ABSTRACT
The retinoblastoma (RB) tumor suppressor protein, pRB, plays an important role in the regulation of mammalian cell cycle. Furthermore, several lines of evidence suggest that pRB also involves in the regulation of apoptosis. In the present study, the degradation of pRB was observed in apoptotic gastric tumor cells treated with a new potent anti-tumor component, tripchlorolide (TC). The inhibition of pRB degradation by a general cysteine protease inhibitor IDAM resulted in the reduction of the apoptotic cells. Furthermore, the survival of the gastric tumor cells under the TC treatment was enhanced by an over-expression of exogenous pRB. These results suggest that the pRB degradation of the gastric tumor cells under the TC treatment involves in the apoptotic progression. In addition, the same extent of TC-induced pRB-degradation was detected in the gastric tumor cells containing a p53 dominant-negative construct, indicating that this kind of pRB degradation is p53-independent.
Similar content being viewed by others
INTRODUCTION
The retinoblastoma (RB) tumor suppressor gene was first cloned in 1986 1. As a suppressor of G1/S progression, the RB gene product, pRB, plays a pivotal role in regulating cell cycle of the normal mammalian cells 2. Moreover, pRB protein acts as a critical effector with DNA damage checkpoint function in response of cell stresses and DNA damage. pRB is hypophosphorylated following genotoxic stress, which finally elicits G1-phase cell cycle arrest due to the suppression of the E2F family of transcription factors 3, 4, 5, 6.
In addition to the regulation of the cell growth, pRB may also involve in the regulation of apoptosis. It was observed the proteolytical degradation of pRB in the apoptotic cells induced by a number of stimuli such as TGF-b, UV irradiation, ionizing irradiation and p53 over-expression, whereas the exogenous overexpression of pRB could reduce the apoptotic progression 7, 8, 9, 10. Moreover, mouse embryos lacking functional pRB showed the higher levels of apoptosis in many kinds of tissues 11. Recently, Chau et al showed that the degradation of pRB was associated with the increased level of apoptosis directly in the RB-MI knockin mice 12. Taken together, these data suggest that the degradation of pRB promotes the progression of apoptosis.
There are two elucidated mechanisms for the degradation of pRB: one is proteolysis via proteasomal pathway, and the other is caspase-dependent cleavage. It was identified several proteins having the capability to bind to pRB and stimulating its degradation via proteasome-dependent proteolysis, such as human papillomavirus (HPV) oncoprotein E7 13, 14, 15, 16, human cytomegalovirus pp71 protein 17, hepatocellular carcinoma-associated protein gankyrin 18, 19, human T-cell leukemia virus type-I (HTLV-I) oncoprotein Tax 20 and MdM2 21. As to the caspase-mediated pathway, several caspase-cleavage sites of pRB have been mapped. For example, Fattman CL revealed that the pRB underwent cleavage by caspase after Asp residues those located between amino acids 341 and 421. Any cleavage of these sites would generate a C-terminal fragment p48 and a N-terminal containing fragment p68 22. It has been elucidated that pRB is cleaved by caspase-3 at the DXXD recognition motif at extreme C-terminal, which can be blocked by z-DEVD-FMK 23, 24. This is consistent with the result from the RB-MI knockin mice generated by introducing a caspase-resistant RB 12.
Tripchlorolide (TC) is a diterpenoid component isolated from extracts of Chinese herb Tripterygium wilfordii Hook f 25. Several reports showed that TC inhibited the proliferation of peripheral blood mononuclear cells 26, mesangial cells 27 and spermatocytes 28. It was also reported that TC could induce apoptosis of cells in both p53-dependent and p53-independent manner 29, 30. Moreover, our group showed that TC caused DNA damage in CHO cells and induced the degradation of c-Myc protein 31. The present study demonstrated that the degradation of pRB involved in the progression of apoptosis of human gastric tumor cells under the TC treatment. Further analysis showed that the degradation of pRB was p53-independent and may be mediated by cysteine protease rather than the proteasome or the caspases.
MATERIALS AND METHODS
Cell culture and materials
AGS, a human gastric tumor cell line (ATCC), was maintained in Dulbecco's Modified Eagle's Medium (DMEM, Gibco BRL) supplemented with 10% fetal bovine serum (FBS, Gibco BRL) at 37°C in 5% CO2. Ac-DEVD-CHO (cell permeable), Ac-IETD-CHO (cell permeable), and Ac-LEHD-CMK were purchased from Calbiochem (Calbiochem); MG132 and iodoacetamide (IDAM) were purchased from Sigma (Sigma); z-VAD-FMK was purchased from Promega (Promega).
