ABSTRACT
Human c-myc cDNA was fused with the hormone-binding domain (HBD) cDNA of murine estrogen receptor gene and the chimeric gene was introduced into the CHO cells. The fusion protein, c-MycER, becomes activated when the synthetic steroid, 4-hydroxy-tamoxifen (OHT), binds HBD. Activated c-MycER, likely c-Myc, can induce quiescent CHO cells reentry into S phase and subsequent cell death under serum-free condition. In addition, the expression of some proposed c-myc target genes such as ODC, MrDb, cad, rcc1 and rcl were found to increase upon OHT induction before S phase entry and apoptosis, indicating that these target genes are involved in cell cycle regulation and/or apoptosis control. However, the mutant D106-143c-MycER protein does not have above activities.
Similar content being viewed by others
INTRODUCTION
The c-myc proto-oncogene, the cellular homologue of the gene born by the retrovirus MC29, is responsible for chicken leukemia1, and it originally got its name for the ability to causemyelocytomatosis in susceptible animals and cultured cells2. c-myc is one of the most widely studied and the best-characterized proto-oncogene. Although c-myc has been shown closely related to cell growth, differentiation, cell cycle progression, tumorigenesis and apoptosis, the authentic functions of c-Myc protein have still been obscure. However, the elucidation of bHLHLZ structure in c-Myc and the discoveries of partner proteins of c-Myc in recent years promoted a new approach to understand this “old” proto-oncogene. C-Myc protein, likely other bHLHLZ proteins, possesses specific DNA-binding activity to CACGTG (E-box) sequence. C-Myc is therefore supposed to function as a transcription regulator, while cooperating with its partner(s) including Max and Mad family3. It is agreed that c-Myc dimerizes with Max to bind CACGTG or similar consensus sequence of down stream target genes thereby regulating transcription and subsequently promoting cell proliferation and apoptosis, as well as inhibiting cell differentiation. Dozen of c-myc target genes have been proposed including p534, ODC5, α-prothymosin6, ECA397, cad8, eIF-2a9, cdc25A10, MrDb11, rcc112, rcl13 and TERT14,15. The list is increasing each year.
To understand how c-myc involved in cell growth and death, we established a function-inducible c-myc over expression system in CHO cells in which human c-myc was fused with the hormone binding domain (HBD) cDNA of estrogen receptor gene16. The produced fusion protein is called c-MycER. The c-MycER system was first brought forward by Eilers in 19916 and modified by Littlewood in 199516. It has been proven that c-MycER fusion protein exhibited all c-Myc activities, which was closely dependent upon the synthetic steroid 4-hydroxy-tamoxifen (OHT), yet was totally unresponsive to endogenous steroid and phenol red in medium. D106-143c-myc, encoding a 106-143 amino acid deleted c-Myc, was also fused with HBD cDNA and transfected CHO cells. D106-143c-Myc protein showed no c-Myc activities. We found that the transcriptional expressions of some proposed c-myc target genes including ODC, MrDb, cad, rcc1 and rcl were increased upon OHT induction in CHO-c-mycER cells. In addition, only wild type quiescent CHO-c-mycER cells, but not CHO-D106-143c-mycER, CHO-neo or parent CHO cells, could reenter the S phase and subsequently showed apoptotic phenotype after OHT activation. Most importantly, the transcription activation of c-myc target genes was much earlier than the cell behavior change in CHO-c-mycER cells.
MATERIALS AND METHODS
Construction of expression plasmid
The 2.4 Kb wild type c-mycER or 2.3 Kb mutant type D106-143c-mycER cDNAs (gifts of Littlewood) were inserted to the EcoRI site of pcDNA 3 (invitrogen) respectively. The correct orientation was confirmed by sequencing with T7 sequencingTM kit (Pharmacia) according to producer's instruction except for using T7 sequencing primer. The resulting plasmids were named as pcDNA3/c-mycER and pcDNA3/D106-143c-mycER.
