Abstract
MicroRNAs (miRNAs) are important post-transcriptional regulators of their target genes in plants and animals. miRNAs are usually 20-24 nucleotides long. Despite their unusually small sizes, the evolutionary history of miRNA gene families seems to be similar to their protein-coding counterparts. In contrast to the small but abundant miRNA families in the animal genomes, plants have fewer but larger miRNA gene families. Members of plant miRNA gene families are often highly similar, suggesting recent expansion via tandem gene duplication and segmental duplication events. Although many miRNA genes are conserved across plant species, the same gene family varies significantly in size and genomic organization in different species, which may cause dosage effects and spatial and temporal differences in target gene regulations. In this review, we summarize the current progress in understanding the evolution of plant miRNA gene families.
Similar content being viewed by others
Introduction
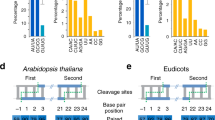
The commencing of the 21st century witnessed the explosion of a new class of regulatory genes that significantly expanded our knowledge about gene regulation: the discovery of small RNAs. Small RNAs, microRNAs (miRNAs), and small interfering RNAs (siRNAs) are 20-24 nucleotides long. They play important roles in post-transcriptional regulation of protein-coding genes via the RNA interference pathway 1. In animals, the appearance of miRNA genes coincided with the three major developmental innovations pivotal in animal evolution: the advent of the bilaterians, the vertebrates, and the placental mammals 2. In plants, miRNAs and their target genes have been conserved since the last common ancestor of bryophytes and seed plants more than 400 million years ago 3. Functionally, plant miRNAs are involved in many fundamental biological processes 4, such as leaf polarity 5, 6, floral identity 5, 6, stress responses 7, and auxin responses 8. Although the phenomena related to miRNA and siRNA functions were initially observed in plants 9, the first miRNA ever cloned was from an animal, Caenorhabditis elegans10, 11, 12. Similarly, the evolutionary history of miRNA genes was characterized earlier in animals, such as the formation of the miR-17 cluster 13, 14 and the exceptionally imprinted miR-134 gene cluster at human locus 14q32 15, 16.
Since the discovery of the first miRNA gene lin-4 in C. elegans 10, more than 500 different miRNAs have been identified in animals and plants. The number of miRNA genes is expected to increase to 500-1000 per species, which would amount to ∼2-3% of protein-coding genes 17. The Release 8.1 of the miRNA Registry (February, 2006) stores more than 3 963 entries including 114 C. elegans, 78 fruit fly, 372 zebra fish, 144 chicken, 462 human, 118 Arabidopsis, 178 rice, and 213 poplar miRNA genes 18 (http://www.sanger.ac.uk/Software/Rfam/mirna/). miRNAs are produced from larger precursors, which usually form a stem-loop structure, whereas siRNAs are generally derived from double-stranded RNAs. In plants, a ribonuclease III-like protein in the nucleus, DICER-LIKE 1 (DCL1), is responsible for processing primary miRNA gene transcripts, also known as pri-miRNAs 19. The subsequent products, the pre-miRNAs, are eventually processed into mature miRNA:miRNA* duplex 17. The mature miRNA duplex is then transported into the cytoplasm, where it is unwound and incorporated into the RNA-induced silencing complex 17. miRNAs and siRNAs represent a negative gene regulatory system. In animals, nearly all miRNAs suppress target gene functions by interfering with their translation. In plants, several possible mechanisms are applied by the miRNA genes. In some cases, the pairing of an miRNA with its target gene transcript causes the cleavage of the transcript 5, 20, 21. Plant miRNAs can also interfere with the ability of target transcripts in translation in a manner similar to animal miRNAs 6, 22. They can also initiate RNA-dependent RNA polymerase-mediated second-strand synthesis and trans-acting siRNA production 23, 24, 25, 26.