DNA fragmentation assay
AGS cells were harvested at indicated time and resuspended in cell lysis buffer (10 mM Tris-HCl, pH 8.0, 25 mM EDTA, 0.25% Triton X-100). After 30 min incubation on ice, the lysates were centrifuged at 13,500×g at 4°C for 15 min. The supernatant were collected and incubated with 100 μg/ml RNase A at 37°C for 30 min, and then with 200 μg/ml protease K at 56°C overnight. The DNA fragments were extracted with phenol and subjected to 1.5% agarose gel.
Flow cytometric analyses
For the DNA content analysis, the cells were collected and fixed with 70% ethanol, treated with RNase A and stained with propidium iodide (PI, Sigma). The DNA contents were measured with a flow cytometer (FACScan). For M30-apoptotic analysis, the cells were subjected to the CytoDEATH M30 assay according to the manufacture's instruction (Roche Molecular Biochemicals). Briefly, the cells were fixed in ice-cold methanol at −20°C for 30 min, washed with washing buffer (PBS containing 0.1% Tween 20), and incubated with 100 μl M30 CytoDEATH fluorescein antibody working solution (PBS, 1% BSA, 0.1% Tween 20 and 0.4% CytoDEATH fluorescein antibody) at 25°C for 1 h. After the washing, the cells were subjected to flow cytometric analysis.
Isolation of cytosolic and mitochondrial fractions
The cells were treated with a digitonin-buffer (20 mM Hepes-KOH, pH 7.3, 110 mM KAc, 5 mM NaAc, 2 mM MgAc2, 1 mM EGTA and 200 μg/ml digitonin) on ice for 10 min to permeabilize the cell membrane. The cell lysate was then centrifugated at 10,000×g at 4 °C for 15 min. The supernatant was collected as a cytosolic fraction, and the pellet was as a mitochondria-containing fraction.
Western blot analysis
AGS cells were lysed with 1×SDS-loading buffer (50 mM Tris-Cl, pH 6.8, 100 mM DTT, 2% SDS, 10% glycerol and 0.1% bromophenol blue) as the whole-cell sample. In the case of digitonin-permeabilized cells, the cytosolic fraction was mixed with equal volume of 2×SDS-loading buffer, and the mitochondria-fraction was suspended with 1×SDS-loading buffer. The protein samples were subjected to SDS-PAGE gel electrophoresis. Immunoblottings were carried out with primary antibodies (anti-Bax, anti-Cytochrome c, anti-β-actin, Santa Cruz; anti-Bcl-2, Sigma; anti-COX 4, Molecular Probes; anti-p53, Cell Signaling; anti-RB, BD transduction Lab), respectively. The proteins were detected by enhanced chemiluminescence (ECL-plus, Amersham Pharmacia Biotech).
Comet assay
The Comet assay was carried out as described 31. Briefly, the cells were suspended in 0.5% low melting point agarose at a concentration of 1×104 cells/ ml and applied to the surface of a microscope slide. The slides were immersed in a lysis buffer (2.5 M NaCl, 100 mM Na2EDTA, 10 mM Tris, pH 10.0, 1% N-lauroyl-sarcosine, and 1% Triton X-100) for 1 h, and then in 0.3 M NaOH, 1 mM Na2EDTA for 20 min, both at 4°C. After slides were put to horizontal gel electrophoresis, they were washed and stained with 1 mg/ml ethidium bromide. The cells were analyzed with fluorescence microscope. The cells were analyzed with fluorescence microscope and the proportion of the cells containing damaged DNA strand breaks as the comet-cells was scored. DNA-damage-induced Comet cells were also analyzed with free download Visual Cometassay Software as described 32. Each cell was assigned a value of 0 (without tail) to 4 (almost all DNA as tail), and the total score of 100 cells represented the DNA damage level, which would be located in the area between 0 and 400 “arbitrary units”.
Plasmid construction and generation of stable cell lines
The plasmid p44-2L RB containing full length of human RB was a kind gift from Dr. Xue Liang Zhu. After the conserved caspase cleavage site DEADGSKHL was mutated to DEAAESKHL 12 with QuikChange Site-Directed Muatagenesis Kit (Stratagene), the caspase3-resistant RB insert was cloned into SalI/BamHI sites of pEGFP-C1 (Clontech) to generate the plasmid pEGFP-HIRB. AGS cells were transfected with pEGFP-HIRB by LipofectamineTM 2000 system (Gibco BRL) and the stable cell line HI-Rb was selected in the presence of 500 μg/ml G418 (Roche Molecular Biochemicals).
For the stable cell line AER, AGS cells was transfected with pEGFP-cgTP53, an EGFP fused p53 dominant-negative construct 30, by LipofectamineTM 2000 system. AER was further selected in the presence of 500 μg/ml G418.
