ABSTRACT
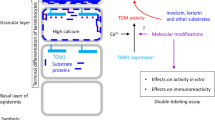
Epidermal-type transglutaminase 3 (TGM3) is involved in the cross-linking of structural proteins to form the cornified envelope in the epidermis. In the present study, we detected the expression of TGM3 in the mouse embryo using RT-PCR. TGM3 mRNA is weakly presented from E11.5 to E14.5 and increases significantly from E15.5 to birth. Then we determined the spatial and temporal expression pattern of TGM3 in the skin and other organs by in situ hybridization. We found a deprivation of TGM3 in skin at E11.5, while a rich supply in periderm cells and a weak expression in basal cells from E12.5 to E14.5. From the period of E15.5 to E16.5, after keratinization in the epidermis, TGM3 was expressed in the granular and cornified layers. The electron microscopic observation of the C57BL/6J mouse limb bud skin development provided several morphological evidences for the epidermal differentiation. The above findings suggest that the expression of TGM3 plays a important role in the epidermis differentiation in embryogenesis.
Similar content being viewed by others
INTRODUCTION
Transglutaminases (TGase: EC 2.3.2.13) are enzymes that catalyze the Ca2+ dependent cross-linking reaction between a g-carboxyamide group of glutamine and an o-amino group of lysine or other primary amines 1, 2, 3, 4. To date, eight human TGase isozymes (Factor XIII, TGases 1-/7) have been found comprising a large protein family 5. Transglutaminase 3 (TGM3; epidermal-type TGM), a member of the transglutaminase family, is found to be expressed during the late stages of the terminal differentiation of the epidermis and in certain cell types of the hair sheath cells 6, 7, 8. TGM3 is predominantly involved in the formation of the cornified cell envelope, where it cross-links with various structural proteins such as involucrin, loricrin and small proline-rich proteins during epidermal terminal differentiation 9, 10. Therefore, TGM3 takes a critical part in skin formation because the cornified envelope acts as a barrier function. The native TGM3 protein molecule contains 692 amino acids and has a molecular mass of 77 kD, both in human and mouse 11. Similar to TGase 1 and Factor XIII, the activation of TGM3 during keratinocyte differentiation requires the cleavage of a 77 kD zymogen by an unknown protease to release the 50 kD and 27 kD components, which remain associated together in the active enzyme 12. Although TGM3 mRNA represents less than 2% of the TGase transcripts, the activated TGM3 accounts for up to 75% of the total TGase activity in mammalian epidermis 13.
Although the structural and biochemical characterization of recombinant TGM3 has been reported, the distribution of TGM3 in different embryonic periods and various organs development has not been reported so far. Previous studies detected TGM3 mRNA and protein expression and their enzymatic regulation in most organs of male balb/c mice (12-14 w) 14. In order to further analyze the gene expression and enzymatic regulation of TGM3 in embryogenesis, we attempted to clarify the tissue distribution and to establish the expressing orderliness of TGM3 in mouse embryonic development.
In this study, we investigated TGM3 tissue distribution in developing mouse embryos and adult male mouse tissues by RT-PCR and in situ hybridization analysis. Then the skin ultrastructure in different embryonic periods was examined by electron microscopy.
MATERIALS AND METHODS
Animals and tissues
Time-mated C57BL/6J mice were obtained from the Laboratory Animal Institute, Chinese Academy of Medical Sciences. Mice were obtained from in-house breeding programs and maintained on a 12:12 h light-dark cycle and provided standard laboratory food and water. All manipulations of mice were done in accordance with policies of the Institute Animal Care and Use Committee. Timed mating was set up between adult mice. Females were inspected for plugs on the following day to ensure successful mating. The day of plug detection was considered as E0.5. Embryos were dissected from pregnant mice at the age of embryonic day 11.5, 12.5, 13.5, 14.5, 15.5, 16.5, 17.5, 18.5, and postnatal day 1 (P1). The adult male C57BL/6J mice used in this study was about 26 g. Three samples per time point were taken and analyzed for statistical significance test. The tissues were taken and immediately frozen in liquid nitrogen until analysis.
RNA extraction
Total RNA was extracted from frozen tissues by using TRIzol reagent following the protocols of the manufacturer (GIBCO/BRL, NY, USA). The integrity of the RNA was assessed by denaturing agarose gel electrophoresis and spectrophotometry.
