Abstract
Colon cancers arising in most patients with hereditary nonpolyposis colorectal cancer (HNPCC) show microsatellite instability (MSI). BAT26, a quasimonomorphic polyA stretch located just 3′ of MSH2 exon 5, is considered the most sensitive and specific marker of MSI. A total of 10 HNPCC families with large intragenic MSH2 deletions, encompassing exon 5 and intron 5, identified by multiplex ligation-dependent probe amplification (MLPA) were included in this study. The deletions under study were del1-16, del1-8, del1-7, del1-6, and del3-6, detected in 3, 1, 2, 3, and 1 families, respectively. Although all patients examined from these 10 families developed unstable tumours, 13/19 MSI-H tumours (68 %) surprisingly showed stability of BAT26. By MLPA and MSH2 sequence analyses of the BAT26-stable tumours, we demonstrated that the wild-type MSH2 allele was somatically inactivated by an identical large deletion, with complete loss of intron 5/BAT26 sequences at the tumour DNA level. We could infer that the apparent stability of BAT26 was due to the complete absence of target BAT26 sequences in the tumour sample, which results in exclusive amplification of contaminant normal DNA, containing a single copy of a wild-type stable BAT26 sequence. Identification of a subset of MSH2-related unstable tumours that are not recognized by analysis of BAT26 instability indicates that this marker should never be used alone for rapid MSI screening of HNPCC tumours. Moreover, our findings indicate that BAT26 stability in the context of MSI is strongly suggestive of the presence of a large intragenic MSH2 deletion.
Similar content being viewed by others
Introduction
Hereditary nonpolyposis colorectal cancer (HNPCC), the most common form of hereditary colon cancer, is an autosomal dominant syndrome characterized by a high incidence of early onset colorectal cancer and an excess of cancers of extracolonic sites, such as endometrium, small intestine, urinary and hepatobiliary tracts, ovary, and stomach. 1, 2
HNPCC predisposition is caused by germline mutations in one of the known human mismatch repair (MMR) genes, namely MLH1, MSH2, MSH6, and PMS2 (InSiGHT Mutation Database: http://www.insight-group.org/). Their inactivation may result from point mutations, identifiable in approximately 50–80% of the HNPCC families, or from genomic rearrangements, which can be detected in up to 20% of cases.3, 4, 5, 6, 7, 8, 9 Large deletions are especially frequent in MSH2, and they account for up to one-third of MSH2 mutations in some populations.10
Colon cancer arising in patients with a defective MMR system usually shows microsatellite instability (MSI), determined by the occurrence of frameshift mutations in simple sequence repeats. This molecular hallmark is useful for selecting patients and families eligible for MMR mutation analysis. To this aim, a standard panel of microsatellite markers for MSI testing of tumour tissues was proposed at the 1997 NCI international consensus meeting.11 This panel is comprised of five markers, including two mono- (BAT26 and BAT25) and three dinucleotide repeats (D2S123, D5S346 and D17S250). Conventionally, all samples showing altered length in ⩾2/5 markers are classified as MSI-H (high degree of instability), whereas samples with 1/5 (or less than 30%, if additional markers are tested) unstable markers and no altered markers are defined as MSI-L (low level of instability) and MSS (microsatellite stable), respectively.
More recently, an alternative panel of five mononucleotide repeats has been suggested.12, 13 This still includes BAT26 and BAT25, the most widely used markers, which are deemed to be the most sensitive and specific markers for the detection of MSI-H tumours.14, 15, 16, 17, 18
BAT26 is a quasimonomorphic marker formed by a poly-A tract located immediately downstream of MSH2 exon 5. For its high sensitivity, comprised between 93 and 100%, it has been proposed that it could be used as a single marker for MSI testing.14, 15, 18, 19, 20 Here we show that HNPCC tumours from a subset of patients who are heterozygous for MSH2 deletions are characterized by frequent stability of BAT26.