RESULTS
TC induced DNA damage and apoptosis in AGS cells
To address the effect of TC on tumor cells, we treated the exponential growing gastric tumor cells, AGS, with 20 ng/ml TC for 24 h. An apoptotic population was identified from the TC-treated AGS cells as measured by DNA fragmentation or M30 assay (Fig. 1A and Fig. 1B). The results from M30 assay further revealed that the TC induction of apoptosis was in both time- (Fig. 2A) and dose- ( data not shown) dependent manners.
Induction of apoptosis of AGS cells by TC treatment. AGS cells were treated with 20 ng/ml TC for 24 h. (A) Apoptotic cells measured by DNA fragmentation assay. The DNA fragments were extracted and separated by 1.5% Agarose gel. Lane 1, marker; lane 2, no TC treatment; lane 3, TC treatment for 24 h. (B) Apoptotic cells measured by M30 assay. The cells were labeled with M30 CytoDEATH antibody and measured by flow cytometry. (C) Western-blotting analysis of Cytochrome c, Bax and Bcl-2 of AGS cells. The cells were collected and separated as the mitochondrial and cytosolic fractions as described in Material and Methods. Total cell lysates (lane 1, 4), the cytosolic fractions (lane 2, 5) and the mitochondrial fractions (lane 3, 6) were analyzed by Western blot. The mitochondrial protein COX4 served as a quality control of the fractionation.
Degradation of pRB concomitant with apoptosis of TC-treated AGS cells. (A) Time course of apoptotic AGS cells by TC treatment. TC-treated AGS cells were analyzed by M30 assay. Data shown are representative of three independent experiments (mean ± SD). (B) Detection of pRB by Western blot. AGS cells were treated with 20 ng/ml TC for indicated times, respectively. The cell lysates were separated by 8% SDS-PAGE and subjected to Western-blotting analysis. Actin was used as sample-loading control.
In order to understand the molecular mechanism in the TC induced apoptosis of AGS cells, we monitored the changes of mitochondrial pathway in TC treated AGS cells. All TC-treated cells were collected and fractionationed as the cytosolic fraction and the mitochondria-containing fraction by digitonin-permeabilization assay (see Materials and methods). Western-blotting analysis showed that cytochrome c was released from mitochondria into the cytosol in the TC treated cells, while it remained in the mitochondria of the cells without TC treatment (Fig. 1C, compare lanes 2 to 5). Furthermore, Bax was significantly translocated from the cytosol to mitochondria in the TC treated cells, whereas most Bax proteins of the untreated cells remained in the cytosol (Fig. 1C, compare lanes 5, 6 to 2 and 3). COX4 (cytochrome c oxidase IV), a mitochondrial membrane locating protein, served as an internal control in order to show no cross contamination between mitochondrial and cytosolic fractions (Fig. 1C). Taken together, these results indicated that TC induced the apoptosis of AGS cells through the mitochondrial-regulated apoptotic pathway.
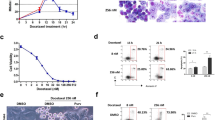
Comet assay was performed to detect whether DNA damage was caused in TC- treated AGS cells. The cells treated with TC for 8 h and 16 h, respectively, were collected and subjected to Comet assay, of which no significant apoptotic cells were detected by M30 assay (Fig. 2A). Significant Comet cells were observed in the treated cells (Fig. 3A). The percentage of DNA strand breaks was increased significantly from 20.83% at 8 h to 53.33% at 16 h (Fig. 3B). In addition, the quantitatively measurement showed that the Comet-tail length was doubled in the cells treated with TC for 16 h than those treated with TC for 8 h (see Materials and methods). These data consisted with the previous report that TC caused DNA damage in CHO cells 31.
Induction of DNA damage of AGS cells by TC treatment. (A) Identification of comet cells. AGS cells were treated with 20 ng/ml TC for 16 h and subjected to comet assay. The cells with DNA strand breaks (comet cells) were pointed out by arrowheads. (B) Calculation of comet cells. AGS cells were treated with 20 ng/ml TC for 8 h and 16 h separately and subjected to comet assay. The result was scored for more than 200 cells per sample. Data shown are representative of three independent experiments (mean ± SD).
pRB was degradated during TC-induced apoptosis
Since pRB is an effector in response to DNA damage, we analyzed the status of pRB of AGS cells in the presence of TC. After the cells were treated with TC for indicated times, respectively, the cell lysates were subjected to SDS-PAGE gel 33. The Western-blotting analysis showed that the total amount of pRB decreased significantly in the population treated with TC for 24 h, and hardly detected in the population treated with TC for 36 h (Fig. 2B). Moreover, the dynamics of the degradation of pRB was evaluated by comparing Fig. 2A with Fig. 2B. The results indicated that pRB diminished along with the apoptotic progress of AGS cells induced by TC, both in a time dependent manner. In other words, the more degradation of pRB was identified, the more apoptotic cells were detected.