In both animal and plant genomes, multiple precursors are found to produce similar mature miRNA products 13, 27, 28, 29 i.e. miRNA genes also form gene families. The questions then arise: how did the miRNA genes evolve into large families? Was the evolutionary history of these genes with tiny final products similar to their protein-coding counterparts? Since the evolution of animal miRNA gene families has been a subject of many papers 14, 30, 31, in this review, we attempt to summarize the recent progress in the evolutionary study of miRNA gene families in plant genomes.
The miRNA gene families
Based on the number of their target sites, animal miRNAs are estimated to directly regulate more than a third of all protein-coding genes 4, 32, whereas in plants the numbers of known miRNAs and their targets are much smaller 33. Correspondingly, animal genomes have a large number of small miRNA gene families, whereas plant genomes have fewer, but larger miRNA gene families. As shown in Table 1, the average size of plant miRNA gene families is about 2.5 genes, while most miRNA gene families from animal species have an average size of less than two members. The zebra fish (Danio rerio), however, is an exception. It has a huge miRNA family, the miR-430 gene family, which is comprised of nearly 100 members 34, 35. MiR-430 genes are identical at nucleotides 2-8, which seems to be the most important segment for target recognition 32. They also have strong homology in their 3′ region, but differ in their central and terminal nucleotides. If this family were subtracted, the average size of zebra fish miRNA gene families would still stand around 2.72, similar to that in Arabidopsis.
Unlike animal miRNA genes where divergence has occurred even on the mature miRNA sequences, plant miRNAs derived from the same gene family are often highly similar. The similarities not only lie at the mature miRNA regions but also throughout the genes, indicating that the expansion of plant miRNA gene families has a recent origin and may be still ongoing. The model plant A. thaliana contains at least 22 miRNA gene families, most of which are conserved between monocots and eudicots 7, 18. The rice genome is more than three times larger than that of Arabidopsis and it has a third more miRNA genes too. Nevertheless, the MicroRNA Registry 8.1 records 47 and 46 miRNA gene families from the Arabidopsis and rice genomes, respectively (Table 1), suggesting a comparable number of unique miRNA gene pools in both monocots and eudicots.
Most miRNAs have shown to be conserved among related species and homologs were even found among those distantly related. For example, at least a third of C. elegans miRNAs are also present in the human genome 36, 37, suggesting their functional conservation among various animal lineages. Similar observation was also reported in plant genomes. Recent work by Zhang et al. 38 has shown, from more than 6 million plant EST sequences, a total of 481 miRNAs that belong to 37 miRNA families from 71 different plant species. Plant miRNAs have been demonstrated to be conserved across gymnosperms, fern, moss, and liverwort 3. Members of some miRNA gene families are physically clustered in plant genomes 27, 28, 29, 39. Such clusters in animal genomes often contain non-homologous miRNA genes 27, 28, 29, 39. Therefore, miRNA gene families in different species are characterized by different genome organizations.
The origin of plant miRNA genes
To date, only limited evidence is available about the origination of miRNA genes in plant genomes. The special stem-loop structure and the functional mode by which miRNAs pair with their target genes support the hypothesis that the de novo generation of miRNA genes was related to their target genes 40. In spite of this, cases of similarities between miRNAs and their target genes remain rare and have only been observed in the non-conserved miRNA genes, such as the Arabidopsis miR161 and miR163. In contrast to conserved miRNA genes that often have multiple copies, these miRNA genes are usually a single copy. MiR161 and miR163 target pentatricopeptide repeat proteins and S- adenosylmetheonine-dependent methyltransferases, respectively. Significant similarity to their target genes was found at the regions outside the mature miRNA and its pairing sequence (miRNA*) of these two genes. A putative mechanism was put forward for the arising and evolution of miRNA genes with unique target specificities 40. The hypothesis proposed that head-to-head or tail-to-tail gene configuration was generated by inverted gene duplication events from one founder gene with or without the founder gene's promoter. Sequence divergence at the inverted duplication locus occurred under constraints to maintain both the fold-back structure and the recognition by DCL1. Sequence degeneration continued until the point that only the miRNA or miRNA-complementary sequences were maintained for matching the founder gene sequence 40.