Cell culture and establishment of cell lines
Chinese Hamster Ovarian (CHO) cells (Cell Bank, CAS) were grown in Dulbecco's modified Eagle's medium (DMEM, Gibco) supplemented with 10% heat-inactivated newborn bovine serum (NBS, Evergreen), 100 units/ml penicillin, and 100 μg/ml streptomycin. Cells were passaged by standard trypsinization at 70-80% confluence and seeded at 1:4 dilution directly onto tissue culture dishes or flasks. Culture medium was changed every 2 or 3 d to maintain good growth condition. To generate quiescent CHO cells, 2 × 105 exponentially growing cell were seeded to 25 cm2 flask, medium was changed at 30% confluence (2 d). After 100% confluence (4-5 d), cells were kept in the serum-free medium for another 2 d before they were used for the experiments.
To establish CHO cells with highly expressed wild type c-mycER or mutant type D106-143c-mycER, CHO cells were transfected with pcDNA3/c-mycER or pcDNA3/D106-143c-mycER by the calcium-phosphate method17. The cells were cultured for 3 weeks in selection medium [800 μg/ml Geneticin (Gibco)] to select stable transfectants. CHO cells were also transfected with pcDNA3 to obtain the CHO-neo control cells. Individual clones were isolated by cloning rings method18. Expansion of isolated clones was carried out sequentially in 96-well plate, 24-well plate and finally 25 cm2 flask in selection medium. The expression of c-mycER or D106-143 c-mycER mRNA in the clones was determined by Northern blot analysis. The c-mycER or D106-143 c-mycER positive clones were then subcloned to get genetically stable and homogeneous cell lines.
Blot analysis
Total cellular RNA was isolated from cells by acid guanidinium thiocyanate phenol-chloroform extraction method19. 30 μg of total RNA was size-fractionated on formaldehyde agarose gel, transferred to nylon filter. To do reverse Northern dot hybridization, 100 ng of each cDNA fragment was denatured and dotted directly onto nylon filter. The filter containing DNA or RNA was baked at 80°C for 2 h. The cross-linked membrane was incubated in prehybridization solution (6 × SSC, 5 × Denhardt's, 0.5% SDS and 100 μg/ml denatured salmon DNA) at 68°C for 4 h, then hybridized with 32P-labelled, denatured DNA probe in hybridization solution (6 × SSC, 0.5% SDS, 100μg/ml denatured salmon DNA) at 65°C for 16-24 h. After hybridization, the filter was washed extensively twice in 2 × SSC/0.1%SDS at 37 °C for 20 min, then twice in 0.2 × SSC/0.1%SDS at 42 °C for 20 min, and finally in 0.1 × SSC/0.1 % SDS at 55°C for 15 min. The washed membrane was autoradiographed for 1-2 d at −70 °C using X-ray film (Kodak) with intensifying screen. The c-myc probe was labeled with a-32P-dCTP (Amersham) using Primer-a-Gene labeling system (Promega) according to producer's instruction. To prepare total cDNA probes for reverse Northern dot hybridization, 5m g mRNA was isolated from total RNA using Quickprep MicromRNA purification kit (pharmacia) and a-32P-dCTP was integrated into the cDNA by reverse transcription with 5 μg oligo(dT)18 as primer.
Morphological, biochemical and quantified analysis of apoptosis
Cells were starved to quiescence as described previously. Medium was changed to serum-free DMEM with or without 200 nM 4-hydroxy-tamoxifen (OHT, Sigma). After 24 h culture, microphotographs were taken and then cells were harvested using trypsin, fixed with 75% ethanol in PBS, and stained for 15 min with 20 μg/ml propidium iodide (PI, Sigma) and 180 μ/ml RNaseA. PI-stained cells were observed with fluorescence microscopy to identify apoptosis by nuclear morphology, or were analyzed by flow cytometry to quantify apoptosis. To identify apoptosis by biochemical character, genomic DNA was extracted from 1 × 106 quiescent, OHT-induced cells and electrophoresed in 2.0% agarose. The DNA “laddering” was observed by ethidium bromide (EB, Sigma) staining.
RESULTS
Establishment of c-myc overexpressing CHO cell line
Plasmid pcDNA3, pcDNA3/c-mycER or pcDNA3/D106-143c-mycER was introduced respectively to CHO cells by calcium phosphate-DNA coprecipitation method. After G418 selection, individual clones were isolated and expanded. Cellular RNA was extracted and Northern blot hybridization was followed. The representative results were shown in Fig 1. In comparison with parent CHO cells (Fig 1, lane 1) and CHO-neo cells (Fig 1, lane 2), CHO-c-mycER cells overexpressed c-mycER mRNA (Fig 1, lane 3) and CHO-D106-143c-mycER cells overproduced D106-143c-mycER transcript (Fig 1, lane-4). However, quiescent parent CHO cells and CHO-neo cells have weakly detected endogenous c-myc expression.