Cysteine protease mediated the TC-induced degradation of pRB
It has been well reported that two molecular mechanisms involving in the degradation of pRB. The first is that pRB might be degraded through proteasomal pathway, in which pRB was targeted to 26S proteasome and underwent proteolytic degradation 15, 34, 35. Therefore, we examined whether the presence of proteasome inhibitor MG132 was capable to prevent the degradation of pRB induced by TC-treatment. Western-blotting analysis showed that the pRB degradation of TC-treated AGS cells in the presence of 0.5 μM MG132 was to the same extent as the samples without MG132 treatment (Fig. 4A), whereas the ubiquitin-proteasome dependent degradation of c-Myc, which was used as a positive control, was completely prevented under MG132 treatment (data not shown). These data indicate that the degradation of pRB is independent of proteasomal pathway in the TC-treated AGS cells.
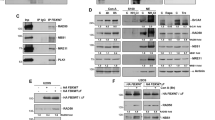
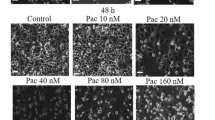
The pRB degradation involves in cysteine protease rather than proteosome. (A) Effects of protease inhibitors on TC-induced pRB degradation. The cells were treated with 20 ng/ml TC for indicated times, respectively. Protease inhibitors were co-administrated as indicated. The cells only treated with TC were used as control, which was indicated as “none”. The final concentration of the inhibitors was: MG132, 0.5 μM; Ac-DEVD-CHO, 50 μM; Ac-IETD-CHO, 40 μM; Ac-LEHD-CMK, 20 μM; z-VAD-FMK, 75 μM; IDAM, 50 μM. The cell lysates were separated by 8% SDS-PAGE and subjected to Western-blotting analysis. Actin was used as sample-loading control. (B) Effects of protease inhibitors on TC-induced apoptosis. AGS cells were co-treated with 20 ng/ml TC and the indicated inhibitor for 24 h. The condensed apoptotic nuclei stained with DAPI were determined under the fluorescent microscope. Data shown are representative of three independent experiments (mean ± SD).
Another reported possible pathway for the degradation of pRB is caspase-3/7-dependent 22, 23, 24. To detect this possibility, the TC-treated AGS cells were administrated with a caspase3/7 specific inhibitor, Ac-DEVD-CHO, at the concentration of 50 μM 24. The result showed little effect on the prevention of the pRB degradation of TC-treated AGS cells (Fig. 4A), whereas 50 μM Ac-DEVD-CHO sufficiently inhibited the cleavage of PARP (data not shown). To further confirm this observation, an AGS-derivative cell line HI-Rb containing the mutated conservative caspase-3 cleavage site of pRB was constructed (see Materials and methods). Comet assay showed that HI-Rb cells had the similar amounts of DNA strand breaks comparing to AGS cells when cells were treated with TC for 16 h (Fig. 5D), while the Western-blot data presented that the amount of EGFP-fused caspase3-resistant pRB in HI-Rb cells treated with TC was also reduced (Fig. 5A). Taken together, the results suggest that the caspase3-like activity does not play the key role in the process of the pRB degradation in the TC-treated AGS cells.
Inhibition of the TC-induced apoptosis by the over-expression of pRB. (A) Detection of caspase-3-resistant pRB (Casp-3-r pRB) by Western blot. HI-Rb cells were treated with 20 ng/ml TC for indicated times, respectively. EGFP antibody was used for detecting the protein product of the pEGFP-caspase3-r pRB. (B) Effects of the pRB over-expression on TC-induced apoptosis. AGS and HI-Rb cells were treated with 5 ng/ml TC for 48 h, respectively. The condensed apoptotic nuclei stained with DAPI as representative of the apoptotic cell were determined under the fluorescent microscope. Data shown are representative of three separate experiments (mean ± SD). (C) Comparison of total pRB of AGS cells and HI-Rb cells. AGS and HI-Rb cells were treated with 5 ng/ml TC for 48 h, respectively. The anti-RB antibody was used for detecting the total protein product of RB. Both native pRB and extrinsic Casp-3-r pRB in HI-Rb cells were detected. (D) Visual scoring of comet cells. No.1, AGS cells treated with TC (20 ng/ml); No.2, AGS cells co-treated with TC (20 ng/ml) and IDAM (50 μM); No. 3, AGS cells co-treated with TC (20 ng/ml) and z-VAD-FMK (75 μM); No. 4, HI-Rb cells treated with TC (20 ng/ml). The cells were treated with indicated drugs for 16 h and subjected to comet assay. The Comet cells were measured into 5 classes, from 0 (without tail) to 4 (almost all DNA as tail) with Visual Cometassay Software 32. 100 cells of each sample were measured. Data shown are representative of three independent experiments (mean ± SD).