Such a model, however, may only apply to non-conserved plant miRNA genes since no such similarity has been found between conserved miRNA genes and their targets, nor would the model be able to explain animal miRNA origination because their precursors are too short to provide the information on their founder genes. Animal miRNA regulatory mechanisms are considered to be acquired by so-called “gain-of-interaction” events between miRNAs and their target genes. Most animal miRNAs regulate protein-coding genes by interfering with their translation via binding to the 3′-non-translated regions, during which more mismatches are allowed in the pairing process 31. This is in contrast to the observation that plant miRNAs are nearly identical to their target gene regions and such a pairing may cause cleavage of the target gene transcripts in many cases. The different functional modes in plant and animal genomes therefore imply a divergence in origination mechanisms for miRNA genes in the two kingdoms.
miRNA gene clusters – the formation of miRNA gene families
Clusters of miRNA genes have been found in both animal and plant genomes. Some clusters are so compact that multiple miRNA genes on the cluster can be transcribed as a single polycistron 29, 39. The history of miRNA gene clusters may represent general evolutionary experiences during the formation of most miRNA gene families. The animal miR-17 gene cluster, e.g., consists of miR-17, miR-18, miR-19a, miR-19b, miR-20, miR-25, miR-92, miR-93, miR-106a, and miR-106b. Some of these genes are not homologous, although evolutionarily related. MiR-17 genes confer important functions including negative regulation of expression of the E2F1 gene, which is involved in human cell cycle progression 41. Phylogenetic reconstruction indicated that the history of this cluster was governed by an initial phase of local (tandem) duplications, a series of duplications of entire clusters and subsequent loss of individual miRNA genes from the resulting paralogous clusters. The complicated history of the miR-17 gene family appears to be closely linked to the early evolution of the vertebrate lineage 13, 14, 27. The clusters are conserved across vertebrates: from teleost fish to human 13. The fact that they have been fixed in many modern animal genomes implies a selection advantage on such miRNA gene family organizations.
The observation in plant genomes is different. Although plant miRNA gene families are much larger, few of their members have been found to be clustered in a range of several kilobases 28, 29. miRNA genes of the same family are often scattered throughout the genome, indicating that plant genomes have experienced significant shuffling since the amplification of these families. One miRNA gene family, the miR395 gene family, is distinguished. Members of the miR395 gene family are clustered in several plant genomes with various cluster sizes and intergenic distances. Unlike animal miRNA gene clusters, plant miRNA gene clusters are comprised of homologous members 7, 29, 38. To date, there has been no report of clusters containing non-homologous, but evolutionarily related, miRNA gene members in plant genomes. Therefore, in contrast to the animal miRNA gene clusters, where the co-transcription of non-homologous miRNA genes could regulate multiple functionally related genes simultaneously, the consequence of the co-transcription of similar or identical miRNA genes on a plant gene cluster would be a dosage effect.
Additional expansion of miRNA gene families via segmental duplications
Many plants are considered to be ancient polyploids, including those with their whole-genome sequence available: Arabidopsis, rice, and poplar. If miRNA gene clusters existed before polyploidization, genome duplication events would produce duplicated miRNA clusters, a situation analogous to the protein-coding gene blocks 42. Segmental duplication may also occur during chromosomal recombination and shuffling. In the model plant Arabidopsis, a plethora of evidence demonstrate that protein-coding gene families arose by gene duplication and diversification 43, 44, 45. Similarly, by investigating contiguous miRNA distribution on the same or neighboring intergenic regions, Maher et al. 28 found 23 tandemly duplicated miRNA gene regions in the Arabidopsis genome. About two-thirds of these miRNAs were on the same strand with an average distance of ∼2 kb between tandem duplicates. Their study also showed that protein-coding genes flanking the miRNA genes were more conserved than those randomly chosen suggesting that the regions where the miRNA genes reside were indeed duplicated blocks. Two duplications, miR159a/miR159b and miR166a/miR166b, have at least four or more conserved flanking protein-coding genes, which fell in the large-scale duplications reported previously 43. Therefore, the seemingly random distributions of miRNA genes in current plant genomes reflect dynamic evolutionary histories, where the occurrence of various genome duplications was followed by chromosomal rearrangements and loss of duplicated genes. It has to be noted that miRNA genes have to co-evolve with their target genes. The target genes, many of which also form gene families, have experienced similar genetic changes. Thus, the unique regulation between miRNAs and their target genes was established in a “try-and-error” manner during dynamic genomic rearrangements. Such processes may have repeated multiple times and a coordinated regulatory network was generated that was advantageous over the old one, and hence was fixed by natural selection. The evolution of the miR395 gene families and their clusters in various plants provide a clear picture of such evolutionary scenarios in plant genomes.