Northern blot hybridization of c-mycER or D106-143c-mycER over-expressed CHO transfectants. Upper part, c-myc cDNA was used as probe to detect the expression of c-mycER or D106-143c-mycER in CHO-c-mycER cells (lane 3), CHO-D106-143c-mycER cells (lane 4), parent CHO cells (lane 1) and control vector transfected CHO cells (lane 2). Lower part, b-actin probe hybridization was used as RNA loading control in different cell samples.
OHT-dependent c-MycER-promoted transcription
As a transcription factor, c-Myc controls cell destiny by regulating the expression of downstream target genes. Northern blot hybridization was performed to detect the mRNA level of a representative c-myc target gene, ODC. Expression of ODC increased within 3 h after addition of OHT to quiescent CHO-c-mycER cells, and reached a maximum by 9 h. In contrast, addition of OHT to quiescent parent CHO cells did not induce expression of ODC (data not shown). The expression of ODC and other proposed c-myc target genes were further investigated by reverse Northern dot hybridization with mRNA at 9 h after OHT induction. The result shown in Fig 2 indicated that the expression of ODC, MrDb, cad, rcc1 and rcl were up-regulated.
OHT-dependent c-MycER-induced apoptosis
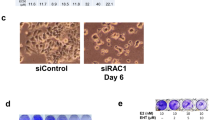
Deregulated expression of c-myc in the absence of growth factors induces apoptosis in fibroblasts. Accordingly, we performed further experiment to test this ability of c-MycER in CHO cells. Quiescent, serum-deprived CHO-c-mycER cells underwent death in response to OHT, as demonstrated by microscopic observation (Fig 3). The rate of cell death was determined by flow cytometry. As shown in Fig 4, CHO-c-mycER cells with OHT displayed a significant rate of apoptosis, whereas CHO-c-mycER cells without OHT and other cell lines with or without OHT all showed low rate of apoptosis which represented inherent low level cell death, not related to OHT induction. The dying cell exhibited nuclear condensation (Fig 5), a classic character of apoptosis. Degradation of cellular DNA into nucleosomal fragments, a frequent feature of apoptosis, was also evident (Fig 6).
DISCUSSION
In our present c-myc overexpressing system, c-Myc is rendered functionally OHT-dependent through fusion with the mutant HBD of estrogen receptor16. c-MycER protein is probably inactive in the absence of the ligand (OHT) because it complexes with a variety of intracellular polypeptides, of which Hsp90 is prototype. OHT's binding to HBD releases the chimeric protein from the inhibitory complexes.
Although other methods have been used to produce inducible c-myc (i. e. heat or Zn2+-inducible) expression system, c-MycER system, which is regulated at protein function level, offers several advantages20. The most significant merit is that the response of c-MycER to OHT is tightly controlled, relatively fast and readily reversible. The only drawback is that OHT can be toxic to some cells at concentrations above 1 μM. However, OHT is usually added to medium at concentrations far below the lethal limit (ranging from 1 to 200 nM).
As an immediate early gene, c-myc expression is rapidly induced following mitogenic stimulation and is absolutely dependent on growth factors. When serum or specific growth factors are withdrawn, c-myc expression is down-regulated and cells eventually accumulate in the G0/G1 phase of the cell cycle21. In order to investigate the roles of exogenous c-MycER on growth and death of CHO cells, it is necessary to arrest CHO cells in G0/G1 phase to repress the endogenous expression background of hamster c-myc. When confluent CHO cells were kept in serum-free medium for 2 days, more than 85% cells were arrested in G0/G1 phase, and endogenous c-myc expression under these conditions was weakly detected by Northern blot analysis (Fig 1). The quiescent cells could reentry into the cell cycle after serum stimulation or c-myc expression.