In order to explore the degradation mechanism of pRB, other proteinase inhibitors were further used in this study. Though a little prevention effect on the pRB degradation was detected in the presence of caspase 9 inhibitor Ac-LEHD-CMK, the caspase 8 inhibitor Ac-IETD-CHO did not prevent the degradtion of pRB in the TC- treated cells (Fig. 4A). We further co-administrated the pan-caspase inhibitor z-VAD-FMK and the general cysteine protease inhibitor IDAM 36, 37, 38 to the TC-treated AGS cells, respectively. As shown in Fig. 4A, the degradation of pRB in the TC-treated AGS cells was fully inhibited by IDAM, whereas the amount of pRB was restored partially by z-VAD-FMK. Since the administration of IDAM or z-VAD-FMK did not significantly inhibit TC induced DNA damage detected by comet assay (Fig. 5D), these results indicate that the unknown caspase(s) may play the major role in the process of the pRB degradation during the apoptosis of TC-treated AGS cells.
Prevention of the pRB degradation or overexpression of pRB reduced the TC- induced apoptosis of AGS cells
To address the question whether the degradation of pRB promotes the apoptotic progression, the amount of apoptotic TC-treated AGS cells in the presence of different protease inhibitors, respectively, were measured by counting the condensed nuclei as the representative of apoptotic cells under the fluorescent microscope. The results showed that the apoptotic cells were significantly reduced to the background level in the presence of the inhibitor IDAM (Fig. 4B), which prevented the degradation of pRB (Fig. 4A), whereas the amount of apoptotic cells in the presence of MG132, Ac-DEVD-CHO, Ac-IETD-CHO or Ac-LEHD-CMK, which did not inhibit the degradation of pRB (Fig. 4A), were to the same extent of only TC-treated apoptotic cells (Fig. 4B). The quantitative analysis showed that the level of pRB protein in the 36h-TC-treated cells was 57.89% in the presence of VAD, but 12.12% in the presence of DEVD, and 26.91% in the presence of LEHD, respectively. Since the apoptotic cells were also significantly reduced by z-VAD-FMK but not reduced by Ac-DEVD-CHO or Ac-IETD-CHO (Fig. 4B), it might exist a threshold that small amount of pRB could not protect cells against apoptosis. Taken together, these results suggest that the degradation of pRB is required in the apoptotic progression of TC-treated AGS cells.
To eliminate the possibility that the inhibition of the TC-induced apoptosis by IDAM might also be due to the general inhibitory ability of IDAM, we treated both AGS and HI-Rb cells, in which exogenous RB was over-expressed (see Materials and methods), with TC for 48 h. The condensed nuclei as the representative of apoptotic cells were counted under the fluorescent microscope, while the protein level of pRB was detected by Western blot using anti-RB antibody. The results showed that the apoptotic population of the RB-over-expressed cells was lower than that of the wild AGS cells in the presence of TC (Fig. 5B), when the certain amount of exogenous RB protein remained in the TC-treated cells (Fig. 5A and Fig. 5C). On the other hand, the existence of exogenous RB protein did not influent the level of DNA damage in the TC-treated cells (Fig. 5D). Taken together, these results support the conclusion that the degradation of pRB involves in the process of the TC-induced apoptosis of AGS cells.
TC-induced pRB degradation of AGS cells was p53 independent
pRB regulation is tightly related to p53, for example, p53 suppresses transcription of RB gene 39. Furthermore, it was reported that p53 facilitated pRB cleavage in interleukin-3-deprived lymphoid cells, which resulted in the apoptotic progression 40. In AGS cells, p53 was functional and accumulated rapidly under the treatment of TC (Fig. 6A and 6D). To address whether the activated p53 involved in the degradation of pRB in the TC treated AGS cells, an AGS-derivative cell line, AER, containing the p53 dominant-negative was constructed (see Materials and methods) 30. When the cells were administrated with 20 ng/ml TC for the indicated times, there was the pRB reduction of the TC-treated AER cells in a time-dependent manner (Fig. 6B). In addition, the TC-induced pRB degradation was also detected in another gastric cancer cell line, MKN28 (data not shown), in which p53 gene was reported to be invalid 41. Taken together, these results suggested that the TC-induced pRB degradation of gastric cancer cells is p53-independent.
