Abstract
As a component of the apoptosome, a caspase-activating complex, Apaf-1 plays a central role in the mitochondrial caspase activation pathway of apoptosis. We report here the identification of a novel Apaf-1 interacting protein, hepatocellular carcinoma antigen 66 (HCA66) that is able to modulate selectively Apaf-1-dependent apoptosis through its direct association with the CED4 domain of Apaf-1. Expression of HCA66 was able to potentiate Apaf-1, but not receptor-mediated apoptosis, by increasing downstream caspase activity following cytochrome c release from the mitochondria. Conversely, cells depleted of HCA66 were severely impaired for apoptosome-dependent apoptosis. Interestingly, expression of the Apaf-1-interacting domain of HCA66 had the opposite effect of the full-length protein, interfering with the Apaf-1 apoptotic pathway. Using a cell-free system, we showed that reduction of HCA66 expression was associated with a diminished amount of caspase-9 in the apoptosome, resulting in a lower ability of the apoptosome to activate caspase-3. HCA66 maps to chromosome 17q11.2 and is among the genes heterozygously deleted in neurofibromatosis type 1 (NF1) microdeletion syndrome patients. These patients often have a distinct phenotype compared to other NF1 patients, including a more severe tumour burden. Our results suggest that reduced expression of HCA66, owing to haploinsufficiency of HCA66 gene, could render NF1 microdeleted patients-derived cells less susceptible to apoptosis.
Similar content being viewed by others
Main
Apoptosis is an evolutionarily conserved cell suicide process that depends on the activation of caspases, a group of cysteine aspartic acid-specific proteases.1 Two main activation pathways (extrinsic and intrinsic) of the initiator procaspases have been identified, involving engagement of death receptors by their ligands or perturbation of mitochondria.2 The intrinsic pathway involves the formation of a large caspase-processing complex termed the apoptosome. The apoptosome is formed by Apaf-1 upon binding to cytochrome c, which is released from the mitochondria by various forms of apoptosis triggers.3 The Apaf-1-cytochrome c complex then recruits the initiator caspase procaspase-9 in a dATP/ATP-dependent manner, resulting in its activation.4, 5
The intrinsic cell death pathway recognizes developmental cues, genomic stress or cytotoxic damage. Indeed, Apaf-1 knockout mice exhibit severe developmental defects that lead to embryonic lethality, and their cells are resistant to death receptor-independent induction of apoptosis.6, 7, 8 In-vivo, loss of expression of Apaf-1 is associated with tumour progression, suggesting that Apaf-1 inactivation may provide a selective survival advantage to neoplastic cells.9 Even though some apoptosis regulated by Bcl-2 can be elicited independently of the apoptosome,10 the above information clearly illustrate the critical role of Apaf-1 in mitochondria-dependent cell death.
Although it is clear that the core of the apoptosome complexes comprises Apaf-1 and caspase-9, other cellular proteins may modulate the formation or the activity of the apoptosome. A number of proteins have been reported to regulate apoptosis through interaction with Apaf-1, including Aven, NAC, Hsp70, Hsp90, Tucan, Nucling or APIP (see Schafer and Kornbluth11 for review). Because of the crucial role on Apaf-1 in the mitochondrial pathway, identifying new regulators of Apaf-1-mediated cell death will be important.
Here, we report the identification of a novel regulator of Apaf-1, HAC66. Hepatocellular carcinoma antigen 66 (HCA66) was able to potentiate drastically Apaf-1-dependent, but not Apaf-1-independent apoptosis whereas an isoform of HCA66 representing the binding domain of HCA66 to Apaf-1 had the opposite effect. HCA66 maps to chromosome 17q11.2 and is among the genes heterozygously deleted in neurofibromatosis type 1 (NF1) microdeletion syndrome patients.12, 13 Because these patients have a distinct phenotype compared to other NF1 patients, including a more severe tumour burden,14, 15, 16, 17, 18 we examined the possibility that reduced expression of HCA66, owing to haploinsufficiency of HCA66, could render NF1 microdeleted patients-derived cell lines less susceptible to apoptosis.
Results
Identification of HCA66
To investigate further the regulation of the apoptosome at the level of Apaf-1, we developed a yeast two-hybrid strategy using the WD40-deleted Apaf-1 (residues 1–591) as bait. Thirty-four interacting cDNA clones were identified from 68 million yeast diploïde using a CEMC7 random primed library. Among them, HCA66, a putative hepatocellular carcinoma tumour antigen19 (GenBankTM accession number NM_018428) was identified by four clones with two independent fragments and was selected for further study.
HCA66 is a protein of 597 residues with unknown function that is highly conserved between human and mouse (Figure 1a). Sequence analysis shows that HCA66 contained putative HAT (Half-A-TPR) repeats at its N-terminus. During the cloning of HCA66, we isolated an alternatively spliced form of HCA66 in which exon 9 of HCA66 is spliced out. This clone contains an open reading frame that generates the first 207 residues of HCA66 followed by a non-specific portion of 54 residues and a stop codon, and was named HCA66-S.
Sequence, tissue distribution and subcellular localization of HCA66. (a) Amino-acid sequence of human HCA66 (hHCA66) was aligned to the sequence of mouse HCA66 (mHCA66). Black and grey shades indicate identical and similar residues, respectively. (b) Expression pattern of HCA66. Human tissue Northern blots (Clontech) containing 2 μg of polyadenylated RNA were probed with radioactive HCA66 cDNA, exposed to X-ray films and photographed. PBL, peripheral blood leucocytes
To determine the distribution of HCA66, various normal human tissues mRNA samples were subjected to Northern blot analysis. As shown in Figure 1b, HCA66 mRNA (∼2.4 kbp) is constitutively expressed in most adult tissues with high expression in heart and in liver.
HCA66 interacts with the CED-4 homologous domain of Apaf-1
To confirm the interaction we detected by yeast two-hybrid analysis, C-terminally Flag-tagged HCA66 was expressed in 293 T cells with T7-tagged Apaf-1 or T7-tagged Ipaf, a CED-4-related protein that we described as a caspase-1 activator.20 Western blot analysis of protein complexes immunoprecipitated with anti-Flag antibody revealed that T7-Apaf-1, but not T7-Ipaf, co-precipitated with HCA66-Flag (Figure 2a). The immunoprecipitation was also performed in reverse. As shown in Figure 2b, HCA66-Flag co-precipitated with T7-Apaf-1 but not T7-Ipaf. To confirm the HCA66-Apaf-1 interaction at the endogenous level, endogenous HCA66 was immunoprecipitated from HeLa cell extracts using an HCA66 polyclonal antibody (antibody HCA66b) or a polyclonal antibody to ASC, a protein involved in the pro-inflammatory response.21 The resulting immune complexes were analysed by SDS-PAGE/immunoblotting with an anti-Apaf-1 antibody. ASC failed to immunoprecipitated any detectable Apaf-1 (Figure 2c, lane 1) whereas immunoprecipitation with HCA66 pelleted Apaf-1 in the absence but not in the presence of competing immunogenic peptide (Figure 2c, lane 2 and 3). These findings indicate that endogenous HCA66 associates with endogenous Apaf-1 in-vivo. To map the regions of HCA66 and Apaf-1 that are required for the interaction between the two molecules, we expressed T7-tagged HCA66 together with several Flag-tagged deletions mutants of Apaf-1. As shown in Figure 2d, a deletion mutant containing only the CED-4 homologous domain of Apaf-1 (residues 87–420) was still able to co-precipitate with HCA66. Of note, HCA66 was unable to interact with other components of the apoptosome (cytochrome c and caspase-9) neither with any human caspase (not shown). To extend the characterization of the HCA66-Apaf-1 interaction, we expressed T7-tagged Apaf-1 (87–420) with several forms of truncated Flag-tagged HCA66. Western blot analysis of Apaf-1 (87–420) complexes with anti-T7 antibody revealed that the first 207 residues of HCA66, corresponding to HCA66-S, were sufficient for interaction with Apaf-1 (Figure 2e). These observations were confirmed with glutathione S-transferase (GST) pull-down assays (Figure 2f). Combined, these data indicate that the interaction between HCA66 and Apaf-1 occurs through the N-terminal half of HCA66 and the CED-4 domain of Apaf-1.