Semi-quantitative RT-PCR
Five micrograms of total RNA of each sample was used to synthesize the first strand cDNA with SuperScript Preamplification System by the First Strand cDNA Synthesis kit (GIBCO/BRL, NY, USA). Then 1 μl of RT product was used as the template to amplify TGM3. The cycle number for PCR was adjusted so that the reactions fell within the linear range of product amplification. The resulting cDNAs were subjected to PCR reaction with primers designed to amplify TGM3 DNAs. 5′-primer and 3′-primer were 5′-CGCAACATCTTCGAGGAATC-3′ (mouse TGM3:1455-1475) and 5′-TCCTTCCACACTTCGTGGACAA-3′ (mouse TGM3:1664-1642), respectively. The expression of housekeeping gene, glyceraldehyde-3-phosphate dehydrogenase (GAPDH), was used as an internal control. Its primers were 5′-ACCACAGTCCATGCCATCAC-3′ and 5′-TCCACCACCCTGTTGCTGTA-3′. The RT-PCR reaction product was analyzed by electrophoresis on a 1.5% agarose gel. The electrophoresis images were scanned by Fluor-S MultiImager (Bio-Rad, CA, USA) and the original intensity of every specific band was quantitated with the software Multi-Analyst (Bio-Rad, CA, USA). The data were compared after being normalized by the intensity of GAPDH.
In situ hybridization of staged murine embryos
For in situ hybridization, paraffin-embedded mouse tissue sections were deparaffinized in xylene and rehydrated through a graded ethanol series. After proteinase K digestion (18 μg/ml), the sections were post-fixed with 4% (w/v) paraformaldehyde in PBS for 10 min and treated with 0.1 M triethanolamine–HCl (pH 8.0) for 1 min. Following acetylation for 10 min, the sections were dehydrated, air-dried and then incubated overnight at 50°C in hybridization buffer composed of 50% formamide, 10 mM Tris–HCl (pH 7.5), 1 mg/ml yeast tRNA (Sigma Co.), 1 × Denhardt's solution (Sigma Co.), 10% PEG6000, 600 mM NaCl, 0.25% SDS, 1 mM EDTA, and 2 μg/ml probe. After hybridization, the sections were washed at 45°C for 1 h in 50% formamide and 2 × SSC, and digested with 20 μg/ml RNase (Sigma Co.) in 10 mM Tris–HCl (pH 8.0) and 500 mM NaCl at 37°C for 30 min. Hybridized DIG-labeled probes were visualized with the Nucleic Acid Detection Kit (Boehringer Mannheim).
Electron microscopy
For ultrastructural evaluation of skin, tissue was rinsed in PBS, fixed for 4 h in 2% glutaraldehyde/2% formaldehyde in 0.05 M cacodylate buffer, pH 7.3, washed 60 min in the buffer, and post-fixed for 2 h in 1% osmium tetroxide. The specimens were dehydrated in ethanol and acetone, then embedded in POLY/BED 812 (Polysciences, Warrington, PA), mounted into flat molds, and polymerized at 65°C. Silver color thin sections were cut on a Reichert Ultracut E, using a diamond knife (Diatome, Switzerland). The sections were stained with 2% uranyl acetate/lead citrate and viewed under H7500 transmission electron microscope.
RESULTS
Tissue distribution of mouse TGM3 detected by RT-PCR analysis
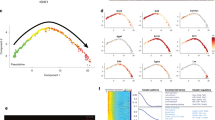
The expression of TGM3 mRNA was analyzed by semi-quantitative RT-PCR. The primers were designed to enable specific amplification of mouse TGM3 cDNA. During the mouse embryonic development, TGM3 mRNA expression was very weak from E11.5 to E14.5 and began to increase significantly from E15.5 to birth (Fig. 1A). Although TGM3 is widely expressed in many tissues, striking changes in its expression were observed between E17.5 and after birth, depending on different organs tested Its expression in liver and heart was stronger in E17.5 than that after birth. On the other hand, the expressions of TGM3 in brain and skin were stronger after birth than that in E17.5, and a similar expression level was detected in lung between the two stages (Fig. 1B). Furthermore, the expression of TGM3 was also analyzed in different organs of adult mouse. We found that besides brain and esophagus, TGM3 was highly expressed in testis. No significant changes in other tissues were detected (Fig. 1C).