Materials and methods
Patients and tumours
A total of 10 HNPCC families were selected from a group of 16 families in which we previously identified large intragenic MSH2 deletions by multiplex ligation-dependent probe amplification (MLPA) (Table 1). Overall, 18 deletions carriers from these 10 families, including the probands, were characterized by the inclusion of exon and intron 5 in the constitutionally deleted tracts. Lack of immunohistochemical MSH2 expression in tumours from nine out of nine tested families was consistent with the underlying MSH2 genetic defect (data not shown). Some patients developed more than one tumour, and MSI status could be investigated on a total of 22 tumour DNA samples.
In the remaining six families with large MSH2 deletions, exon 5/intron 5 were retained in the mutant allele. A total of nine tumours from six patients belonging to this subset were available for MSI analysis.
The research was approved by the local Ethical Committee and written consent was obtained from patients.
MSI analysis
Genomic DNA was obtained from blood and from paraffin-embedded or frozen tissues. Tumour DNA was extracted by the DNAeasy tissue kit (Qiagen GmbH, Hilden, Germany) following the manufacturer's instructions.
Standard MSI analysis was performed on paired tumour–normal tissue DNA samples using the NCI panel of microsatellite markers (BAT26, BAT25, D2S123, D5S346, D17S250).11 Where possible, additional markers were investigated (NR21, NR22, and NR24).12 The fluorescently labelled PCR products were mixed with a size standard (GeneScan 400HD ROX, Applera, Foster City, CA, USA), run on an automatic ABI3100 DNA analyzer, and evaluated with the GeneScan software (Applera). A variation in number and size of peaks of a marker in tumour DNA compared with normal DNA was interpreted as instability for that marker. Tumours were scored as MSI-H, MSI-L, or MSS, according to the international criteria.11
MLPA analysis
MLPA was performed with 200 ng of normal and tumour DNAs using the MRC-Holland (Amsterdam, Holland) HNPCC probe kit, according to the supplier's protocol. Then, 1 μl of the FAM-labelled PCR product was mixed with 1 μl of fluorescent GeneScan 500 LIZ size standard (Applera) in 15 μl of HiDi Formamide, run on an automatic ABI3100 DNA analyser, and evaluated with GeneScan software (Applera). The electropherograms showed specific peaks corresponding to each exon of MSH2 and MLH1, as well as additional peaks corresponding to control sequences mapping on different chromosomes. Dosage analyses based on comparison between deleted and reference wild-type DNA samples were performed on an Excel file following the manufacturer's protocol (www.mrc-holland.com). Briefly, all MSH2 peak areas were normalized by dividing each peak area by the combined area of all other peaks of the electropherogram. Then, these normalized peak areas were divided by the corresponding wild-type normalized peak area (average of three independent DNA samples) of that probe amplification product, in order to obtain a series of 16 aex values.
In constitutional and tumour DNA samples, a 40–55% decrease of the area of an MSH2 exon peak compared to the wild-type control samples was considered as indicative of a heterozygous deletion of that exon. In tumor DNAs, a 60–90% decrease was considered as indicative of a homozygous deletion of that exon.
To quantitatively evaluate deletions in tumour DNAs, the following values were also calculated for each electropherogram of all tumour/normal DNA pairs: Adel, the mean value of normalized deleted MSH2 peak areas; and Aret, the mean value of normalized retained MSH2 peak areas, where present.
Polymorphism analysis
Four sequence polymorphisms of the MSH2 locus were analysed: IVS1(+9)C/G, IVS9(−9)A/T, IVS10(+12)G/A, and IVS12(−6)A/G. Analysis was first performed by direct sequencing of PCR products on normal DNA. In heterozygous patients, the same polymorphism was evaluated on tumour DNA to check for the presence of loss of heterozygosity (LOH).
Results
MSI status was investigated on a total of 22 tumour DNA samples, including 15 colorectal carcinomas, two colorectal adenomas, two kidney tumours, and one tumour each of the stomach, bladder, and skin, from the 18 selected individuals with deleted exon 5/intron 5, and a total of nine tumours, including five colorectal carcinomas, two colorectal adenomas, and one tumour each of the uterus and skin, from the six patients with retained exon 5/intron 5. With the exception of the kidney tumour of patient CFS185 (family A-AV62), of the bladder and skin tumours of patient CFS366 (family A-VA25) and of the adenoma and skin tumour of patient CFS316 (family A-VA16), which were MSS (data not shown), all other tumours were clearly MSI-H. In particular, they all displayed instability at the BAT25 marker. However, 13/19 tumours (68%) in the first group surprisingly showed stability of BAT26. This result was observed in tumours from nine different families, and was absent in four out of five tumours from family A-AV62 (del1-16), in one out of three tumours from family A-VA25 (del1-6), and in the only tumour of family A-PD21 (del1-6) (Table 1). BAT26 was unstable in the seven MSI-H tumours from patients carrying MSH2 deletions not including exon 5 (Table 1).