HCA66 associates, through its N-terminus, with the CED-4 domain of Apaf-1. (a) and (b) 293 T cells were co-transfected with Flag-HCA66 together with either empty vector or T7-tagged Apaf-1 or T7-tagged Ipaf. After 24 h transfection, cells were lysed and the lysates immunoprecipitated with either anti-Flag or anti-T7 antibodies, as indicated. The immunoprecipitates were immunoblotted with anti-T7 or anti-Flag antibodies, respectively. Expression of T7- or Flag-tagged constructs was determined by immunoblotting with anti-T7 or anti-Flag antibodies, respectively. (c) Endogenous HCA66 interacts with endogenous Apaf-1. Cytoplasmic extracts (500 μg of proteins in 0.3 ml) derived from HeLa cells were subject to immunoprecipitation with anti-ASC (lane 1) or anti-HCA66 (antibody HCA66b) in the absence (lane 2) or presence (lane 3) of 10 μg of competing immunogenic peptide. The resulting protein precipitates were analysed by immunoblotting with anti Apaf-1 antibody. (d) HCA66 interacts with the CED-4 domain of Apaf-1. 293 T cells were co-transfected with T7-HCA66 together with either empty vector or the indicated Apaf-1 Flag-tagged constructs. Cells were lysed and the lysates immunoprecipitated with anti-Flag antibody, 24 h after transfection. The immunoprecipitates were immunoblotted with anti-T7 antibody. The expression of different T7- or Flag-tagged chimeras was determined by immunoblotting with anti-Flag or anti-T7 antibodies, respectively. (e) The N-terminal part of HCA66 mediates its interaction with Apaf-1. 293 T cells were co-transfected with T7-Apaf-1 (87–420) together with either empty vector or the indicated HCA66 Flag-tagged constructs. Immunoprecipitation and Western blot analysis was performed as in (d). (f) In vitro interaction of the CED-4 domain of Apaf-1 with the N-terminus part of HCA66. 35S-labeled HCA66 full length, N-terminus (residues 1–207) or C-terminus (residues 208-End; lane 1, 10% input) was incubated with an equal amount of glutathione Sepharose beads bound to an equal amount of GST (lane 2) or Apaf-1 (87–420)-GST (lane 3). Bound proteins were then eluted and analyzed by SDS-PAGE and autoradiography
A role for HCA66 in Apaf-1-mediated induction of apoptosis
To assess the functional significance of the interaction between Apaf-1 and HCA66, plasmids encoding HCA66 or HCA66-S were transiently transfected in HeLa cells. Whereas HCA66-S did not induce apoptosis, the full-length protein caused about 20% cell death in the HeLa-transfected cells (Figure 3a). Moreover, HCA66 was able to increase dramatically, in a dose-dependent manner, etoposide-, cisplatin- or staurosporine-induced apoptosis, stimuli that activate the Apaf-1-dependent mitochondrial pathway.22 In co-expression experiments, HCA66 was also able to potentiate apoptosis induced by Bax whereas a C287A dominant negative pro-caspase-9 mutant efficiently blocked HCA66-induced apoptosis (Figure 3a). Whereas full-length HCA66 induced apoptosis and enhanced Apaf-1-dependent cell death, HCA66-S had the opposite effect, interfering with apoptosis induced by etoposide, cisplatin, staurosporine or co-expression with Bax (Figure 3a). To determine whether HCA66 selectively modulates the Apaf-1 pathway, we transfected MCF7/Fas/casp3 cells with vectors expressing HCA66 or HCA66-S and then treated the cells with etoposide or agonist Fas antibody. These cells are known to behave like type I cells, that is active caspase-8, following death receptor stimulation, directly activates caspase-3 without relying on mitochondrial signal amplification.23 As shown in Figure 3b, HCA66 was able to potentiate staurosporin-induced apoptosis, whereas HCA66-S significantly inhibited apoptosis induced by the cytotoxic drug. However, the agonist Fas antibody was able to induce comparable levels of apoptosis in the presence or in the absence of HCA66 or HCA66-S (Figure 3b). Finally, as expected, neither HCA66 (1–190) nor HCA66 (211-End), forms that are unable to interact with Apaf-1 (Figure 2e), had an effect on drug-mediated apoptosis (Figures 3c). To examine further the function of HCA66 in apoptosis, we generated 293 T cell lines that stably expressed either HCA66 or HCA66-S. We then compared staurosporine-induced activation of downstream caspases (caspase-3 and -7) of the different cell lines. As shown in Figure 3d, overexpression of HCA66 significantly increased downstream caspases activation following treatment with staurosporine compared with the activation in control cells. Conversely, staurosporine-induced caspase activation was markedly reduced in cells expressing HCA66-S (Figure 3d). Collectively, these data demonstrate that the Apaf-1-interacting domain of HCA66 is a dominant negative regulator of Apaf-1-dependent apoptosis and that HCA66 might positively potentiate Apaf-1 function.