The expression of TGM3 mRNA was analyzed by semi-quantitative RT-PCR. (A) Expression of TGM3 in different stages of embryos and newborn of C57BL/6J mice was studied by semi-quantitative RT-PCR. (B) Differential expression of TGM3 in different organs of C57BL/6J mice was studied by semi-quantitative RT-PCR between the period of E17.5 and newborn. H: heart, L1: lung, B: brain L: liver, S: skin. (C) Expression of TGM3 in different organs of adult male C57BL/6J mice was studied by semi-quantitative RT-PCR.
Expression of TGM3 in C57BL/6J mouse embryonic organs
The TGM3 distribution in different stages of mouse embryos were analyzed using in situ hybridization. In the single-layered and early two-layered epidermis (E10.5–E11.5), in situ TGase expression was not observed in the periderm cells of the single-layered epidermis, and begins to be weakly presented at stage E11.5. In the three-layered (E12.5–E13.5) and in four or more layered (E14.5) stratified epidermis, in situ TGase expression was mainly localized in the periderm cells and weakly in the basal cells. After keratinization in the epidermis (E15.5–E17.5), TGase was detected in the granular, cornified layers and hair follicle (Fig. 2). In addition to the skin, the expressions of TGM3 in heart, lung, bone, muscle, testis and blood vessel were also assessed (Fig. 3, 4 and Tab. 1).
Expression of TGM3 in developing mouse epidermis by in situ hybridization. TGM3 was observed mainly in the periderm cells (arrows) and weakly in the basal cells in stages of E12.5 (A) and E13.5 (B). In E16.5, TGM3 activity was detected in the granular, cornified layers (arrows) (C) and hair follicles (arrows) (D).
Ultrastructural changes of epidermis during C57BL/6J mouse embryonic development
Four stages of limb bud skin development were established on the basis of EM observation (Fig. 5). Stage I (E10.5): The epidermis lying on the continuous basement membrane is consisted of a single layer of overlapping flat cells. There is a wide intercellular space between the neighboring cells. Some mitochondria and rough endoplasmic reticulum can be seen in the cytoplasm. Stage II (E11.5): The very flattened periderm cells are formed by differented epidermal cells. Rough endoplasmic reticulum, abundant free ribosome and mitochondria could be found in the cytoplasm of periderm cells. The region between periderm cells and underlying polygonal cells that arranged in a lattice-like pattern is high electron dense. At the same time, there is a wide intercellular space in the polygonal basal cell layer. Stage III (E12.5–13.5): The epidermis differentiated into three or four layers from the deep to the surface. In spite of the cuboid basal cells and flat periderm cells, the polygonal cells of the intermediate layer begin to appear in the epidermis. Hemidesmosomes began to form in this stage. The wide intercellular space that is usually found in the earlier stages during the epidermal differentiation becomes less obvious between the intermediate cells and basal cells. Cytoplasmic glycogen particles can occasionally be found in the intermediate cells. Stage IV (E15.5–17.5): In E15.5, a differentiated horny layer in the epidermis appears and abundant large keratohyalin granules can be seen in the granular cells. The fetal epidermis at E17.5 became keratinized, consisting of a basal layer, 2 or 3 spinous layers of cell, the granular layer with distinct keratohylin, and 5-6 layers stratum corneum. At this stage, no remnant of periderm was left and the epidermis contains all the characteristics of an adult epidermis.
Ultrastructure of the five stages of developing limb bud epidermis. (A) In E10.5, epidermal cells containing mitochondria and endoplasmic reticulum were connected to each other. A wide intercellular space is present in the epidermis. (B) In E11.5, the flattened periderm cell differentiated from the epidermal cells. They contain rough endoplasmic reticulum, abundant free ribosome and mitochondria. Wide intercellular space can still be seen between basal cells. A continuous basement membrane (BM) lied beneath the epidermis. (C) In E13.5, the epidermis was consisted of 3 or 4 layers of cells. An intermediate layer consisted of polygonal cells (PC) began to appear and hemidesmosomes began to form in this stage. (D) In E15.5, a differentiated horny layer (HL) of the epidermis appeared. The wide intercellular space disappeared in this period. (E) Epidermis at E17.5 became keratinized (K) and vesicles could be seen in the cells of horny layer. The structure of this epidermis was quite similar to that of the adult epidermis.