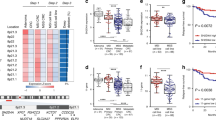
In order to gain some insight into the mechanisms underlying this unexpected phenomenon, we carried out additional molecular tests on tumour DNAs. As a first step, we used MLPA analysis to determine the copy number of each MSH2 exon in tumour DNA samples. Owing to the poor yield and/or quality of some samples, only 13 tumour samples gave reliable results and could be used for tumour/constitutional comparisons. Careful visual inspection of the tumour electropherograms from the 10 BAT26-stable samples showed a further reduction in size of the same peaks deleted in matched normal DNA, without significant changes in the peaks that were not constitutionally deleted (Figure 1a). On the other hand, the electropherograms of the three BAT26-unstable tumours were undistinguishable from those of the corresponding constitutional DNAs (Figure 1b).
Large deletions of MSH2 gene. Representative MLPA electropherograms of wild-type controls (wt) and constitutional (N) and tumour (T) DNAs from a BAT26-stable sample with del1-7 (a) and a BAT26-unstable sample with del1-6 (b). The MLPA control (C), MSH2-exon 5 (5), MSH2-deleted (↓), and MSH2-retained (*) peaks are indicated.
Precise qualitative and quantitative definition of these exonic deletions was carried out by dosage analysis of all single peaks. The exact extension of deletions in the different tumour/normal pairs was confirmed by calculating the aex values, which approximated ∼1 for the retained peaks, but were ∼0.5 or lower for the deleted peaks (Figure 2a and b). Moreover, to better define the borders of the somatically deleted regions, we verified that the flanking peak (ie the first retained peak immediately adjacent to the deleted region) had a normalized area of at least 85% of the expected value in each tumor DNA (data not shown).
Dosage of deleted and retained exons. Representative histograms showing aex values of constitutional (N) and tumour (T) DNAs from a BAT26-stable sample with del1-7 (a) and a BAT26-unstable sample with del1-6 (b). Panel (c) shows Adel values in BAT26 stable (from CFS261 to CFS408) and BAT26 unstable (from CFS356 to CFS262) cases.
Adel ranged from 0.45±0.08 to 0.58±0.09 in constitutional DNAs and from 0.18±0.04 to 0.58±0.13 in tumor DNAs (Figure 2c). However, in the BAT26-stable tumours, these values were in the range of 0.18±0.04–0.37±0.08 and, in each couple of tumor/normal DNA, the tumour Adel value was always significantly lower (P<0.001) than the matched constitutional Adel value. On the contrary, Adel of the BAT26-unstable tumours had values between 0.51±0.05 and 0.58±0.13, overlapping their corresponding constitutional Adel values (0.47±0.08 and 0.56±0.10).
Aret varied from 0.82±0.12 to 1.08±0.04 and from 0.80±0.04 to 1.20±0.04 in constitutional and tumour DNAs, respectively. The Aret values of the eight BAT26-stable tumours with retained exons were not dissimilar from the Aret value calculated for CFS356, the only BAT26-unstable sample retaining some MSH2 exons (data not shown).
Overall, these data indicate that tumour somatic inactivation of the wild-type MSH2 allele was determined by a second (intra)genic deletion, which was limited to the exons already deleted in constitutional DNA, in the BAT26-stable, but not in the BAT26-unstable tumour samples (Table 2).
Polymorphism analyses of some samples were not informative, due to constitutional homozygosity or to PCR difficulty in some paraffin-embedded samples. Only five tumours, four BAT26-stable and one unstable, could be evaluated for at least one of the four MSH2 polymorphisms analysed. Heterozygosity was retained in the BAT26-unstable, but not in the four BAT26-stable tumour DNAs, which displayed total or partial LOH of MSH2 sequence polymorphisms mapping outside the deleted region (Table 2).