HCA66 selectively modulates Apaf-1 function. (a) HeLa cells were cotransfected with either HCA66 or HCA66-S together with empty vector or the indicated constructs in combination with a GFP-reporter plasmid. The cells were exposed to etoposide (ETP, 20 μM), staurosporine (STS, 0.5 μM) or cisplatin (CPT, 20 μM) for 16 h or left untreated, after 24 h transfection. Cells were then stained with Hoechst 33342 and normal and apoptotic GFP-expressing cells were counted using fluorescence microscopy. The percentage of apoptotic cells in each experiment was expressed as the percentage of apoptotic cells as a fraction of the total number of GFP-expressing cells. Each bar represents the mean±S.D. from three independent experiments. (b) MCF7/Fas/casp3 cells were transfected with either HCA66 or HCA66-S together with a GFP-reporter plasmid. The cells were exposed to staurosporine (STS, 0.5 μM) for 16 h or treated with anti-Fas antibody (0.5 μg/ml) plus cycloheximide for 4 h or left untreated, after 24 h transfection and percentages of apoptotic cells were determined as in (a). Each bar represents the mean±S.D. from three independent experiments. (c) HeLa cells were co-transfected with either HCA66 or the indicated HCA66 truncated mutants together with a GFP-reporter plasmid. The cells were exposed to etoposide (ETP, 20 μM) or cisplatin (CPT, 20 μM) for 16 h or left untreated, after 24 h transfection and percentages of apoptotic cells were determined as in (a). Each bar represents the mean±S.D. from three independent experiments. (d) HCA66 and HCA66-S have opposite effects on staurosporine-induced caspase-3/7 activation. 293 T clones stably transfected with HCA66 or HCA66-S constructs or empty vector were treated with different amounts of staurosporine for 5 h. Caspase-3/7 activity was measured using the Apo-ONE Homogeneous Caspase-3/7 Assay kit (Promega) according to the manufacturer's protocol. Assays were measured by detection with a fluorescence microplate reader and the fluorescence was measured at an excitation/emission wavelength of 485/535 nm. Results are presented as the mean values±S.D. of two independent experiments carried out in triplicate. (e) Wild type or Apaf−/− MEFs were transfected with either HCA66 or HCA66-S in combination with a GFP-reporter plasmid. The cells were exposed to etoposide (20 μM) for 16 h or left untreated, after 24 h transfection. Percentages of apoptotic cells were determined as in (a). Each bar represents the mean±S.D. from three independent experiments
To assess further the function of HCA66 in Apaf-1-mediated cell death, we transfected Apaf+/+ or Apaf−/− mouse embryonic fibroblast (MEFs) with HCA66 full length or HCA66-S encoding vectors and then treated the cells with etoposide. As shown in Figure 3e, expression of HCA66 induced apoptosis in Apaf+/+ but not in Apaf−/− MEFs, indicating that Apaf-1 is crucial for HCA66-induced cell death. Moreover, HCA66 synergized etoposide-induced apoptosis in Apaf+/+ cells, but etoposide treatment of Apaf−/− cells did not induce apoptosis, even in the presence of HCA66 (Figure 3e).
We then further assessed the function of HCA66 in Apaf-1-mediated apoptosis using RNA interference experiments. Small interfering RNA (siRNA) directed against HCA66, but not mock siRNA, reduced completely the level of HCA66 after 36 h whereas the levels of Apaf-1, pro-caspase-9 or cytochrome c did not change (Figure 4a and not shown). We next treated the transfected cells with etoposide and evaluated cell death. As shown in Figure 4b, siRNA directed against HCA66, but not mock siRNA, significantly prevented etoposide-induced apoptosis. However, silencing of HCA66 did not block cytochrome c release from mitochondria into cytosol following etoposide treatment (Figure 4c). To verify whether the observed reduced Apaf-1-dependent apoptosis was due specifically to the loss of HCA66, we performed rescue experiments by ectopically expressing a form of HCA66 that cannot be degraded by the siRNA. For this purpose, we introduced two silent mutations into the HCA66 cDNA in the region that is targeted by the siRNA. As shown in Figure 4a, siRNA transfection abolished completely the expression of ectopic wild-type HCA66, whereas the nondegradable HCA66 was readily detectable. Moreover, transfection of the mutated HCA66 together with HCA66 siRNA restored etoposide-induced apoptosis (Figure 4b), indicating that the effect of the siRNA could be explained by specific silencing of HCA66. Finally, we compared staurosporine-induced caspase-3/7 activity in 293 T cells transfected with control or HCA66-specific siRNA. As shown in Figure 4d, depletion of HCA66 resulted in a large reduction of effector caspases activity following staurosporine treatment compared with control cells. Thus, taken together, the above data strongly suggest a role for HCA66 in the apoptotic process and strengthen the notion that HCA66 specifically modulates the Apaf-1-mediated pathway at the level of, but not above, Apaf-1.
Deregulation of HCA66 protects from apoptosis induced by etoposide. (a) HeLa cells were transfected with Flag-tagged HCA66 or HCA66 in which two silent mutations preventing interaction between siRNA and the HCA66 mRNA were introduced (mutHCA66) in the absence or together with siRNA to HCA66. Cells were lysed and the lysates immunoblotted with anti-Flag or anti-tubulin antibodies to estimate the amounts of HCA66 or mutHCA66, after 36 h transfection. (b) Silencing of the HCA66 gene significantly prevents etoposide-induced apoptosis. HeLa cells were co-transfected with siRNA to HCA66 or mock (scrambled sequence) siRNA together with empty vector, Flag-tagged HCA66 or Flag-tagged mutHCA66, in combination with a GFP-reporter plasmid. The cells were exposed to etoposide (20 μM) for 16 h or left untreated, after 36 h transfection. Cells were then stained with Hoechst 33342 and percentages of apoptotic cells were determined as in Figure 3 (a) and (b). Each bar represents the mean±S.D. from three independent experiments. (c) Silencing of the HCA66 gene did not alter etoposide-induced cytochrome c release from mitochondria into the cytosol. HeLa cells were transfected with siRNA to HCA66 or mock siRNA.The cells were exposed to etoposide (20 μM) or left untreated, after 36 h transfection. Cell extracts were separated into membrane and cytosolic fractions, and the fractionated extracts were subsequently immunobloted with the indicated antibodies. (d) Reduction of staurosporine-induced caspase-3/7 activation following deregulation of HCA66. 293 T cells were transfected with siRNA to HCA66 or mock siRNA. The cells were exposed to various amounts of staurosporine for 5 h, after 36 h transfection,. Caspase-3/7 activity was measured as in Figure 3 (c). Results are presented as the mean values±S.D. of two independent experiments carried out in triplicate
Activation of caspases by cytochrome c is a multistep process, where Apaf-1 first oligomerizes, in the presence of cytochrome c and dATP, to form a functional apoptosome and then recruits pro-caspase-9 to the complex. We therefore examined which events could be influenced by HCA66. To assess the effects of HCA66 on oligomerization of Apaf-1 in vivo, T7-tagged Apaf-1 expressing HeLa cells were transfected with empty vector or Flag-Apaf-1 in the absence or together with siRNA directed to HCA66 or mock siRNA. Cell lysates were incubated with or without cytochrome c/dATP and immunoprecipitated with an anti-T7 antibody. Immunoblot analysis of the immunoprecipitates with an anti-Flag antibody indicated that, as expected, only in the presence of cytochrome c and dATP an association between T7- and the Flag-tagged Apaf-1 is detected (Figure 5a). This association still occurred after silencing of the HCA66 gene, indicating that HCA66 might not affect the oligomerization of Apaf-1 in response to cytochrome c/dATP. We next sought to determine if HCA66 could influence the recruitment of pro-caspase-9 to the apoptosome. Lysates from HeLa cells depleted or not of HCA66 by siRNA treatment were incubated with or without cytochrome c/dATP, in the presence of the pan-caspase inhibitor z-VAD-fmk and the endogenous Apaf-1 was immunoprecipitated with an Apaf-1 antibody. The immunoprecipitates were then analysed with a caspase-9 or Apaf-1 antibody. Here again, an association between Apaf-1 and caspase-9 was detected only in the presence of cytochrome c and dATP (Figure 5b). However, depletion of HCA66 significantly inhibited the amount of caspase-9 interacting with Apaf-1 (Figure 5b). Taken together, these data support a model in which HCA66 may regulate Apaf-1-dependent apoptosis by potentiating caspase-9 recruitment by Apaf-1.