The miR395 gene family, a case study
Among plant miRNA gene families, the miR395 gene family is interesting because the members of this family form clusters of various sizes in different species, from as compact as < 1 kb in rice to as large as 70 kb in Medicago truncatula! MiR395 is predicted to target mRNAs for ATP sulfurylases that catalyze the first step of inorganic sulfate assimilation 7. MiR395 transcripts are not detectable in Arabidopsis under normal growing conditions, but are inducible under low-sulfate stress. A database search showed that the miR395 matches sulfurylase mRNAs from several other plants, including rice, maize, Brassica juncea(brown mustard), and Allium cepa(garden onion), indicating that miR395/sulfurylase pairing is conserved in many plant genomes (data not shown).
In Arabidopsis, six miR395 genes are broken into two groups of tandem duplications (Figure 1A), as suggested by their high sequence similarity within the loop regions 7, 28. For each group of miRNA genes, two are on the same strand while the third is on the opposite strand, suggesting an intrachromosomal duplication event that probably occurred after the tandem gene duplications 28. Two genes, miR395b and miR395c, are highly conserved with only two base pair differences in their loop sequences. The rest also showed a high similarity with each other, suggesting that they are recent tandem duplicates.
Clusters of miR395 gene families from five different plant genomes. Thin black lines represent DNA fragments. Vertical bars denote the locations of miR395 genes on each cluster. The gene locations are roughly proportional to their real physical distances. Straight broken lines indicate two miR395 genes on different linkage groups with high sequence similarity. Curved broken lines with arrows at both ends indicate two miR395 genes on the same linkage group with high sequence similarity. Note that the unit on the top scale bar represents 1 kb, whereas for the bottom scale bar it represents 10 kb. (A) Two clusters of six miR395 genes on the Arabidopsis chromosome 1. (B) A cluster of four Solanum demissum miR395 genes. (C) Four compact clusters of miR395 genes in the rice genome. Clusters a, b, and d are segmental duplicates, whereas cluster c may have a different history of origination 29. (D) Two Medicago miR395 gene clusters. Note the large intergenic distances of Medicago miR395 genes and the high sequence similarity between the two clusters. The chromosome locations of these two fragments are unknown. (E) A cluster of poplar miR395 genes. Identical miR395 genes are indicated. (F) A multiple sequence alignment showing high sequence similarity between poplar miR395 genes.
This gene family is particularly interesting in rice for three reasons: first, it is significantly expanded when compared with Arabidopsis (24 vs 6); second, up to seven genes are compacted in a range of ∼1 kb; and third, they exhibit clear tandem and segmental duplication histories 29. The 24 rice miR395 genes are organized into four compact clusters that each could be transcribed as a single transcript (Figure 1B). The sequence similarity and the distribution of miRNA genes on these clusters indicate that three clusters, a, b, and d, were derived from segmental duplications, whereas the fourth cluster c may have originated separately 29. Clusters a and b are located about 815 kb apart on chromosome 4, whereas clusters c and d are on chromosomes 8 and 9, respectively. These clusters are distinguished by their sizes and the similarity of the miR395 gene members residing on them. The a, b, and d clusters bare miR395 precursors of ∼66 and ∼120 bp, whereas cluster c is comprised of miR395 genes of ∼79 and ∼92 bp. Some members on the clusters display high sequence similarity, while a gradual decrease in sequence similarity among other gene members suggests that duplication events occurred at various time points. Tandem duplication events involving one gene as well as two genes were observed on these duplicated clusters, demonstrating that duplication events occur at various scales. Compared with the Arabidopsis miR395 gene clusters that span ∼4 kb, rice miR395 gene clusters are more compact, each containing up to seven miR395 genes in a range of around 1 kb. Therefore, it is not surprising that these miRNA genes are transcribed in one polycistronic transcript, as supported by a cDNA sequence from the rice EST database 7.