In the present experiment, wild type c-mycER or D106-143c-mycER was introduced into CHO cells to study their respective role in CHO cell growth, transformation and apoptosis. We find that (1) activation of c-MycER by OHT can and is sufficient to drive quiescent serum-deprived CHO cells into the cell cycle (Fig 7) and (2) OHT can promote anchorage-independent growth through activation of c-MycER (data not shown). The observation also suggests that c-MycER protein can induce apoptosis. All of the activities are closely dependent upon the OHT addition and upon the intact c-myc sequence because the D106-143c-mycER does not show any of them, due to its deletion in transactivation domain of c-Myc. However, D106-143c-MycER has a complete DNA binding domain, so it can function as a dominant-negative mutant to suppress the apoptosis caused by low expressed endogenous c-myc under serum deprived condition. As shown in Fig 4, CHO-D106-143c-mycER cells displayed a little lower cell death percentage compared to parent and CHO-neo cells. D106-143c-MycER also exerted a dominant-negative effect on S phase entry of CHO-D106-143c-mycER cells. C-myc controls both proliferation and apoptosis in mammalian cells. Deregulated expression of c-myc induces cells to undergo apoptosis, unless specific signals provided either by cytokines or oncogenes blocking the apoptotic pathway. C-MycER activation in quiescent CHO-c-mycER cells induced S phase reentry and subsequent apoptosis under serum-free medium. Our observation tends to support the “conflict model” theory which proposed that apoptosis is a conflict between “stop” (no growth factors) and “go” (but c-myc is on) signals22.
The expression of some c-Myc target genes was also determined by mRNA analysis. ODC is one of the most important Myc target genes. ODC is a rate-limiting enzyme in polyamine biosynthesis, required for DNA synthesis5. When overproduced, ODC can induce apoptosis (as Myc does) in cells lacking survival factor(s). Furthermore, inhibitors of ODC enzyme activity interfere with Myc-induced apoptosis5. We find that ODC is induced to express by OHT addition to the CHO-c-mycER cells. The expression reaches to the maximum at 9 h after OHT induction. The expression of other c-Myc target genes, including MrDb, cad, rcc1 and rcl, also have great change after OHT's adding to CHO-c-mycER cells. As downstream target genes, they may all contribute to the function of c-myc played in cell apoptosis in CHO-c-mycER system. Dozen of articles indicated that c-myc target genes were involved in DNA synthesis(ODC and cad), RNA metabolism (eIF-2α, eIF-4E, MrDb and rcc1), cell cycle control(p53 and cdc25A), and chromatin modification (μ-prothymosin and TERT) (for review see reference 23). However, the function of these genes and the cross-reaction of these genes in the apoptosis induction in CHO-c-mycER cells remained to be understood. Although some c-myc target genes show similar functions to c-myc signal pathway, they all can not substitute c-myc in cell growth and death. The cloning and characterization of new c-myc target gene(s) would contribute to the understanding of the authentic function of c-myc.
References
Lemaitre JM, buckle RS, Mechali M . c-myc in the control of cell proliferation and embryonic development. Adv in Cancer research 1996; 70:95–144.
Henrisksson M, Luscher B . Proteins of the myc network: essential regulators of ell growth and differentiation. Adv in Cancer Research 1996; 6:109–82.
Shi C, Wang P, Hu YJ, Xu L . Affinity chromatography-dependent selection (ACDS) of genomic DNA fragments bound specifically to bacterial synthesized Myc/Myn proteins. Cell Research 1995; 5:25–34.
Reisman D, Elkind NB, Roy B, Beamon J, Rotter V . c-myc transactivates the p53 promoter through a required downstream CACGTG motif. Cell Growth and Differentiation 1993; 4:57–65.
Bell-Fernander C, Packham G, Cleveland JL . The ornithine decarboxylase gene is a transcriptional target of c-myc. Proc Natl Acad Sci USA 1993; 90:7804–8.
Eilers M, Schism S, Bishop JM . The Myc protein activates transcription of the alpha-prothymosin gene. EMBO J 1991; 10:133–41.
Benvenisty N, Leder A, Kuo A, Leder P . An embryonically expressed gene is a target for c-my regulation via the c-myc-binding sequence. Genes Dev 1992; 6:2513–23.
Miltenberger RJ, Sukow KA, Farnham PJ . An E-box-mediated increase in cad transcription at the G1/S-phase boundary is suppressed by inhibitory v-myc mutants. Mol Cell Biol 1995; 15:2527–35.