Reduced amount of caspase-9 in the apoptosome following deregulation of HCA66. (a) T7-tagged Apaf-1 expressing HeLa cells were transfected with empty vector or Flag-tagged Apaf-1 in the absence or together with siRNA to HCA66 or mock siRNA. Cytosolic lysates were incubated for 30 min at 30°C in the presence or absence of 1 μM cytochrome c and 0.5 mM dATP and immunoprecipitated with anti-T7 antibody, after 36 h transfection. The immunoprecipitates were immunoblotted with anti-Flag antibody. The expression of Flag-tagged Apaf-1 as well as T7-Apaf-1 was determined by immunoblotting with anti-Flag or anti-T7 antibodies, respectively. (b) HeLa cells were transfected with siRNA to HCA66 or mock siRNA. Cytosolic lysates were incubated with or without cytochrome c and dATP, in the presence of 100 μM of z-VAD-fmk, and immunoprecipitated with anti-Apaf-1 antibody, after 36 h transfection. The immunoprecipitates were immunoblotted with antibodies against caspase-9 and Apaf-1. The expression of Apaf-1 as well as caspase-9 was determined by immunoblotting with anti-Apaf-1 or anti-caspase-9 antibodies, respectively. (c) HeLa cells were transfected with siRNA to HCA66 or mock siRNA. After 36 h transfection, aliquots (0.5 ml) of S100 extracts (2.5 mg ml−1) were incubated with or without cytochrome c and dATP and fractionated on a Superdex 200 HR column. HCA66, Apaf-1 and caspase-9 were detected in the various fractions by immunoblotting. Pro, unprocessed pro-caspase-9. The elution profiles and sizes of selected standards are indicated by arrowheads on the top. Lower panel, fractions 8–13 from cytochrome c/dATP-activated extracts were run side by side on the same gel. Apaf-1 and caspase-9 were detected in the various fractions by immunoblotting. (d) The Apaf-1-containing fractions from cytochrome c/dATP-activated extracts were analysed for their caspase-3 processing activity as described in ‘Material and Methods'. The produced DEVD-AMC cleaving activity was measured and expressed as arbitrary units (normalized fluorescence at 465 nm)
To confirm the immunoprecipitation results, we analysed apoptosome formation by size exclusion chromatography of S100 from HeLa depleted or not of HCA66. In the absence of cytochrome c and dATP, Apaf-1 was mostly in a monomeric form in both extracts (Figure 5c). Of note, HCA66 was detected along with Apaf-1 as well as in lower molecular weight fractions, indicating that HCA66 could associate with monomeric Apaf-1. After incubating with cytochrome c and dATP, most of the Apaf-1 shifted to a size of ∼650 000–1 000 000, indicating apoptosome formation. This shift was similar in the presence or in the absence of HCA66 (Figure 5c), confirming that the apoptosome was still efficiently formed even after depletion of HCA66. However, only a reduced amount of caspase-9 was associated with the apoptosome formed in the absence of HCA66 (Figure 5c). By running the fractions 8–13 of both cell extracts side by side in the same gel, we were able to estimate that there was a roughly sixfold decrease in the amount of associated caspase-9 in the absence of HCA66 (Figure 5c, lower panel). Interestingly, in mock siRNA-treated cells, a significant proportion of HCA66 now co-eluted with Apaf-1 in fractions 8–11, corresponding to the apoptosome. This indicates that HCA66 can associate with both monomeric and oligomerized Apaf-1. Finally, the ability of the apoptosome to activate exogenous caspase-3 was found to be dramatically reduced in the absence of HCA66 compared to that of control extracts (Figure 5d). Combined, these results indicate that HCA66 does not act by promoting Apaf-1 oligomerization but rather by increasing the amount of caspase-9 bound to the apoptosome following cytochrome c/dATP stimulation.
Haploinsufficiency of HCA66 may causes apoptosis defects in cell lines derived from patients with NF1 gene microdeletion
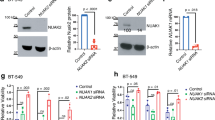
In 5–10% of the NF1 patients, a microdeletion is found that encompasses the NF1 gene and neighbouring genes, including HCA66.12, 13 Haploinsufficiency of the deleted genes results in the so called NF1 microdeletion syndrome, which is often characterized by a severe clinical phenotype for the microdeleted patients compared to the NF1 patients, with a more severe tumour burden, lower average IQ and facial dysmorphies.14, 15, 16, 17, 18 The outcome for microdeleted patients is probably caused by haploinsufficiency of one or more of the genes in the deletion interval, which spans at least 12 genes.24 Because of its role in apoptosis, we tested the possibility that lower expression of HCA66, caused by haploinsufficiency, might result in a lower Apaf-1-dependent apoptosis in NF1 microdeleted patients. For this purpose, Epstein-Barr virus (EBV) B lymphoblastoid cell lines were derived from healthy donor (Bwt), NF1 patients (B1, B2 and B3) and NF1 microdeleted patients (BM1, BM2 and BM3). We first sought to investigate the expression level of HCA66 in these cell lines using real-time quantitative RT-PCR. As shown in Figure 6a, expression of HCA66 was markedly reduced in the 3 cell lines derived from NF1 microdeleted patients compared to the cell lines from healthy donor or non-microdeleted NF1 patients. Conversely, HCA66 expression in NF1 non-microdeleted patients cell lines was at least equal or higher to that of control cells. Western blot analysis confirmed that protein levels of HCA66 were reduced in microdeleted patients cell lines compared to non-microdeleted NF1 patients or healthy donor cell lines (Figure 6b). However, Apaf-1, caspase-9 or caspase-3 protein levels were not markedly different among all the cell lines (Figure 6b).