DISCUSSION
It was found by RT-PCR and in situ hybridization that TGM3 was expressed in the periderm from E11.5. The electron microscopy showed that the periderm first appeared as a layer of simple squamous cells, underwent a series of changes that alter both of the surface morphology and the subcellular organization, and then disappeared when epidermal keratinization was complete. At the same time, the sequential expression patterns of cornified cell envelop (CCE) precursor proteins were closely associated with the thickening of periderm cell membrane, as revealed by electron microscopy 15. According to the expression pattern of TGM3, we hypothesize that CCE must begin to form in the periderm cells from the stage of two-layered epidermis.
Although TGM3 is required for cross-linking structural proteins in the skin and the hair follicle, its expression in other organs has not been sufficiently investigated 16. A rich supply of TGM3 in brain and testis tissues were detected in this study. In other studies, TGase expression in human brain tissue has revealed that not only the expression of TGase 2 but of TGases 1 and TGases 3 are commonly found in stratified squamous epithelia 17, 18. In brain, TGase was demonstrated to be involved in various signal transduction pathways, particularly in the regulation of neurotransmitter release 19. Our results agree with the presence of TGM3 and its important roles in mouse brain. However, elucidation of the differential expression of TGM3 in different regions of the brain during its development will provide a better understanding of the nature of the TGase enzyme.
At the stage of E12.5 to E14.5, TGM3 was mainly expressed in the testis tunica albuginea. As we know, tunica albuginea, a dense fibrous connective tissue capsule forms a covering for the testis itself. To our knowledge it is the first time showing the higher level expression of TGM3 in adult testis, which indicats an important role in spermatogenesis. In testis, TGase activity was also been reported to be involved in the interaction of Follicle-Stimulating Hormone (FSH) with its receptor 20. Furthermore, it was reported that rat epididymal sperm immunogenicity could be suppressed by transglutaminase both in vivo and in vitro. The suppression of the epididymal sperm immunogenicity is suggested to be of crucial importance to prevent the immune response when the sperm is introduced into the immunocompetent female genital tract during coitus 21.
The electron microscopic observation of the C57BL/6J mouse limb bud skin development provided several morphological evidences for the epidermal differentiation. The epidermal thickening becomes apparent during its development, and is mainly due to elongation of basal cells and the development of intermediate layer that was proliferated from basal cells. Wide intercellular space was observed and some phagosomes were present in the epidermis during its early developmental period. Therefore, we would conclude that in its early development period, the epidermal cells may exchange substances with the amniotic fluid to change its composition, rather than to act as a barrier against the amniotic fluid. With the differentiation of the epidermis processes, the wide intercellular space that is usually found in the earlier stages becomes to be reduced. When a horny layer is formed in the E15.5, the permeability of the epithelium decrease. Similar results have been found in humans 22, rabbits 23 and rats 24.
Abbreviations
- TGM3:
-
transglutaminase 3
- CCE:
-
cornified cell envelope
- DEPC:
-
diethylpyrocarbonate
- DIG:
-
digoxigenin
- GAPDH:
-
glyceraldehyde-3-phosphate dehydrogenase
References
Folk JE . Transglutaminases. Annu Rev Biochem 1980; 49: 517–31.
Aeschlimann D, Paulsson M . Transglutaminases: protein crosslinking enzymes in tissues and body fluids. Thromb Haemost 1994; 71:402–15.
Greenberg CS, Birckbichler PJ, Rice RH . Transglutaminases: multifunctional crosslinking enzymes that stabilize tissues. FASEB J 1991; 5:3071–7.
Ichinose A, Bottenus RE, Davie EW . Structure of transglutaminases. J Biol Chem 1990; 265:13411–4.
Grenard P, Bates MK, Aeschlimann D . Evolution of transglutaminase genes: identification of a transglutaminase gene cluster on human chromosome 15q15. J Biol Chem 2001; 276:33066–78.
Martinet N, Kim HC, Girard JE, et al. Epidermal and hair follicle transglutaminases. Partial characterization of soluble enzymes in newborn mouse skin. J Biol Chem 1988; 263:4236–41.