Rosenwald IB, Rhoads DB, Callanan LD, Isselbacher KJ, Schmidt EV . Increased expression of eukaryotic translation initiation factors eIF-4Eand eIf-2alpha in response to growth induction by c-myc. Proc Natl Acad Sci USA 1993; 90:6175–8.
Galaktionov K, Chen XC, Beach D . Cdc25A cell cycle phosphatase as a target of c-myc. Nature 1996; 382:511–7.
Grandori C, Mac J, Siebelt F, Ayer DE, Eisenman RN . Myc-max heterodimers activate a DEAD box gene and interact with multiple E-box-related sites in vivo. EMBO J 1996; 15:4344–57.
Tsuneoka M, Nakano F, Ohgusu H, Mekada E . c-myc activates RCC1 gene expression through E-box elements. Oncogene 1997; 14:2301–11.
Lewis BC, Shim H, Li Q, Wu CS, Lee LA, Maity A, Dang CV . Identification of putative-myc-responsive genes: characterization of rcl, a novel growth-related gene. Mol Cell Biol 1997; 17:4967–78.
Wu KJ, Grandori C, Amacker M, Nathalie SV, Polack A, Lingner J, Riccardo DF . Direct activation of TERT transcription by c-MYC. Nature Genetics 1999; 21:220–4.
Greenberg RA, O'Hagan RC, Deng HY, Xiao QR, Hann SR, Adams RR, Lichtsteiner S, Chin L, Morin GB, DePinho RA . Telomerase reverse transcriptase gene is a direct target of c-Myc but is not functionally equivalent in cellular transformation. Oncogene 1999; 18:1219–26.
Littlewood TD, Hancock DC, Danielian PS, Parker MG, Evan GI . A modified oestrogen receptor ligand-binding domain as an improved switch for the regulation of heterologous proteins. Nucleic Acids Research 1995; 23:1686–90.
Pari GS, Keown WA . Experimental strategies in efficient transfection of mammalian cells: calcium phosphate and DEAE-Dextran. In: Tuan RS eds Recombinant gene expression protocols, Methods in Molecular Biology. Human Press, New Jersey: 1997; 62:301–6.
Freshney RI . Culture of animal cells, a manual of basic technique (second ed.), ARL Inc., New York, 1987:144–6.
Chomczynski P, Sacchi N . Single-step method of RNA isolation by acid guanidinium thiocyanate phenol chloroform extraction. Anal Biochem. 1987; 162:156–9.
Walker AK, Enrietto PJ . Regulatable chimeric oncogene, Oncogene Techniques, Methods in Enzymology, Academic Press 1995; 254:469–80.
Packham G, Cleveland JL . c-Myc and apoptosis, Biochimica et Biophysica Acta 1995; 1442:11–28.
Desbarats L, Schneider A, Muller D, Burgin A, Eilers M . Myc: a single gene controls both proliferation and apoptosis in mammalian cells. Experientia 1996; 52:1123–9.
Grandori C, Eisenman RN . Myc target genes. TiBS 1997; 22:177–81.
Acknowledgements
The authors thank Dr. Evan, Dr. Eisenman, Dr. Farnnam, Dr. Tsuneoka and Dr. Dang for their kind gifts of cDNAs of c-mycER, D106-143c-mycER, ODC, MrDb, cad, Rcc1 and rcl. This work was supported by research grants from National Natural Science Foundation of China (C Shi), Laboratory of Molecular Cell Biology, SICB, CAS (C-M Fang), Human Genome Program of CAS (Y-H Xu) and 863 High-tech Program (Y-H Xu).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
FANG, C., SHI, C. & XU, Y. Deregulated c-myc expression in quiescent CHO cells induces target gene transcription and subsequent apoptotic phenotype. Cell Res 9, 305–314 (1999). https://doi.org/10.1038/sj.cr.7290029
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7290029
Keywords
This article is cited by
-
Geochemistry of the Lower Cretaceous limestones in the Eastern Tethys Gyabula Formation (Himalaya, southern Tibet): implications for the depositional environment and tectonic setting
Arabian Journal of Geosciences (2021)
-
Effect of heparin on apoptosis in human nasopharyngeal carcinoma CNE2 cells
Cell Research (2001)
-
Improved safety through tamoxifen-regulated induction of cytotoxic genes delivered by Ad vectors for cancer gene therapy
Gene Therapy (2000)