Reduced HCA66 expression and Apaf-1-dependent apoptosis in NF1 microdeleted patients-derived cell lines. (a) Quantitative real-time PCR for HCA66 in control (Bwt), NF1 patients (B1, B2 and B3) and NF1 microdeleted patients (BM1, BM2 and BM3) derived lymphoblastoid cell lines. Quantification was performed by real-time PCR using cyclosporine A as the endogenous control. Values are means±S.E. of three experiments expressed in arbitrary units. (b) Expression of HCA66 and apoptosome-related molecules in used cell lines. Cytoplasmic cell extracts from the different cell lines were immunobloted with the indicated antibodies. (c) Reduced staurosporine-induced apoptosis in NF1 microdeleted patients-derived cell lines. The indicated cell lines were exposed to staurosporine (0.2 μM) for 16 h or left untreated and the percentage of cells that had become annexin V positive/PI negative was assessed by flow cytometry. Results are presented as the fold induction of cell death over untreated. Each bar represents the mean±S.D. from three independent experiments. (d) Reduced staurosporine-induced caspase-3/7 activation in NF1 microdeleted patients-derived cell lines. The indicated cell lines were exposed to staurosporine (0.2 μM) for 5 h or left untreated and caspase-3/7 activity was measured as in Figure 3(c). Results are expressed as fold increase in caspase-3/7 activation over untreated. Each bar represents the mean±S.D. from two independent experiments done in triplicate. (e) Staurosporine-induced reduction in mitochondrial transmembrane potential is not modified in NF1 microdeleted patients-derived cell lines. The indicated cell lines were exposed to staurosporine (0.2 μM) for 16 h or left untreated. Mitochondrial transmembrane potential loss (ΔΨm) was then determined by flow cytometry using DiOC6.3 Each bar represents the mean±S.D. from three independent experiments. (f) and (g) Deregulation of HCA66 protects NF1 patients-derived cell lines from apoptosis induced by staurosporine. (f) NF1 patients-derived cell lines were transfected with siRNA to HCA66 or mock siRNA using the Amaxa biosystem according to the manufacturer's protocol. Cytoplasmic extracts were immunobloted using anti-HCA66 (antibody HCA66a) or anti-tubulin, after 36 h transfection. (g) NF1 patients-derived cell lines were transfected with siRNA to HCA66 or mock siRNA. After 36 h transfection, the cells were exposed to staurosporine (0.2 mM) for 16 h or left untreated and apoptosis induction was estimated as in (c)
To determine if this reduction in HCA66 expression could affect the Apaf-1 pathway, we treated the different cell lines with staurosporine and evaluated cell death using annexin V/propidium iodide staining. Microdeleted patients cell lines clearly showed a lower rate of staurosporine-induced apoptosis compared with the healthy donor or non-microdeleted NF1 patients cells (Figure 6c). Moreover, induction of caspase-3/7 activity following staurosporine treatment was markedly reduced in NF1 microdeleted patients-derived cell lines compared to non-microdeleted NF1 patients or healthy donor cell lines (Figure 6d). Measure of 3,3′ dihexyloxacarbocyanine iodide (DiOC6)3 uptake indicated that staurosporine-induced reduction in mitochondrial transmembrane potential was comparable in all cell lines (Figure 6e), demonstrating that the observed differences in apoptosis did not result from variations in mitochondrial damage. Finally, silencing of HCA66, with specific siRNA, in NF1 patients-derived cell lines resulted in a clearly lower staurosporine-induced apoptosis (Figure 6f and g). Collectively, these results indicate that the level of HCA66 expression could be a crucial determinant of the sensitivity of the used cells to Apaf-1-mediated apoptosis.
Discussion
In this report, we describe the identification, through a yeast two-hybrid approach, of a novel interactor of Apaf-1, HCA66. HCA66 was found to positively regulate Apaf-1-dependent apoptosis whereas the N-terminus part of HCA66, which mediates its interaction with Apaf-1, had the opposite effect, interfering specifically with Apaf-1-dependent cell death. Silencing the HCA66 gene markedly inhibited Apaf-1-dependent apoptosis and reduced downstream effector caspases activation. Because the siRNA used did not downregulate expression of other apoptosome-related molecules nor prevent cytochrome c release following treatment with cytotoxic agent, our data indicate that HCA66 appears to regulate apoptosis at the level of Apaf-1-induced activation of caspase-9. Indeed, we found that, whereas the apoptosome was still efficiently formed, depletion of HCA66 resulted in a marked decrease of the amount of caspase-9 in the apoptosome following cytochrome c/dATP stimulation. This decrease was accompanied by a reduced ability for the apoptosome to activate the effector caspase-3. Therefore, our results indicate that HCA66 promotes apoptosis by increasing the level of active apoptosome. It possible that the binding of HCA66 can both induce a conformational change that would increase the recruitment of caspase-9, maybe through a more favourable orientation of the caspase recruitment domain of Apaf-1, and retain activated caspase-9 in the apoptosome, thus fine-tuning Apaf-1 function. Of note, the formation of the apoptosome in the absence of HCA66 is very similar to what was observed in the human ovarian carcinoma cell line SKOV3, which exhibit analogous diminished apoptosome activity.25 It is therefore possible that HCA66 could be missing in these cells, thus explaining the observed phenotype. Because HCA66 and HCA66-S bind to Apaf-1 in the same way, it is likely that HCA66-S, when overexpressed, acts as a dominant-negative inhibitor by competing with endogenous HCA66 and interfering with its binding to Apaf-1, resulting in a reduction of caspase-9 activation by the apoptosome. Although HCA66 binds to the CED-4 domain of Apaf-1, neither the long nor the short form (data not shown) modulates Apaf-1 oligomerization. At this moment, one can only speculate that HCA66-binding site on Apaf-1 is not in a domain essential for the formation of the Apaf-1 oligomer, like the binding pocket for dATP, which is crucial for apoptosome assembly. However, determination of the detailed structural organization of the HCA66-containing apoptosome as well as the precise mechanism of action of HCA66 awaits structural determination.
The gene HCA66 is located at 17q11.2, a region that is found to be deleted in 5–10% of NF1 patients. This heterozygous deletion, usually a 1.5 Mb region flanked by large paralogous sequences called NF1REPs,12, 26 involves 12 genes, including NF1 and HCA66. Microdeleted patients usually exhibit a more severe clinical phenotype, including lower average IQ, dysmorphic features and numerous and precocious neurofibromas.14, 15, 16, 17 These patients also appear to have a more severe malignant tumours burden, both NF1 associated and those not known to be associated with NF1.14, 16, 17, 18, 27, 28, 29, 30, 31, 32 It has long been suspected that deletion of NF1 gene and one of the adjacent genes favours the development of these tumours. We found that B lymphoblastoid cell lines from microdeleted patients exhibited a lower level of HCA66 expression compared to non-microdeleted NF1 patients or control cells. Interestingly, apoptosis response following staurosporine treatment as well as staurosporine-induced caspase-3/7 activity was found to be significantly reduced for microdeleted patients-derived cell lines. NF1 patients exhibit a mutation of the gene encoding neurofibromin. Loss of function of neurofibromin results in deregulation of Ras signaling owing to failure to inactivate Ras signal transduction.33 Ras controls Raf, MEK/ERK and PI-3K/Akt pathways, which regulate basic cell biologic functions such as cell proliferation, differentiation, survival and apoptosis. However, in our conditions, staurosporine-induced apoptosis from cells derived from non-microdeleted NF1 a patient was not reduced compared to that of the control cells, in contrast to microdeleted patients cell lines. Moreover, silencing of HCA66 in non-microdeleted NF1 patients cell lines with specific siRNA resulted in a significant reduction of staurosporine-induced apoptosis. Therefore, it is possible that insufficient expression of HCA66 protein could explain the observed default in apoptosis. Even although the precise role of HCA66 in the physiopathology of microdeleted patients remains inconclusive at the moment, one can suspect that the severe phenotype of microdeleted patients could be at least partially explained by a default in the apoptotic response, and in particular the Apaf-1-dependent response, owing to a reduction in HCA66 expression. However, in a syndrome as complex as NF1, all dosage-sensitive genes are likely to contribute to particular features of the phenotype.24 Therefore, one has to be careful in drawing potential phenotype-genotype correlation as for the impact of haploinsufficiency for HCA66 in microdeleted patients, and mouse models will have to be generated to address the role of HCA66 in the disease.