Discussion
This is the first study investigating the correlations between BAT26 status and mutation type in MSH2. We observed an association between stability of the BAT26 microsatellite marker and presence of large intragenic MSH2 deletions encompassing exon and intron 5. Since BAT26 stability was present in 13 out of 19 MSI-H tumours with such deletions, it appears to be a very frequent (68%) and specific phenomenon. In fact, BAT26 was unstable in seven out of seven tumour samples from subjects who were heterozygous for MSH2 deletions not encompassing exon and intron 5, and in our 10 year-laboratory experience, BAT26 stability was very rarely observed in tumour DNAs from patients with MMR gene point mutations and never observed in patients with large intragenic MLH1 deletions. Only recently it has become evident that large rearrangements of MMR genes cause Lynch syndrome in about 20% of cases. MLPA, a new high-resolution, robust and quick assay for detecting copy number variations in genomic sequences, has provided an important contribution for the detection of several new MSH2 and MLH1 deletions in some cohorts of families.4, 5, 8
Owing to its high sensitivity and specificity, BAT26 was initially proposed as an unambiguous marker of MSI,14 and therefore it has been included in both microsatellite panels proposed at the first11 and second13 Bethesda consensus meetings. Recently, it has been reported that BAT26, together with BAT25, accurately detects MSI-H tumours and can also be used to predict tumour content (the percentage of tumour cells) in DNA samples.18
BAT26 is a quasimonomorphic poly-A stretch located just 3′ of MSH2 exon 5. We hypothesized that BAT26 stability could be due to somatic deletions, with complete loss of intron 5/BAT26 sequences in tumour cells, and, consequently, amplification of a wild-type allele present in infiltrating leucocytes and/or stromal cells. By MLPA analysis, we demonstrated that the wild-type MSH2 allele in BAT26-stable tumours, but not in the BAT26-unstable samples, is somatically inactivated by another large deletion encompassing intron 5/BAT26 sequences, thus confirming our hypothesis. We found that MLPA is a very easy and efficient test to detect large MSH2 rearrangements also in the majority of DNA samples extracted from paraffin-embedded tumours. Visual and quantitative evaluation of the MLPA peaks areas showed that normal and tumour DNAs had similar patterns in the BAT26 unstable cases, with Adel values ≈0.5, as expected. On the other hand, the MLPA patterns of the BAT26-stable tumours were different from those of the corresponding normal tissues, showing on average Adel <0.5 and Aret ≈1. This suggests that somatic inactivation of the wild-type MSH2 allele was determined by a second identical deletion, which leads to homozygosity of the constitutional mutation. However, only DNA sequencing of breakpoints on both alleles could definitively confirm this hypothesis. Hemizygosity, as alternative to homozygosity, was excluded because the height of retained MSH2 peaks relative to the other internal peaks did not significantly vary between normal and tumour DNAs. However, reduction to homozygosity was never complete, showing smaller deleted peaks of variable height, likely due to the presence of contaminating normal DNA.
We then investigated the status of MSH2 intragenic polymorphic markers in tumour/constitutional DNA pairs. Data from these polymorphisms were concordant with those obtained with MLPA analysis, showing the presence of two different alleles in the only informative BAT26-unstable tumour DNA sample (CFS356), and LOH in all four BAT26-stable samples, possibly due to the presence of two identical MSH2 alleles. Two mechanisms can account for the reduction to homozygosity for MSH2 deletions observed in the majority of the samples with a stable BAT26 marker: loss of the wild-type chromosome with duplication of the whole chromosome 2 containing the mutant allele or, alternatively, a somatic recombination, possibly associated with DNA repair failure, with copy/duplication of a DNA tract restricted to a region of the short arm surrounding the MSH2 locus. Further analyses will be necessary to discriminate between these two hypotheses.
