Abstract
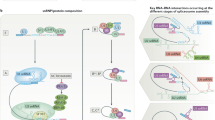
In order to determine the potential of alternative splicing as a means of targeting the expression of therapeutic genes to tumor cells in vivo, a series of episomal plasmid-based “splice-activated gene expression” (pSAGE) vectors was generated, which contain minigene cassettes composed of various combinations of the three alternatively spliced exons present in the differentially expressed adhesion protein CD44R1 (v8, v9, and v10) with or without their corresponding intronic sequences, positioned in-frame between the CD44 leader sequence and a “leaderless” human liver/bone/kidney alkaline phosphatase (ALP) cDNA. Because both the v8–v9 and v9–v10 introns contain multiple in-frame stop codons, the expression and enzymatic activity of ALP are dependent upon the accurate removal of intronic sequences from the pre-mRNA transcripts encoded by these constructs. The various pSAGE constructs were introduced into CD44H-positive (T24) and CD44R1-positive (PC3) target cells by electroporation and transfectants selected in hygromycin B. ALP expression was determined by staining with the ALP substrate, BCIP/INT, and the transfected cells tested for their sensitivity to the inactive prodrug, etoposide phosphate. ALP-mediated dephosphorylation of etoposide phosphate generates the potent topoisomerase II inhibitor etoposide. The data obtained indicate that whereas the v8–v9 intron is spliced in both CD44H- and CD44R1-positive cells, the v9–v10 intron is efficiently and accurately removed only in CD44R1-positive cells. Furthermore, only CD44R1-positive cells were sensitized to etoposide phosphate when transfected with the v9–v10.ALP construct. These data emphasize the potential usefulness of alternative splicing as a novel means of targeting gene expression to tumor cells in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Brenner MK . Gene transfer and the treatment of haematological malignancy J Intern Med 2001 249: 345–358
Dougherty GJ, Chaplin DJ, Dougherty ST et al. In vivo gene therapy of cancer Tumor Targeting 1996 2: 1–10
Dougherty GJ, Peters CE, Dougherty ST et al. Gene therapy–based approaches to the treatment of cancer. Development of targetable retroviral vectors Transfus Sci 1996 17: 121–128
Galanis E, Vile R, Russell SJ . Delivery systems intended for in vivo gene therapy of cancer: targeting and replication competent viral vectors Crit Rev Oncol Hematol 2001 38: 177–192
Dachs GU, Patterson AV, Firth JD et al. Targeting gene expression to hypoxic tumor cells Nat Med 1997 3: 515–520
Dachs GU, Dougherty GJ, Stratford IJ et al. Targeting gene therapy to cancer: a review Oncol Res 1997 9: 313–325
Cooper DL, Dougherty GJ . To metastasize or not? Selection of CD44 splice sites Nat Med 1995 1: 635–637
Dreyfuss G, Hentze M, Lamond AI . From transcript to protein Cell 1996 85: 963–972
Lopez AJ . Alternative splicing of pre-mRNA: developmental consequences and mechanisms of regulation Annu Rev Genet 1998 32: 279–305
Smith CW, Patton JG, Nadal-Ginard B . Alternative splicing in the control of gene expression Annu Rev Genet 1989 23: 527–577
Cooper TA, Mattox W . The regulation of splice-site selection, and its role in human disease Am J Hum Genet 1997 61: 259–266
Naot D, Sionov RV, Ish-Shalom D . CD44: structure, function, and association with the malignant process Adv Cancer Res 1997 71: 241–319
Lesley J, Hyman R, Kincade PW . CD44 and its interaction with extracellular matrix Adv Immunol 1993 54: 271–335
Finn L, Dougherty G, Finley G et al. Alternative splicing of CD44 pre-mRNA in human colorectal tumors Biochem Biophys Res Commun 1994 200: 1015–1022
Sneath RJ, Mangham DC . The normal structure and function of CD44 and its role in neoplasia Mol Pathol 1998 51: 191–200
Ghaffari S, Smadja-Joffe F, Oostendorp R et al. CD44 isoforms in normal and leukemic hematopoiesis Exp Hematol 1999 27: 978–993
Herrera-Gayol A, Jothy S . Adhesion proteins in the biology of breast cancer: contribution of CD44 Exp Mol Pathol 1999 66: 149–156
Senter PD, Saulnier MG, Schreiber GJ et al. Anti-tumor effects of antibody–alkaline phosphatase conjugates in combination with etoposide phosphate Proc Natl Acad Sci USA 1988 85: 4842–4846
Senter PD, Schreiber GJ, Hirschberg DL et al. Enhancement of the in vitro and in vivo antitumor activities of phosphorylated mitomycin C and etoposide derivatives by monoclonal antibody–alkaline phosphatase conjugates Cancer Res 1989 49: 5789–5792
Hande KR . Etoposide: four decades of development of a topoisomerase II inhibitor Eur J Cancer 1998 34: 1514–1521
Kaighn ME, Lechner JF, Narayan KS et al. Prostate carcinoma: tissue culture cell lines Natl Cancer Inst Monogr 1978 17–21