In summary, our results show that HCA66 acts as a positive regulator of Apaf-1-dependent apoptosis, and that reduced expression of this protein in NF1 microdeleted patients-derived cell lines might causes apoptosis defects. More and more data indicate that the apoptosome may be altered in various pathological conditions. Targeting HCA66 could be a potential mean for effective therapy for tumour and degenerative or developmental disorders.
Materials and Methods
Yeast two-hybrid screening procedure
Two-hybrid screens was performed using a cell to cell mating protocol as described previously34 using WD40-deleted Apaf-1 (residues 1–591) as bait and a random cDNA library from T-cell leukaemia cells CEMC7 poly(A+) RNA. Positive clones were obtained using the HIS3 reporter gene reporter gene selection. Inserts of the positive clones were amplified by PCR and sequenced on an ABI 3700 automatic sequencer (Applied Biosystem).
Northern blot analysis
Tissue distribution analysis of HCA66 mRNA was performed by Northern blot analysis on normal MTN Blot (Clontech). Each lane contains 2 μg of poly(A)+ RNA from specific tissues or cell lines. The blots were probed with a radiolabelled riboprobe prepared from a full-length human HC66 cDNA template and then subjected to autoradiography.
Materials and antibodies
Protease inhibitor cocktail, staurosporine, etoposide, cisplatin and DiOC63 were purchased from Sigma. Agonist Fas antibody was from Roche. Antibodies used for immunoprecipitation and Western blotting were rabbit anti-Apaf-1 (BD Pharmingen), rabbit anti-caspase-9 (Cell Signaling Technology), rabbit anti-caspase-3 (Cell Signaling Technology), mouse anti-cytochrome c (BD PharMingen), mouse anti-β-tubulin c (BD Pharmingen), rabbit anti-Cox4 (Abcam), rabbit anti-ASC (Abcam). Polyclonal antibodies to HCA66 were generated using keyhole limpet haemocyanin (KLH)-conjugated synthetic peptides with sequences corresponding to residues aa 477–491 (antibody HCA66a) or aa 35–53 (antibody HCA66b) of HCA66. Antibodies were purified by antigen-affinity chromatography using a NHS-activated Sepharose 4B column (Pharmacia).
Expression vectors
Constructs encoding full-length HCA66, Apaf-1, or truncated mutants were generated by PCR using modified complementary PCR adapter-primers. Flag- and T7-epitope tagging was carried out by cloning the PCR-generated cDNAs of the respective genes in-frame into pFLAG CMV-2 (IBI Kodak) and pcDNA3-T7 (Invitrogen) vectors, respectively. GST tagging was done by cloning the PCR-generated cDNAs together with GST cDNA in-frame into pET-28 vector (Novagen). To introduce silent mutations into the region of the HCA66 cDNA targeted by the siRNA used in this study, the codons at amino acid positions 142 and 143 were mutated from GCT to GCC (codon 142) and TTG to TTA (codon 143).
Cell culture
Cells were cultivated either in Dulbecco's modified Eagle medium (DMEM; HeLa, MCF-7/Fas/Casp3 or 293 T cells) or RPMI 1640 (B lymphoblastoid cells) (Gibco), supplemented with 10% fetal bovine serum, 200 μg ml−1 penicillin and 100 μg ml−1 streptomycin sulphate.
Transfection, immunoprecipitation and Immunoblot analysis and GST pull-down analysis
293 T cells (5 × 105 cells) in 60 mm dishes were transiently transfected with the expression plasmids using the JetPei™ (Polyplus Transfection) method as per the manufacturer's instructions. Cells were lysed, and the cellular lysates were immunoprecipitated and Western blotted as described before.20 For endogenous immunoprecipitation, cells were lysed by syringing ( × 20) in buffer A (20 mM HEPES-KOH, pH 7.5, 10 mM KCl, 1.5 mM MgCl2, 1 mM sodium EDTA, 1 mM sodium EGTA, 1 mM dithiothreitol and 0.1 mM PMSF) and centrifuged at 15 000 g for 15 min. NP-40 and KCl were then added to a final concentration of 0.5% and 100 mM, respectively and the protein complexes were immunoprecipitated from the lysate with an anti-HCA66 (antibody HCA66b) or the indicated antibodies immobilized on protein G-sepharose beads for 3 h at 4°C. The bead-bound proteins were then fractionated by SDS-PAGE and immunoblotted with an anti-Apaf-1 antibody. For GST pull-down assays, GST fusion proteins were overexpressed in Escherichia coli strain BL21 and equal amounts of proteins were immobilized on glutathione-Sepharose beads. Labelled interacting proteins were prepared by in vitro transcription and translation in the presence of 35[S]methionine using Promega's T7-coupled transcription/translation TNT kit according to the manufacturer's recommendations. After translation, equal amounts of the translation reactions were incubated with the protein-bound Sepharose beads in 100 μl of binding buffer (50 mM Tris-HCl, pH 7.6, 100 mm NaCl, 0.5% Brij, and protease inhibitors) for 3 h at 4°C. The beads were washed four times with the same buffer, boiled in SDS sample buffer. The proteins were then resolved on SDS-PAGE and visualized by autoradiography.
SiRNA transfection assay
The 21-nt-long interfering RNA duplexes with two 3′-end overhanging dT nucleotides on the antisense strand were synthesized by Eurogentech. The sequences of the siRNA were for HCA66: 5′-(CCAGCUUUGUGGAUUAUGG)dTT-3′ (nucleotides 421–439) and for control siRNA: 5′-(CGUUUAUUAUGAGCUGCGG)dTT-34. This control siRNA possess the same nucleotide composition as HCA66 siRNA but which lacks significant sequence homology to the genome. 293 T or HeLa cells were transfected with siRNAs or with siRNA together with the indicated plasmids using the JetSi endo transfection kit (Q-Biogene) as described by the manufacturer. B lymphoblastoid cells were transfected with siRNA with the Amaxa biosystem according to the B-cell transfection protocols provided by the manufacturer. HCA66 expression was analysed by Western blotting 36–48 h post transfection.
Apoptosis and in vitro caspase-3/7 assays
HeLa or MCF7 cell lines (1 × 105 cells/well) in 12-well plates were transfected with 0.1 μg of pEGFP-N1 (Clontech) reporter plasmid and the specified expression plasmids using the JetPei™ (Polyplus Transfection) method as per the manufacturer's instruction, and treated as indicated. Cells were then stained with Hoescht 33342 and normal and apoptotic GFP-expressing cells were counted using fluorescent microscopy. The percentage of apoptotic cells in each experiment was expressed as the mean percentage of apoptotic GFP-expressing cells as a fraction of the total number of GFP-expressing cells (n=3). FACS analysis of apoptosis and mitochondrial membrane potential modifications for HeLa cells transfected with siRNAs or B lymphoblastoid cell lines from NF1 patients were realized as described previously.35 Caspase-3/7 activities were measured using the Apo-ONE Homogeneous Caspase-3/7 Assay kit (Promega) according to the manufacturer's protocol. Assays were measured by detection with a fluorescence microplate reader (FLUOstar OPTIMA, BMG Labtechnologies), and the fluorescence was measured at an excitation/emission wavelength of 485/535 nm. Results are presented as the mean values±S.D. of two independent experiments carried out in triplicate.
