682–688.
Mund T, Gewies A, Schoenfeld N, Bauer MK, Grimm S . Spike, a novel BH3-only protein, regulates apoptosis at the endoplasmic reticulum. FASEB J 2003; 17: 696–698.
Bos JL . ras oncogenes in human cancer: a review. Cancer Res 1989; 49: 4682–4689.
Khwaja A, Rodriguez-Viciana P, Wennstrom S, Warne PH, Downward J . Matrix adhesion and Ras transformation both activate a phosphoinositide 3-OH kinase and protein kinase B/Akt cellular survival pathway. EMBO J 1997; 16: 2783–2793.
Hanahan D, Weinberg RA . The hallmarks of cancer. Cell 2000; 100: 57–70.
Best CJ, Tanzer LR, Phelps PC, Merriman RL, Boder GG, Trump BF et al. H-ras-transformed NRK-52E renal epithelial cells have altered growth, morphology, and cytoskeletal structure that correlates with renal cell carcinoma in vivo. In Vitro Cell Dev Biol Anim 1999; 35: 205–214.
Nishiwaki T, Daigo Y, Tamari M, Fujii Y, Nakamura Y . Molecular cloning, mapping, and characterization of two novel human genes, ORCTL3 and ORCTL4, bearing homology to organic-cation transporters. Cytogenet Cell Genet 1998; 83: 251–255.
Lee W, Kim RB . Transporters and renal drug elimination. Annu Rev Pharmacol Toxicol 2004; 44: 137–166.
Bahn A, Hagos Y, Reuter S, Balen D, Brzica H, Krick W et al. Identification of a new urate and high affinity nicotinate transporter, hOAT10 (SLC22A13). J Biol Chem 2008; 283: 16332–16341.
Jiang CC, Chen LH, Gillespie S, Wang YF, Kiejda KA, Zhang XD et al. Inhibition of MEK sensitizes human melanoma cells to endoplasmic reticulum stress-induced apoptosis. Cancer Res 2007; 67: 9750–9761.
Miyake H, Hara I, Arakawa S, Kamidono S . Stress protein GRP78 prevents apoptosis induced by calcium ionophore, ionomycin, but not by glycosylation inhibitor, tunicamycin, in human prostate cancer cells. J Cell Biochem 2000; 77: 396–408.
Imreh S, Klein G, Zabarovsky ER . Search for unknown tumor-antagonizing genes. Genes Chromosomes Cancer 2003; 38: 307–321.
Zabarovsky ER, Lerman MI, Minna JD . Tumor suppressor genes on chromosome 3p involved in the pathogenesis of lung and other cancers. Oncogene 2002; 21: 6915–6935.
Kaelin Jr WG . The concept of synthetic lethality in the context of anticancer therapy. Nat Rev Cancer 2005; 5: 689–698.
Rao RV, Ellerby HM, Bredesen DE . Coupling endoplasmic reticulum stress to the cell death program. Cell Death Differ 2004; 11: 372–380.
McCormick TS, McColl KS, Distelhorst CW . Mouse lymphoma cells destined to undergo apoptosis in response to thapsigargin treatment fail to generate a calcium-mediated grp78/grp94 stress response. J Biol Chem 1997; 272: 6087–6092.
Luo S, Mao C, Lee B, Lee AS . GRP78/BiP is required for cell proliferation and protecting the inner cell mass from apoptosis during early mouse embryonic development. Mol Cell Biol 2006; 26: 5688–5697.
Gupta P, Walter MR, Su ZZ, Lebedeva IV, Emdad L, Randolph A et al. BiP/GRP78 is an intracellular target for MDA-7/IL-24 induction of cancer-specific apoptosis. Cancer Res 2006; 66: 8182–8191.
Jones NC . Transformation by the human adenoviruses. Semin Cancer Biol 1990; 1: 425–435.
DuBridge RB, Tang P, Hsia HC, Leong PM, Miller JH, Calos MP . Analysis of mutation in human cells by using an Epstein-Barr virus shuttle system. Mol Cell Biol 1987; 7: 379–387.
Thomas M, Ge Q, Lu JJ, Chen J, Klibanov AM . Cross-linked small polyethylenimines: while still nontoxic, deliver DNA efficiently to mammalian cells in vitro and in vivo. Pharm Res 2005; 22: 373–380.
Grimm S, Baeuerle PA . Failure of the splicing variant p65 delta of the NF-kappa B subunit p65 to transform fibroblasts. Oncogene 1994; 9: 2391–2398.
Zarei MM, Eghbali M, Alioua A, Song M, Knaus HG, Stefani E et al. An endoplasmic reticulum trafficking signal prevents surface expression of a voltage- and Ca2+-activated K+ channel splice variant. Proc Natl Acad Sci USA 2004; 101: 10072–10077.
Andersson AM, Pettersson RF . Targeting of a short peptide derived from the cytoplasmic tail of the G1 membrane glycoprotein of Uukuniemi virus (Bunyaviridae) to the Golgi complex. J Virol 1998; 72: 9585–9596.
Acknowledgements
This work was supported by a grant from Cancer Research UK to SI. A-L M-M was supported by Breast Cancer Campaign, NK by the EU grant ‘Hermione’ and AL by a grant from Cancer Research UK. We thank Dr. W Abdul Salam for advice on statistical evaluation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by RA Knight
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Rights and permissions
About this article
Cite this article
Irshad, S., Mahul-Mellier, AL., Kassouf, N. et al. Isolation of ORCTL3 in a novel genetic screen for tumor-specific apoptosis inducers. Cell Death Differ 16, 890–898 (2009). https://doi.org/10.1038/cdd.2009.21
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2009.21
Keywords
This article is cited by
-
Genetic screening for anticancer genes highlights FBLN5 as a synthetic lethal partner of MYC
Cell Communication and Signaling (2023)
-
The anticancer gene ORCTL3 targets stearoyl-CoA desaturase-1 for tumour-specific apoptosis
Oncogene (2015)
-
Cell death assays for drug discovery
Nature Reviews Drug Discovery (2011)