In contrast, 16 miR395 genes in M. truncatula are distributed on two segments of larger than 45 kb with distances between neighboring genes up to 15 kb (Figure 1D) 29. A few corresponding genes on the two clusters are highly similar, suggesting that, in spite of their large sizes, these two DNA fragments were highly possible to be derived from segmental duplications. The sizes of miR395 genes, as measured from miRNA to miRNA*, are heterogeneous in the Medicago genome. Similar genomic organization of the miR395 gene family was also found in Populus trichocarpa(poplar) whose whole genome sequence became available recently. Genomic mapping of 10 recently cloned miR395 genes 46 showed that six miR395 genes were located on one segment of ∼22 kb (Figure 1E), whereas the rest were located elsewhere in the genome. The lengths of the poplar miR395 genes were either 85 bp (Figure 1E: genes g, h, i, and j) or 98 bp (Figure 1E: genes e and f). The distances between neighboring poplar miR395 genes were large, similar to some of those in Medicago. Despite the large intergenic distances, poplar miR395 genes are highly similar or even identical (Figure 1F), clearly suggesting a very recent origin of these miRNA genes.
In addition to the plants described above, a cluster of miR395 genes has also been observed on a BAC from the wild potato, Solanum demissum. In S. demissum, four miR395 genes were clustered in a region less than 4 kb, similar to that in Arabidopsis. Sequence similarity among the genes suggested that they were derived from a series of gene duplications, and at different time points in evolution (Figure 1B). A better picture of the miR395 gene family in S. demissum awaits the availability of more genomic sequences.
Genomic evolution and functional diversification of miRNA genes
The fact that miRNA sequences have to complement with their target gene transcripts for carrying out their functions suggests that miRNA genes have to co-evolve with their target genes 40. Once generated, miRNA genes would amplify through duplication events similar to those that drive the evolution of protein-coding genes: tandem gene duplications, segmental duplications, and chromosomal duplications or polyploidization 28, 29. Genes derived from tandem duplication events would diverge in sequences outside the mature miRNA and its pairing sequences. On the other hand, the extended intergenic sequences provide locations for novel promoters. This would allow precursors that produce the same miRNA to take on novel spatial and temporal features. Like their protein-coding counterparts, the acquisition of new promoters and the accumulation of mutations in the genes provide miRNA genes with opportunities for subfunctionalization or even neofunctionalization 44, 47. Therefore, it is reasonable to suggest that the variation of genomic organizations of miR395 gene families, and maybe other miRNA gene families too, will generate different regulatory patterns in each plant. In addition, the increase or the decrease in miRNA gene copy numbers will cause a dosage effect. Therefore, future studies should assess such effects on the target gene regulations caused by the genomic divergence of miRNA gene families. However, understanding the evolution of plant miRNA gene families will help us to better understand the complexities of the ancient gene regulatory mechanisms underlined by these small RNA molecules.
References
Bartel B, Bartel DP . MicroRNAs: at the root of plant development? Plant Physiol 2003; 132:709–717.
Hertel J, Lindemeyer M, Missal K, et al. The expansion of the metazoan microRNA repertoire. BMC Genomics 2006; 7:25.
Floyd SK, Bowman JL . Gene regulation: ancient microRNA target sequences in plants. Nature 2004; 428: 485–486.
Mallory AC, Vaucheret H . Functions of microRNAs and related small RNAs in plants. Nat Genet 2006; 38:S31–S36.