Our findings indicate that BAT26 stability in a context of MSI-H is strongly suggestive of the presence of a large intragenic MSH2 deletion, especially when supported by absence of MSH2 immunoistochemical expression and positive family history. In fact, 68% of MSI-H tumours developed by patients with different MSH2 deletions encompassing exon/intron 5 were BAT26 stable, because of a second hit leading the constitutional mutation to homozygosity. Only two deletions escaped this rule: del1-16 (BAT26 instability in 4/7 MSI-H tumours) and del1-6 (two of the five samples examined were unstable). Although literature data on the types of somatic mutations in MSH2-related tumours are scanty, point mutations have also been reported in addition to allelic loss/LOH mutations.21, 22, 23, 24 Our data demonstrate that in a small subgroup of patients with specific MSH2 constitutional mutations, LOH associated with homozygosity of the mutant allele is the preferred mechanism of somatic inactivation, and suggest that the first MSH2 hit can drive the molecular events associated with the second hit.
Finally, our findings bear important consequences for the clinical setting. Assessment of MSI status is used for the diagnosis of HNPCC and may also be relevant for disease prognosis25 and for predicting response to chemotherapy.26 As a consequence of an increasing demand for this molecular test, it is very important to arrange rapid and cost-effective tests with panels of microsatellite markers able to satisfy both clinical requirements. Identification of a subset of MSH2-related MSI-H tumours that are not recognized by BAT26 indicates that this marker should never be used alone for the evaluation of instability in a rapid MSI screening of HNPCC tumours.
References
Lynch HT, de la Chapelle A : Hereditary colorectal cancer. N Engl J Med 2003; 348: 919–932.
Lucci-Cordisco E, Zito I, Gensini F, Genuardi M : Hereditary nonpolyposis colorectal cancer and related conditions. Am J Med Genet 2003; 122A: 325–334.
Viel A, Petronzelli F, Della Puppa L et al: Different molecular mechanisms underlie genomic deletions in the MLH1 gene. Hum Mutat 2002; 20: 368–374.
Gille JJP, Hogervorst FBL, Pals G et al: Genomic deletions of MSH2 and MLH1 in colorectal cancer families detected by a novel mutation detection approach. Br J Cancer 2002; 87: 892–897.
Nakagawa H, Hampel H, de la Chapelle A : Identification and characterization of genomic rearrangements of MSH2 and MLH1 in Lynch syndrome (HNPCC) by novel techniques. Hum Mutat 2003; 22: 258.
Taylor CF, Charlton RS, Burn J, Sheridan E, Taylor GR : Genomic deletions in MSH2 or MLH1 are a frequent cause of hereditary non-polyposis colorectal cancer: identification of novel and recurrent deletions by MLPA. Hum Mutat 2003; 22: 428–433.
Wagner A, Barrows A, Wijnen JT et al: Molecular analysis of hereditary nonpolyposis colorectal cancer in the United States: high mutation detection rate among clinically selected families and characterization of an American founder genomic deletion of the MSH2 gene. Am J Hum Genet 2003; 72: 1088–1100.
Bunyan DJ, Eccles DM, Sillibourne J et al: Dosage analysis of cancer predisposition genes by multiplex ligation-dependent probe amplification. Br J Cancer 2004; 91: 1155–1159.
Di Fiore F, Charbonnier F, Martin C et al: Screening for genomic rearrangements of the MMR genes must be included in the routine diagnosis of HNPCC. J Med Genet 2004; 41: 18–20.
Wijnen J, van der Klift H, Vasen H et al: MSH2 genomic deletions are a frequent cause of HNPCC. Nat Genet 1998; 20: 326–328.
Boland CR, Thibodeau SN, Hamilton SR et al: A national cancer institute workshop on microsatellite instability for cancer detection and familial predisposition: development of international criteria for the determination of microsatellite instability in colorectal cancer. Cancer Res 1998; 58: 5248–5257.
Suraweera N, Duval A, Reperant M et al: Evaluation of tumor microsatellite instability using five quasimonomorphic mononucleotide repeats and pentaplex PCR. Gastroenterology 2002; 123: 1804–1811.
Umar A, Boland CR, Terdiman JP et al: Revised Bethesda Guidelines for Hereditary Nonpolyposis Colorectal Cancer (Lynch Syndrome) and Microsatellite Instability. J Natl Cancer Inst 2004; 96: 261–268.