p53-independent degradation of pRB in TC-treated AGS cells. (A) Detection of p53 in TC-treated AGS cells by Western blot analysis. AGS cells were treated with 20 ng/ml TC for indicated times, respectively, and subjected to Western blot analysis. Actin was used as sample-loading control. (B) Detection of pRB in TC-treated AER cells by Western blot analysis. AER cells, which stably express a p53 dominant negative product, were treated with 20 ng/ml TC for indicated times, respectively, and subjected to Western blot analysis. Actin was used as sample-loading control. (C) Effects of the p53 dominant-negative on TC-induced apoptosis. AGS and AER cells were treated with 20 ng/ml TC for 24 h, respectively. The condensed apoptotic nuclei stained with DAPI as representative of the apoptotic cell were determined. Data shown are representative of three independent experiments (mean ± SD). (D) Comparison of total p53 of AGS cells and AER cells. AGS and AER cells were treated with 20 ng/ml TC for 24 h, respectively. The anti-p53 antibody was used for detecting the total protein product of p53 in both cell lines.
Since our previous data reported that TC could induce apoptosis of HeLa S3 cells in a p53 dependent manner 30, we analyzed the apoptosis of AER cells containing the p53 dominant-negative under the TC-treatment. The results showed the significant reduction of the apoptotic population observed in the TC treated AER cells (Fig. 6C), indicating that p53 still plays other important role in the progression of TC-induced apoptosis of AGS cells although the degradation of pRB is p53-independent.
DISSCUSSION
It is well known that the dephosphorylation of pRB in G1 phase of the cell cycle is a key event for the transition of G1/S phase. Our study showed that pRB of AGS cells was dephosphorylated after 8 hours' TC treatment, which was prior to the detectable degradation of pRB (data not shown). This observation was similar to the finding of Dou's lab 42. They documented that pRB underwent degradation after dephosphorylated under the tamoxifen treated human breast carcinoma cells. These results suggest that the population of the dephosphorylated pRB is the main target for the degradation, which may play a role for preventing the progression of G1 phase into S phase under the apoptotic stimuli.
Ubiquitin-proteasomal and caspase3-like pathways are two elucidated mechanisms involved with the disruption of pRB so far [15, 16, 17, 20, 22, 23]. In the present study, by using protease inhibitors, we revealed that the degradation of pRB is proteasome independent, and there's other unknown cysteine protease but not caspase3-like and caspase 8/9 protease cleaved the protein RB. Since the pan caspase inhibitor z-VAD-FMK can partially inhibit the TC induced degradation of pRB, while the general cysteine protease inhibitor IDAM can totally inhibit the TC induced degradation of pRB, we conclude that two different mechanisms involve in the pRB degradation progression: one involves in the activity of caspase(s), and the other involve the activity of unknown cysteine protease.
The signal pathways controlled by the two prominent tumor suppressors RB and p53 are interconnected in several levels 39, 43, 44, 45, 46. Although it was reported that p53 suppressed transcription of RB gene 39, our study demonstrated the activated p53 did not suppress RB expression in AGS cells under the TC treatment (data not shown). Oren's group found that the p53-facilitated pRB cleavage was due to the activity of p53-dependent caspases 40. Since our work showed that TC-induced pRB was cleaved by an unknown cysteine protease rather than caspase3-like protease and this pRB degradation was p53-independent, it should be concluded that the activity of this kind of cysteine protease is independent the presence of functional p53 within the cells.
DNA-damaging agents are used frequently in cancer therapy. Our previous works showed that TC could induce apoptosis by degrading c-Myc protein 31 or via p53-dependent apoptotic pathway 30 in different cell lines. In the present study, we revealed the p53-independent degradation of pRB might be one of the causations of TC-induced apoptosis of AGS cells, which suggests a novel mechanism of action that TC exhibits its antineoplastic activity.
References
Friend SH, Bernards R, Rogelj S, et al. A human DNA segment with properties of the gene that predisposes to retinoblastoma and osteosarcoma. Nature 1986; 323:643–6.
Weinberg RA . The retinoblastoma protein and cell cycle control. Cell 1995; 81:323–30.
Dyson N . The regulation of E2F by pRB-family proteins. Genes Dev 1998; 12:2245–62.
Trimarchi JM, Lees JA . Sibling rivalry in the E2F family. Nat Rev Mol Cell Biol 2002; 3:11–20.
Nevins JR . Toward an understanding of the functional complexity of the E2F and retinoblastoma families. Cell Growth Differ 1998; 9:585–93.