Palatnik JF, Allen E, Wu X, et al. Control of leaf morphogenesis by microRNAs. Nature 2003; 425:257–263.
Chen X . A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 2004; 303:2022–2025.
Jones-Rhoades MW, Bartel DP . Computational identification of plant MicroRNAs and their targets, including a stress-induced miRNA. Mol Cell 2004; 14:787–799.
Wang JW, Wang LJ, Mao YB, et al. Control of root cap formation by MicroRNA-targeted auxin response factors in Arabidopsis. Plant Cell 2005; 17:2204–2216.
Hamilton AJ, Baulcombe DC . A species of small antisense RNA in posttranscriptional gene silencing in plants. Science 1999; 286:950–952.
Lee RC, Feinbaum RL, Ambros V . The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993; 75:843–854.
Pasquinelli AE, McCoy A, Jimenez E, et al. Expression of the 22 nucleotide let-7 heterochronic RNA throughout the Metazoa: a role in life history evolution? Evol Dev 2003; 5:372–378.
Pasquinelli AE, Reinhart BJ, Slack F, et al. Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature 2000; 408:86–89.
Tanzer A, Stadler PF . Molecular evolution of a microRNA cluster. J Mol Biol 2004; 339:327–335.
Tanzer A, Stadler PF . Evolution of MicroRNAs. In microRNA protocols, methods in molecular biology. Edited by: Yig SY. Totowa. NJ: Humana Press Inc., 2006, in press.
Lagos-Quintana M, Rauhut R, Yalcin A, et al. Identification of tissue-specific microRNAs from mouse. Curr Biol 2002; 12:735–739.
Kim J, Krichevsky A, Grad Y, et al. Identification of many microRNAs that copurify with polyribosomes in mammalian neurons. Proc Natl Acad Sci USA 2004; 101:360–365.
Bartel DP . MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004; 116:281–297.
Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ . miRBase: microRNA sequences, targets and gene nomenclature. Nucl Acids Res 2006; 34:D140–D144
Papp I, Mette MF, Aufsatz W, et al. Evidence for nuclear processing of plant MicroRNA and short interfering RNA precursors. Plant Physiol 2003; 132:1382–1390.
Kasschau KD, Xie Z, Allen E, et al. P1/HC-Pro, a viral suppressor of RNA silencing, interferes with Arabidopsis development and miRNA function. Dev Cell 2003; 4:205–217.
Llave C, Xie Z, Kasschau KD, Carrington JC . Cleavage of Scarecrow-like mRNA targets directed by a class of Arabidopsis miRNA. Science 2002; 297:2053–2056.
Aukerman MJ, Sakai H . Correction: Regulation of flowering time and floral organ identity by a microRNA and its APETALA2-like target genes. Plant Cell 2004; 16:555.
Volpe TA, Kidner C, Hall IM, et al. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 2002; 297:1833–1837.
Peragine A, Yoshikawa M, Wu G, Albrecht HL, Poethig RS . SGS3 and SGS2/SDE1/RDR6 are required for juvenile development and the production of trans-acting siRNAs in Arabidopsis. Genes Dev 2004; 18:2368–2379.
Vazquez F, Vaucheret H, Rajagopalan R, et al. Endogenous trans-acting siRNAs regulate the accumulation of Arabidopsis mRNAs. Mol Cell 2004; 16:69–79.
Allen E, Xie Z, Gustafson AM, Carrington JC . MicroRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell 2005; 121:207–221.
Tanzer A, Amemiya CT, Kim CB, Stadler PF . Evolution of MicroRNAs located within Hox gene clusters. J Exp Zool: Mol Dev Evol 2005; 304B:75–85.
Maher C, Stein L, Ware D . Evolution of Arabidopsis microRNA families through duplication events. Genome Res 2006; 16:510–519.
Guddeti S, Zhang dC, Li AL, et al. Molecular evolution of the rice miR395 gene family. Cell Res 2005; 15:631–638.
Bompfünewerer AF, Flamm C, Fried C, et al. Evolutionary patterns of non-coding RNAs. Theor Biosci 2005; 123:301–369.