Kim HC, Lewis MS, Gorman JJ, et al. Protransglutaminase E from guinea pig skin. Isolation and partial characterization. J Biol Chem 1990; 265:21971–8.
Kim HC, Idler WW, Kim IG, et al. The complete amino acid sequence of the human transglutaminase K enzyme deduced from the nucleic acid sequences of cDNA clones. J Biol Chem 1991; 266:536–9.
Kalinin AE, Kajava AV, Steinert PM . Epithelial barrier function: assembly and structural features of the cornified envelope. BioEssays 2002; 24:789–800.
Kalinin A, Marekov LN, Steinert PM . Assembly of the epidermal cornified cell Envelope. J Cell Sci 2001; 114:3069–70.
Kim IG, Gorman JJ, Park SC, Chung SI, Steinert PM . The deduced sequence of the novel protransglutaminase E (TGase3) of human. J Biol Chem 1993; 268:12682–90.
Ahvazi B, Kim HC, Kee SH, Nemes Z, Steinert PM . Three dimentional structure of the human transglutaminase 3 enzyme: binding of calcium ions changes structure for activation. EMBO J 2002; 21:2055–67.
Kim SY, Chung SI, Steinert PM . Highly active soluble processed forms of the transglutaminase 1 enzyme in epidermal keratinocytes. J Biol Chem 1995; 270:18026–35.
Hitomi K, Horio Y, Ikura K, Yamanishi K, Maki M . Analysis of epidermal-type transglutaminase (TGM3) expression in mouse tissues and cell lines. Int J Biochem Cell Biol 2001; 33:491–8.
Akiyama M, Smith LT, Yoneda K, et al. Periderm cells form cornified cell envelope in their regression process during human epidermal development. J Invest Dermatol 1999; 112:903–9.
Hitomi K, Presland RB, Nakayama T, et al. Analysis of epidermal-type transglutaminase (transglutaminase 3) in human stratified epithelia and cultured keratinocytes using monoclonal antibodies. J Dermatol Sci 2003; 32:95–103.
Selkoe DJ, Abraham C, Ihara Y . Brain transglutaminase in vitro crosslinking of human neurofilament proteins into insoluble polymers. Proc Natl Acad Sci U S A 1982; 79:6070–4.
Kim SY, Grant P, Lee JH, Pant HC, Steinert PM . Differential expression of multiple transglutaminases in human brain. Increased expression and cross-linking by transglutaminases 1 and 2 in Alzheimer's disease. J Biol Chem 1999; 274:30715–21.
Pastuszko A, Wilson DF, Erecinska M . A role for transglutaminase in neurotransmitter release by rat brain synaptosomes. J Neurochem 1986; 46:499–508.
Grasso P, Reichert LE Jr . Stabilization of follicle-stimulating hormone-receptor complexes may involve calciumdependent transglutaminase activation. Mol Cell Endocrinol 1992; 87:49–56.
Peluso G, Porta R, Esposito C, et al. Suppression of rat epididymal sperm immunogenicity by a seminal vesicle secretory protein and transglutaminase both in vivo and in vitro. Biol Reprod 1994; 50:593–602.
Holdrook KA, Odland GF . The fine structure of developing human epidermis: light, scanning, and transmission electron microscopy of the periderm. J invest Derm 1975; 65:16–38.
Maruyama T, Yoshizuka M, Fujimoto S . Light and electron microscopy of fetal rabbit skin with special reference to the role of mesenchymal cells in epidermal differentiation. Acta Anat (Basel) 1988; 133:143–55.
Bauer FW . Differentiation and keratinization of fetal rat skin. II. Ultrastructural study of the epidermis in vivo and in vitro. Dermatologica 1972; 145:17–36.
Acknowledgements
This work was supported by Major State Basic Research Development program of China (2004CB518604), the National High Technology Research and Development Program of China (2004AA231041), and the National Natural Science Foundation of China (30425027). We also thank Prof. Chuan Nong Zhou for his kind assistance in preparing this manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
ZHANG, J., ZHI, H., DING, F. et al. Transglutaminase 3 expression in C57BL/6J mouse embryo epidermis and the correlation with its differentiation. Cell Res 15, 105–110 (2005). https://doi.org/10.1038/sj.cr.7290274
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7290274