Nevins JR . The Rb/E2F pathway and cancer. Hum Mol Genet 2001; 10:699–703.
Tan X, Martin SJ, Green DR, Wang JY . Degradation of retinoblastoma protein in tumor necrosis factor- and CD95-induced cell death. J Biol Chem 1997; 272:9613–6.
Bowen C, Birrer M, Gelmann EP . Retinoblastoma protein-mediated apoptosis after gamma-irradiation. J Biol Chem 2002; 277:44969–79.
Haas-Kogan DA, Kogan SC, Levi D, et al. Inhibition of apoptosis by the retinoblastoma gene product. EMBO J 1995; 14:461–72.
Haupt Y, Rowan S, Oren M . p53-mediated apoptosis in HeLa cells can be overcome by excess pRB. Oncogene 1995; 10:1563–71.
Zacksenhaus E, Jiang Z, Chung D, et al. pRb controls proliferation, differentiation, and death of skeletal muscle cells and other lineages during embryogenesis. Genes Dev 1996; 10:3051–64.
Chau BN, Borges HL, Chen TT, et al. Signal-dependent protection from apoptosis in mice expressing caspase-resistant Rb. Nat Cell Biol 2002; 4:757–65.
Munger K, Werness BA, Dyson N, et al. Complex formation of human papillomavirus E7 proteins with the retinoblastoma tumor suppressor gene product. EMBO J 1989; 8:4099–105.
Boyer SN, Wazer DE, Band V . E7 protein of human papilloma virus-16 induces degradation of retinoblastoma protein through the ubiquitin-proteasome pathway. Cancer Res 1996; 56:4620–4.
Berezutskaya E, Bagchi S . The human papillomavirus E7 oncoprotein functionally interacts with the S4 subunit of the 26 S proteasome. J Biol Chem 1997; 272:30135–40.
Berezutskaya E, Yu B, Morozov A, Raychaudhuri P, Bagchi S . Differential regulation of the pocket domains of the retinoblastoma family proteins by the HPV16 E7 oncoprotein. Cell Growth Differ 1997; 8:1277–86.
Kalejta RF, Shenk T . Proteasome-dependent, ubiquitin-independent degradation of the Rb family of tumor suppressors by the human cytomegalovirus pp71 protein. Proc Natl Acad Sci U S A 2003; 100:3263–8.
Nagao T, Higashitsuji H, Nonoguchi K, et al. MAGE-A4 interacts with the liver oncoprotein gankyrin and suppresses its tumorigenic activity. J Biol Chem 2003; 278:10668–74.
Krzywda S, Brzozowski AM, Higashitsuji H, et al. The crystal structure of gankyrin, an oncoprotein found in complexes with cyclin-dependent kinase 4, a 19 S proteasomal ATPase regulator, and the tumor suppressors Rb and p53. J Biol Chem 2004; 279:1541–5.
Kehn K, Fuente Cde L, Strouss K, et al. The HTLV-I Tax oncoprotein targets the retinoblastoma protein for proteasomal degradation. Oncogene 2005; 24:525–40.
Uchida C, Miwa S, Kitagawa K, et al. Enhanced Mdm2 activity inhibits pRB function via ubiquitin-dependent degradation. EMBO J 2005; 24:160–9.
Fattman CL, An B, Dou QP . Characterization of interior cleavage of retinoblastoma protein in apoptosis. J Cell Biochem 1997; 67:399–408.
Boutillier AL, Trinh E, Loeffler JP . Caspase-dependent cleavage of the retinoblastoma protein is an early step in neuronal apoptosis. Oncogene 2000; 19:2171–8.
Fattman CL, Delach SM, Dou QP, Johnson DE . Sequential two-step cleavage of the retinoblastoma protein by caspase-3/-7 during etoposide-induced apoptosis. Oncogene 2001; 20:2918–26.
Lu X . The isolation and structure of tripchlorolide (T4) from Tripterygium wilfordii. Zhongguo Yi Xue Ke Xue Yuan Xue Bao 1990; 12:157–61.
Yao Q, Zhang N . Effects of tripchlorolide (T4) of Tripterygium wilfordii Hook on the proliferation of peripheral blood mononuclear cells of rheumatoid arthritis patients. Zhongguo Yi Xue Ke Xue Yuan Xue Bao 1994; 16:352–5.
Zhang L, Bi Z, Li X . Inhibitory effects of monomer T4 from Tripterygium wilfordii hook on cultured mesangial cells proliferation and IL-1 production. Zhongguo Yi Xue Ke Xue Yuan Xue Bao 1994; 16:270–4.