Gel filtration analysis of apoptosome assembly
S100 extracts were prepared from HeLa cells transfected with mock or HCA66 siRNA in buffer A and incubated at 30°C for 30 min in the presence or absence of 1 μM cytochrome c and 0.5 mM dATP. After treatment, lysates were fractionated by gel-filtration chromatography using a Superdex-200 HR column (Amersham) pre-equilibrated with buffer A at a flow rate of 0.4 ml/min. The column was calibrated with a Sigma HMW gel-filtration protein standards kit. 0.4 ml fractions were collected. Aliquots of 40 μl from each fraction were then analysed for Apaf-1, caspase-9 and HCA66 by SDS-PAGE and immunoblotting. The biological activity of the apoptosome complexes was assayed as published previously.36, 37 Briefly, 100 μl of the Apaf-1-containing fractions were incubated with 100 nM of purified procaspase-3 for 30 min at 37°C. Ac-DEVD-AMC (100 μM) were then added and AMC release was measured, at emission wavelength of 465 nm with an excitation wavelength of 360 nm, at various times. The DEVD-AMC cleaving activity was expressed as normalized fluorescence produced after 3 h incubation at 37°C.
Genotyping of NF1 patients
A two-step approach was used to identify NF1 gene large deletions. In the first step, patients were genotyped using four intragenic microsatellite markers located in introns 1, 27 and 38. In the second step, patients hemi-homozygous for all markers were analysed with a real-time quantitative PCR-based gene dosage targeted on exons 6, 21, 28 and 48 of the NF1 gene with the ALB gene as endogenous control, as described previously38 (primers sequences available upon request). Results were reported to a normal DNA control. Samples with a value of 0.5 are considered to be deleted at least from exon 6 to 48.
Establishment of B-cell lines from PBMCs of NF1 patients
B lymphoblastoid cell lines were generated from peripheral blood mononuclear cells (PBMCs) by Epstein–Barr virus (EBV) immortalization. The cells were then passaged by partial medium change every 3–4 days.
Nucleic acid extraction and preparation of cDNA templates
RNA was isolated from lymphoblastoid cells using an RNeasy™ Mini Kit and an RNase-Free DNase Set (QIAGEN). Oligo(dT) primed cDNA was prepared from this RNA by reverse transcription using a SuperScript™ III First-Strand Synthesis System for RT-PCR kit (Invitrogen).
Real-time quantitative RT-PCR
The relative expression level of transcripts for HCA66 was quantified by real-time RT-PCR using the SYBR-Green PCR Master Mix on an ABI PRISM 7300 Sequence Detection System (Applied Biosystems, France), according to the manufacturer's instructions. A nontemplate control was run with each assay and the cyclophilin A housekeeping gene was used as an endogenous control (primers sequences available upon request). The relative amount of each mRNA was estimated using a standard curve constructed from serial dilutions of cDNA. The results are expressed in arbitrary units and are the average of three separate experiments (Mean±S.D., n=3).
Accession codes
Abbreviations
- DiOC6(3):
-
3,3′ dihexyloxacarbocyanine iodide
- GST:
-
glutathione S-transferase
- HCA:
-
hepatocellular carcinoma antigen
- MPNST:
-
malignant peripheral nerve sheath tumour
- NF1:
-
neurofibromatosis type 1
- PI:
-
propidium iodide
- Δψm:
-
transmembrane mitochondrial potential
References
Song Z, Steller H . Death by design: mechanism and control of apoptosis. Trends Cell Biol 1999; 9: M49–M52.
Nicholson DW . Caspase structure, proteolytic substrates, and function during apoptotic cell death. Cell Death Differ 1999; 6: 1028–1042.
Green DR, Reed JC . Mitochondria and apoptosis. Science 1998; 281: 1309–1312.
Hu Y, Benedict MA, Ding L, Nunez G . Role of cytochrome c and dATP/ATP hydrolysis in Apaf-1-mediated caspase-9 activation and apoptosis. EMBO J 1999; 18: 3586–3595.
Li P, Nijhawan D, Budihardjo I, Srinivasula SM, Ahmad M, Alnemri ES et al. Cytochrome c and dATP-dependent formation of Apaf-1/caspase-9 complex initiates an apoptotic protease cascade. Cell 1997; 91: 479–489.
Cecconi F, Alvarez-Bolado G, Meyer BI, Roth KA, Gruss P . Apaf1 (CED-4 homolog) regulates programmed cell death in mammalian development. Cell 1998; 94: 727–737.
Hakem R, Hakem A, Duncan GS, Henderson JT, Woo M, Soengas MS et al. Differential requirement for caspase 9 in apoptotic pathways in vivo. Cell 1998; 94: 339–352.
Yoshida H, Kong YY, Yoshida R, Elia AJ, Hakem A, Hakem R et al. Apaf1 is required for mitochondrial pathways of apoptosis and brain development. Cell 1998; 94: 739–750.
Igney FH, Krammer PH . Death and anti-death: tumour resistance to apoptosis. Nat Rev Cancer 2002; 2: 277–288.
Marsden VS, O'Connor L, O'Reilly LA, Silke J, Metcalf D, Ekert PG et al. Apoptosis initiated by Bcl-2-regulated caspase activation independently of the cytochrome c/Apaf-1/caspase-9 apoptosome. Nature 2002; 419: 634–637.
Schafer ZT, Kornbluth S . The apoptosome: physiological, developmental, and pathological modes of regulation. Dev Cell 2006; 10: 549–561.
Jenne DE, Tinschert S, Dorschner MO, Hameister H, Stephens K, Kehrer-Sawatzki H . Complete physical map and gene content of the human NF1 tumor suppressor region in human and mouse. Genes Chromosomes Cancer 2003; 37: 111–120.
De Raedt T, Brems H, Lopez-Correa C, Vermeesch JR, Marynen P, Legius E . Genomic organization and evolution of the NF1 microdeletion region. Genomics 2004; 84: 346–360.
Leppig KA, Kaplan P, Viskochil D, Weaver M, Ortenberg J, Stephens K . Familial neurofibromatosis 1 microdeletions: cosegregation with distinct facial phenotype and early onset of cutaneous neurofibromata. Am J Med Genet 1997; 73: 197–204.
De Raedt T, Brems H, Wolkenstein P, Vidaud D, Pilotti S, Perrone F et al. Elevated risk for MPNST in NF1 microdeletion patients. Am J Hum Genet 2003; 72: 1288–1292.
Wu BL, Austin MA, Schneider GH, Boles RG, Korf BR . Deletion of the entire NF1 gene detected by the FISH: four deletion patients associated with severe manifestations. Am J Med Genet 1995; 59: 528–535.