Kaighn ME, Narayan KS, Ohnuki Y et al. Establishment and characterization of a human prostatic carcinoma cell line (PC-3) Invest Urol 1979 17: 16–23
O'Toole C, Perlmann P, Unsgaard B et al. Cellular immunity to human urinary bladder carcinoma: I. Correlation to clinical stage and radiotherapy Int J Cancer 1972 10: 77–91
O'Toole C, Unsgaard B, Almgard LE et al. The cellular immune response to carcinoma of the urinary bladder: correlation to clinical stage and treatment Br J Cancer 1973 28: Suppl 1 266–275
Dougherty GJ, Cooper DL, Memory JF et al. Ligand binding specificity of alternatively spliced CD44 isoforms. Recognition and binding of hyaluronan by CD44R1 J Biol Chem 1994 269: 9074–9078
Droll A, Dougherty ST, Chiu RK et al. Adhesive interactions between alternatively spliced CD44 isoforms J Biol Chem 1995 270: 11567–11573
Ghaffari S, Dougherty GJ, Lansdorp PM et al. Differentiation-associated changes in CD44 isoform expression during normal hematopoiesis and their alteration in chronic myeloid leukemia Blood 1995 86: 2976–2985
Ghaffari S, Dougherty GJ, Eaves AC et al. Altered patterns of CD44 epitope expression in human chronic and acute myeloid leukemia Leukemia 1996 10: 1773–1781
Dougherty GJ, Landorp PM, Cooper DL et al. Molecular cloning of CD44R1 and CD44R2, two novel isoforms of the human CD44 lymphocyte “homing” receptor expressed by hemopoietic cells J Exp Med 1991 174: 1–5
Melton RG, Sherwood RF . Antibody–enzyme conjugates for cancer therapy J Natl Cancer Inst 1996 88: 153–165
Ruffner KL, Matthews DC . Current uses of monoclonal antibodies in the treatment of acute leukemia Semin Oncol 2000 27: 531–539
Shabat D, Lode HN, Pertl U et al. In vivo activity in a catalytic antibody–prodrug system: antibody catalyzed etoposide prodrug activation for selective chemotherapy Proc Natl Acad Sci USA 2001 98: 7528–7533
Brandt DW, Wachsman W, Deftos LJ . Parathyroid hormone-like protein: alternative messenger RNA splicing pathways in human cancer cell lines Cancer Res 1994 54: 850–853
Moolenaar CE, Pieneman C, Walsh FS et al. Alternative splicing of neural-cell-adhesion molecule mRNA in human small-cell lung-cancer cell line H69 Int J Cancer 1992 51: 238–243
Itoh H, Hattori Y, Sakamoto H et al. Preferential alternative splicing in cancer generates a K-sam messenger RNA with higher transforming activity Cancer Res 1994 54: 3237–3241
Reale MA, Hu G, Zafar AI et al. Expression and alternative splicing of the deleted in colorectal cancer (DCC) gene in normal and malignant tissues Cancer Res 1994 54: 4493–4501
Xu CF, Chambers JA, Nicolai H et al. Mutations and alternative splicing of the BRCA1 gene in UK breast/ovarian cancer families Genes, Chromosomes Cancer 1997 18: 102–110
Saito H, Nakatsuru S, Inazawa J et al. Frequent association of alternative splicing of NER, a nuclear hormone receptor gene in cancer tissues Oncogene 1997 14: 617–621
Carstens RP, Eaton JV, Krigman HR et al. Alternative splicing of fibroblast growth factor receptor 2 (FGF-R2) in human prostate cancer Oncogene 1997 15: 3059–3065
Genuardi M, Viel A, Bonora D et al. Characterization of MLH1 and MSH2 alternative splicing and its relevance to molecular testing of colorectal cancer susceptibility Hum Genet 1998 102: 15–20
Sato H, Hiyama K, Ishioka S et al. Alternative splicing, but not allelic loss, of the FHIT gene increases with development of lung cancer Int J Oncol 1999 15: 81–88
Kwabi-Addo B, Ropiquet F, Giri D et al. Alternative splicing of fibroblast growth factor receptors in human prostate cancer Prostate 2001 46: 163–172
Mercatante DR, Bortner CD, Cidlowski JA et al. Modification of alternative splicing of Bcl-x pre-mRNA in prostate and breast cancer cells. Analysis of apoptosis and cell death J Biol Chem 2001 276: 16411–16417
Lukas J, Gao DQ, Keshmeshian M et al. Alternative and aberrant messenger RNA splicing of the mdm2 oncogene in invasive breast cancer Cancer Res 2001 61: 3212–3219
Acknowledgements
We acknowledge the excellent technical assistance of Simon McCallum. This work was supported, in part, by Grant RO1 CA82296 from the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hayes, G., Carpenito, C., Davis, P. et al. Alternative splicing as a novel of means of regulating the expression of therapeutic genes. Cancer Gene Ther 9, 133–141 (2002). https://doi.org/10.1038/sj.cgt.7700427
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cgt.7700427
Keywords
This article is cited by
-
Exploiting the tumor microenvironment in the development of targeted cancer gene therapy
Cancer Gene Therapy (2009)
-
Molecular mechanisms regulating the tumor-targeting potential of splice-activated gene expression
Cancer Gene Therapy (2004)
-
Establishment and application of minigene models for studying pre-mRNA alternative splicing
Science in China Series C: Life Sciences (2004)