Ye W, Den Y, Huang Y, Xue S . Antispermatogenic effect of Tripterygium wilfordii and tripchlorolide (T4) on rat gametogenesis and spermatozoa. Chin Med Sci J 1994; 9:110–3.
Ren Y, Xiong L, Wu JR . Induction of mitochondrion-mediated apoptosis of CHO cells by tripchlorolide. Cell Res 2003; 13:295–300.
Jin Y, Wei Y, Xiong L, Yang Y, Wu JR . Differential regulation of survivin by p53 contributes to cell cycle dependent apoptosis. Cell Res 2005; 15:361–70.
Jiang MR, Li YC, Yang Y, Wu JR . c-Myc degradation induced by DNA damage results in apoptosis of CHO cells. Oncogene 2003; 22:3252–9.
Collins AR . The comet assay for DNA damage and repair: principles, applications, and limitations. Mol Biotechnol 2004; 26:249–61.
Li D, Day KV, Yu S, et al. The role of adenovirus-mediated retinoblastoma 94 in the treatment of head and neck cancer. Cancer Res 2002; 62:4637–44.
Smith EJ, Leone G, Nevins JR . Distinct mechanisms control the accumulation of the Rb-related p107 and p130 proteins during cell growth. Cell Growth Differ 1998; 9:297–303.
Wang J, Sampath A, Raychaudhuri P, Bagchi S . Both Rb and E7 are regulated by the ubiquitin proteasome pathway in HPV-containing cervical tumor cells. Oncogene 2001; 20:4740–9.
Tsurugi K, Motizuki M, Mitsui K, Endo Y, Shiokawa K . The metabolism of ribosomal proteins microinjected into the oocytes of Xenopus laevis. Exp Cell Res 1988; 174:177–87.
Zhang M, Buckley DJ, Lavoi KP, Buckley AR, Blake MJ . Heat-induced proteolysis of HSF causes premature deactivation of the heat shock response in Nb2 lymphoma cells. Cell Stress Chaperones 1998; 3:57–66.
Hill TW, Loprete DM, Vu KN, Bayat M, Hardin LV . Proteolytic release of membrane-bound endo-(1,4)-beta-glucanase activity associated with cell wall softening in Achlya ambisexualis. Can J Microbiol 2002; 48:93–8.
Shiio Y, Yamamoto T, Yamaguchi N . Negative regulation of Rb expression by the p53 gene product. Proc Natl Acad Sci U S A 1992; 89:5206–10.
Gottlieb E, Oren M . p53 facilitates pRb cleavage in IL-3-deprived cells: novel pro-apoptotic activity of p53. EMBO J 1998; 17:3587–96.
Matozaki T, Sakamoto C, Matsuda K, et al. Missense mutations and a deletion of the p53 gene in human gastric cancer. Biochem Biophys Res Commun 1992; 182:215–23.
Fattman CL, An B, Sussman L, Dou QP . p53-independent dephosphorylation and cleavage of retinoblastoma protein during tamoxifen-induced apoptosis in human breast carcinoma cells. Cancer Lett 1998; 130:103–13.
Bates S, Phillips AC, Clark PA, et al. p14ARF links the tumour suppressors RB and p53. Nature 1998; 395:124–5.
Yap DB, Hsieh JK, Chan FS, Lu X . mdm2: a bridge over the two tumour suppressors, p53 and Rb. Oncogene 1999; 18:7681–9.
Rogoff HA, Pickering MT, Debatis ME, Jones S, Kowalik TF . E2F1 induces phosphorylation of p53 that is coincident with p53 accumulation and apoptosis. Mol Cell Biol 2002; 22:5308–18.
Sherr CJ, McCormick F . The RB and p53 pathways in cancer. Cancer Cell 2002; 2:103–12.
Acknowledgements
We thank Dr. Yuan Chao LI of Institute of Materia Medica, SIBS, CAS who kindly provided TC compound. We thank Dr. Xue Liang ZHU of Institute of Biochemistry and Cell Biology, SIBS, CAS who kindly provided p44-2L RB plasmid. This work was supported by a grant of National Natural Science Foundation #30230110, a grant of Science and Technology Commission of Shanghai Municipality #04DZ14901, a grant of Chinese Academy of Sciences #KSCX2-SW-203, and a grant of the Shanghai-Hong Kong-Anson Research Foundation for CAS and CUHK (Chinese University of Hong Kong) in Molecular Biosciences (SHARF) to Jia Rui WU.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
JIN, Y., LEUNG, W., SUNG, JY. et al. p53-independent pRB degradation contributes to a drug-induced apoptosis in AGS cells. Cell Res 15, 695–703 (2005). https://doi.org/10.1038/sj.cr.7290339
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7290339