Wu BL, Schneider GH, Korf BR . Deletion of the entire NF1 gene causing distinct manifestations in a family. Am J Med Genet 1997; 69: 98–101.
Kayes LM, Burke W, Riccardi VM, Bennett R, Ehrlich P, Rubenstein A et al. Deletions spanning the neurofibromatosis 1 gene: identification and phenotype of five patients. Am J Hum Genet 1994; 54: 424–436.
Wang Y, Han KJ, Pang XW, Vaughan HA, Qu W, Dong XY et al. Large scale identification of human hepatocellular carcinoma-associated antigens by autoantibodies. J Immunol 2002; 169: 1102–1109.
Poyet JL, Srinivasula SM, Tnani M, Razmara M, Fernandes-Alnemri T, Alnemri ES . Identification of Ipaf, a human caspase-1-activating protein related to Apaf-1. J Biol Chem 2001; 276: 28309–28313.
Srinivasula SM, Poyet JL, Razmara M, Datta P, Zhang Z, Alnemri ES . The PYRIN-CARD protein ASC is an activating adaptor for caspase-1. J Biol Chem 2002; 277: 21119–21122.
Lassus P, Opitz-Araya X, Lazebnik Y . Requirement for caspase-2 in stress-induced apoptosis before mitochondrial permeabilization. Science 2002; 297: 1352–1354.
Scaffidi C, Fulda S, Srinivasan A, Friesen C, Li F, Tomaselli KJ et al. Two CD95 (APO-1/Fas) signaling pathways. EMBO J 1998; 17: 1675–1687.
De Raedt T, Maertens O, Chmara M, Brems H, Heyns I, Sciot R et al. Somatic loss of wild type NF1 allele in neurofibromas: Comparison of NF1 microdeletion and non-microdeletion patients. Genes Chromosomes Cancer 2006; 45: 893–904.
Liu JR, Opipari AW, Tan L, Jiang Y, Zhang Y, Tang H et al. Dysfunctional apoptosome activation in ovarian cancer: implications for chemoresistance. Cancer Res 2002; 62: 924–931.
Dorschner MO, Sybert VP, Weaver M, Pletcher BA, Stephens K . NF1 microdeletion breakpoints are clustered at flanking repetitive sequences. Hum Mol Genet 2000; 9: 35–46.
Cnossen MH, van der Est MN, Breuning MH, van Asperen CJ, Breslau-Siderius EJ, van der Ploeg AT et al. Deletions spanning the neurofibromatosis type 1 gene: implications for genotype-phenotype correlations in neurofibromatosis type 1? Hum Mutat 1997; 9: 458–464.
Rasmussen SA, Colman SD, Ho VT, Abernathy CR, Arn PH, Weiss L et al. Constitutional and mosaic large NF1 gene deletions in neurofibromatosis type 1. J Med Genet 1998; 35: 468–471.
Riva P, Corrado L, Natacci F, Castorina P, Wu BL, Schneider GH et al. NF1 microdeletion syndrome: refined FISH characterization of sporadic and familial deletions with locus-specific probes. Am J Hum Genet 2000; 66: 100–109.
Tonsgard JH, Yelavarthi KK, Cushner S, Short MP, Lindgren V . Do NF1 gene deletions result in a characteristic phenotype? Am J Med Genet 1997; 73: 80–86.
Upadhyaya M, Ruggieri M, Maynard J, Osborn M, Hartog C, Mudd S et al. Gross deletions of the neurofibromatosis type 1 (NF1) gene are predominantly of maternal origin and commonly associated with a learning disability, dysmorphic features and developmental delay. Hum Genet 1998; 102: 591–597.
Upadhyaya M, Spurlock G, Majounie E, Griffiths S, Forrester N, Baser M et al. The heterogeneous nature of germline mutations in NF1 patients with malignant peripheral serve sheath tumours (MPNSTs). Hum Mutat 2006; 27: 716.
Bollag G, Clapp DW, Shih S, Adler F, Zhang YY, Thompson P et al. Loss of NF1 results in activation of the Ras signaling pathway and leads to aberrant growth in haematopoietic cells. Nat Genet 1996; 12: 144–148.
Fromont-Racine M, Rain JC, Legrain P . Building protein-protein networks by two-hybrid mating strategy. Methods Enzymol 2002; 350: 513–524.
Sors A, Jean-Louis F, Pellet C, Laroche L, Dubertret L, Courtois G et al. Down-regulating constitutive activation of the NF-kappaB canonical pathway overcomes the resistance of cutaneous T-cell lymphoma to apoptosis. Blood 2006; 107: 2354–2363.
Cain K, Bratton SB, Langlais C, Walker G, Brown DG, Sun XM et al. Apaf-1 oligomerizes into biologically active approximately 700-kDa and inactive approximately 1.4-MDa apoptosome complexes. J Biol Chem 2000; 275: 6067–6070.
Twiddy D, Brown DG, Adrain C, Jukes R, Martin SJ, Cohen GM et al. Pro-apoptotic proteins released from the mitochondria regulate the protein composition and caspase-processing activity of the native Apaf-1/caspase-9 apoptosome complex. J Biol Chem 2004; 279: 19665–19682.
Laurendeau I, Bahuau M, Vodovar N, Larramendy C, Olivi M, Bieche I et al. TaqMan PCR-based gene dosage assay for predictive testing in individuals from a cancer family with INK4 locus haploinsufficiency. Clin Chem 1999; 45: 982–986.
Acknowledgements
We are grateful to Dr. Emad Alnemri (Thomas Jefferson University, Philadelphia, PA, USA) and Dr. Wei-Feng Chen (Peking University Health Science Center, Beijing, People's Republic of China) for sharing reagents. We thank Jocelyne André (INSERM U697) for technical help for real-time quantitative RT-PCR experiments and Banque de cellules de l'Assistance Publique, Hôpital Cochin, Paris, for the establishment of B lymphoblastoid cell lines.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by GM Cohen
Rights and permissions
About this article
Cite this article
Piddubnyak, V., Rigou, P., Michel, L. et al. Positive regulation of apoptosis by HCA66, a new Apaf-1 interacting protein, and its putative role in the physiopathology of NF1 microdeletion syndrome patients. Cell Death Differ 14, 1222–1233 (2007). https://doi.org/10.1038/sj.cdd.4402122
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4402122
Keywords
This article is cited by
-
Correlation between large rearrangements and patient phenotypes in NF1 deletion syndrome: an update and review
BMC Medical Genomics (2024)
-
Identification of epigenetic interactions between miRNA and DNA methylation associated with gene expression as potential prognostic markers in bladder cancer
BMC Medical Genomics (2017)
-
Emerging genotype–phenotype relationships in patients with large NF1 deletions
Human Genetics (2017)
-
Expression of ribosomopathy genes during Xenopus tropicalis embryogenesis
BMC Developmental Biology (2016)
-
Apaf1-deficient cortical neurons exhibit defects in axonal outgrowth
Cellular and Molecular Life Sciences (2015)