Hoang JM, Cottu PH, Thuille B, Salmon RJ, Thomas G, Hamelin R : BAT-26, an indicator of the replication error phenotype in colorectal cancers and cell lines. Cancer Res 1997; 57: 300–303.
Dietmaier W, Wallinger S, Bocker T, Kullmann F, Fishel R, Rüschoff J : Diagnostic microsatellite instability: definition and correlation with mismatch repair protein expression. Cancer Res 1997; 57: 4749–4756.
Bacher JW, Flanagan LA, Smalley RL et al: Development of a fluorescent multiplex assay for detection of MSI-high tumors. Dis Markers 2004; 20: 237–250.
Buhard O, Suraweera N, Lectard A, Duval A, Hamelin R : Quasimonomorphic mononucleotide repeats for high-level microsatellite instability analysis. Dis markers 2004; 20: 251–257.
Brennetot C, Buhard O, Jourdan F, Flejou JF, Duval A, Hamelin R : Mononucleotide repeats BAT-26 and BAT-25 accurately detect MSI-H tumours and predict tumor content: implications for population screening. Int J Cancer 2005; 113: 446–450.
de la Chapelle A : Testing tumors for microsatellite instability. Eur J Hum Genet 1999; 7: 407–408.
Laghi L, Bianchi P, Roncalli M, Malesci A : Re: Revised Bethesda Guidelines for Hereditary Nonpolyposis Colorectal Cancer (Lynch Syndrome) and Microsatellite Instability. J Natl Cancer Inst 2004; 96: 1402–1403.
Wheeler JMD, Beck NE, Kim HC, Tomlinson IPM, McC Mortensen NJ, Bodmer WF : Mechanisms of inactivation of mismatch repair genes in human colorectal cancer cell lines: the predominant role of hMLH1. Proc Natl Acad Sci USA 1999; 96: 10296–10301.
Kruse R, Rutten A, Hosseiny-Malayeri HR et al: ‘Second hit’ in sebaceous tumors from Muir-Torre patients with germline mutations in MSH2: allele loss is not the preferred mode of inactivation. J Invest Dermatol 2001; 116: 463–465.
Yuen ST, Chan TL, Ho JWC et al: Germline, somatic and epigenetic events underlying mismatch repair deficiency in colorectal and HNPCC-related cancers. Oncogene 2002; 21: 7585–7592.
Miyaki M, Iijima T, Yamaguchi T et al: Novel germline hMSH2 genomic deletion and somatic hMSH2 mutations in a hereditary nonplyposis colorectal cancer family. Mutat Res 2004; 548: 19–25.
Gryfe R, Kim H, Hsieh ET et al: Tumor microsatellite instability and clinical outcome in young patients with colorectal cancer. N Engl J Med 2000; 342: 69–77.
Ribic CM, Sargent DJ, Moore MJ et al: Tumor microsatellite-instability status as a predictor of benefit from fluorouracil-based adjuvant chemotherapy for colon cancer. N Engl J Med 2003; 349: 247–257.
Acknowledgements
The study was supported by Alleanza Contro il Cancro (to AV) and by a grant from the Italian Ministry for Education, University and Research (MIUR to MG).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pastrello, C., Baglioni, S., Tibiletti, M. et al. Stability of BAT26 in tumours of hereditary nonpolyposis colorectal cancer patients with MSH2 intragenic deletion. Eur J Hum Genet 14, 63–68 (2006). https://doi.org/10.1038/sj.ejhg.5201517
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201517
Keywords
This article is cited by
-
Left-sided colorectal cancer distinct in indigenous African patients compared to other ethnic groups in South Africa
BMC Cancer (2022)
-
Two-stain immunohistochemical screening for Lynch syndrome in colorectal cancer may fail to detect mismatch repair deficiency
Modern Pathology (2018)
-
MLPAstats: An R GUI package for the integrated analysis of copy number alterations using MLPA data
BMC Bioinformatics (2011)
-
Type A microsatellite instability in pediatric gliomas as an indicator of Turcot syndrome
European Journal of Human Genetics (2009)
-
Probe-specific mixed-model approach to detect copy number differences using multiplex ligation-dependent probe amplification (MLPA)
BMC Bioinformatics (2008)