Bartel DP, Chen CZ . Micromanagers of gene expression: the potentially widespread influence of metazoan microRNAs. Nat Rev Genet 2004; 5:396–400.
Lewis BP, Burge CB, Bartel DP . Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005; 120:15–20.
Jones-Rhoades MW, Bartel DP, Bartel DP . MicroRNAs and their regulatory roles in plants. Annu Rev Plant Biol 2006; 57:19–53.
Chen PY, Manninga H, Slanchev K, et al. The developmental miRNA profiles of zebrafish as determined by small RNA cloning. Genes Dev 2005; 19:1288–1293.
Giraldez AJ, Cinalli RM, Glasner ME, et al. MicroRNAs regulate brain morphogenesis in zebrafish. Science 2005; 308:833–838.
Lim LP, Glasner ME, Yekta S, Burge CB, Bartel DP . Vertebrate microRNA genes. Science 2003; 299:1540.
Lim LP, Lau NC, Weinstein EG, et al. The microRNAs of Caenorhabditis elegans. Genes Dev 2003; 17:991–1008.
Zhang B, Pan X, Cannon CH, Cobb GP, Anderson TA . Conservation and divergence of plant microRNA genes. Plant J 2006; 46:243–259.
Lee Y, Jeon K, Lee JT, Kim S, Kim VN . MicroRNA maturation: stepwise processing and subcellular localization. EMBO J 2002; 21:4663–4670.
Allen E, Xie Z, Gustafson AM, et al. Evolution of microRNA genes by inverted duplication of target gene sequences in Arabidopsis thaliana. Nat Genet 2004; 36:1245–1246.
O'Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT . c-Myc-regulated microRNAs modulate E2F1 expression. Nature 2005; 435:839–843.
Town CD, Cheung F, Maiti R, et al. Comparative genomics of Brassica oleracea and Arabidopsis thaliana reveal gene loss, fragmentation, and dispersal after polyploidy. Plant Cell 2006: tpc.106.041665.
AGI. Analysis of the genome sequence of the flowering plant Arabisopsis thaliana. Nature 2000; 408:796–815.
Prince VE, Pickett FB . Splitting pairs: the diverging fates of duplicated genes. Nat Rev Genet 2002; 3:827–837.
Cannon S, Mitra A, Baumgarten A, Young N, May G . The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol 2004; 4:10.
Lu S, Sun YH, Shi R, et al. Novel and mechanical stress-responsive microRNAs in Populus trichocarpa that are absent from Arabidopsis. Plant Cell 2005; 17:2186–2203.
Force A, Lynch M, Pickett FB, et al. Preservation of duplicate genes by complementary, degenerative mutations. Genetics 1999; 151:1531–1545.
Acknowledgements
We thank Charles H Leseberg for critical reading of this manuscript. The authors thank the support from The HI-TECH Research and Development Program of China (863 program, #2003AA207160), the National Basic Research Program (973 program, #2004CB117200), and the Generation Challenge Program, a CGIAR challenge program cultivating plant diversity for resource poor.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, A., Mao, L. Evolution of plant microRNA gene families. Cell Res 17, 212–218 (2007). https://doi.org/10.1038/sj.cr.7310113
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7310113
Keywords
This article is cited by
-
Delineating the tissue-mediated drought stress governed tuning of conserved miR408 and its targets in rice
Functional & Integrative Genomics (2023)
-
Genome-wide identification and characterization of PdbHLH transcription factors related to anthocyanin biosynthesis in colored-leaf poplar (Populus deltoids)
BMC Genomics (2022)
-
Genome-wide analysis of the apple CaCA superfamily reveals that MdCAX proteins are involved in the abiotic stress response as calcium transporters
BMC Plant Biology (2021)
-
Role of miRNAs in the host–pathogen interaction between sugarcane and Colletotrichum falcatum, the red rot pathogen
Plant Cell Reports (2021)
-
Investigation on the Conserved MicroRNA Genes in Higher Plants
Plant Molecular Biology Reporter (2021)