Abstract
DNaseY, a Ca2+- and Mg2+-dependent endonuclease, has been implicated in apoptotic DNA degradation; however, the molecular mechanisms controlling its involvement in this process have not been fully elucidated. We have obtained evidence from yeast two-hybrid screening and coimmunoprecipitation experiments that DNaseY interacted physically with actinin-α4 and this interaction significantly enhanced its endonuclease activity. Accordingly, simultaneous overexpression of both proteins in PC12 cells dramatically increased the rate of apoptosis in response to teniposide' VM26. However, overexpression of DNaseY alone neither triggered apoptosis nor facilitated cell death in response to VM26 or serum deprivation. Instead, the overexpression of DNaseY increased the production of single-strand DNA breaks and evoked a profound upregulation of DNA repair pathways. Taken together, our results point to a novel regulatory mechanism of DNaseY activity and offer an explanation for why cells must first cleave key DNA repair and replication proteins before the successful execution of apoptosis.
Similar content being viewed by others
Introduction
Progressive DNA cleavage into high molecular weight (HMW) and oligonucleosomal fragments is one of the classical hallmarks of apoptosis. At an early stage of nuclear disassembly, chromatin is cleaved into large domains of 50–300 kilobase pairs (kb). This event has been considered essential for apoptosis and it is, most likely, catalysed by an endonuclease that resides at the matrix attachment regions (MARs) where DNA binds to the nuclear scaffold.1,2 The second stage of nuclear destruction involves more extensive DNA degradation and usually produces oligonucleosomal DNA ladders. Some authors consider this stage as a housekeeping process, since blocking it does not prevent apoptosis;3 however, there are also reports arguing that the inhibition of the internucleosomal ladder formation may reduce, or at least delay apoptosis.4,5 Most apoptotic nucleases characterized to date generate oligonucleosomal DNA fragments; the exceptions are CAD (caspase-3 activated DNase) and DNaseI-like nuclease (DNaseY/DNaseγ in rat, LS-DNase/DNA1IL3 in humans), which have been implicated in both stages of nuclear DNA fragmentation.6,7,8,9,10
Multiple mechanisms have been implicated in the activation of endonucleases in apoptosis. For example, some of the enzymes translocate to the nucleus upon receiving an apoptotic signal, that is, endoG from mitochondria and DNaseI from the rough endoplasmic reticulum,11,12 whereas others, namely L-DNaseII and Helicard, are activated by proteolytic cleavage of their precursors.13,14 Activation of CAD occurs as a result of proteolysis of its inhibitor ICAD. In proliferating cells, ICAD functions as a CAD-specific chaperone, but upon receiving an apoptotic stimulus, caspases cleave it permitting CAD to degrade chromosomal DNA.6 Several reports suggest that protease-mediated early degradation of a specific subclass of DNA-binding proteins, such as the base-unpairing region (BUR)-binding proteins, increases the accessibility of endonuclease(s) to these conformationally unstable DNA sequences. This results in the generation of single-strand breaks (ssb) and, subsequently, double-strand (ds) DNA fragmentation.15,16,17 Recent reports imply that DNaseY is activated by an increase of intracellular Ca2+, which results in early DNA breaks and activation of PARP-1. Poly-ADP-ribosylation of DNaseY by PARP-1 inhibits its endonuclease activity. This negative regulation of DNaseY may be reversed upon the cleavage and inactivation of PARP-1 by a caspase-3 like protease, followed by reactivation of DNaseY by PAR glycohydrolase.18,19
PARP-1 has long been considered a molecular ‘nick sensor’ that is activated at the sites of ssb and synthesizes negatively charged poly-ADP-ribose.20,21,22 Its physical interaction with the base excision repair protein, X-ray repair cross complementing 1 (XRCC1), confirms that PARP-1 is a member of a multiprotein complex involved in the DNA polymerase β, XRCC1 and DNA ligase 3 (Polβ–XRCC1–Lig3) repair pathway.23 This pathway repairs 50–80% of ssb in living cells.24 An alternative pathway of ssb repair involves DNA polymerase δ/ɛ, proliferating cell nuclear antigen and DNA ligase1 (Polδ/ɛ–PCNA–Lig1 pathway). This pathway conducts ssb repair primarily during DNA replication.
The present study was designed to further investigate the molecular mechanisms controlling the involvement of DNaseY in apoptosis and to examine the effects of conditional overexpression of DNaseY in rat adrenal phaeochromocytoma PC12 cells. We have identified an interaction, both in vitro and in vivo, between DNaseY and actinin-α4, which resulted in a significant enhancement of DNaseY activity and have shown that concurrent overexpression of both DNaseY and actinin-α4 in PC12 cells significantly accelerated the rate of apoptosis after exposure to teniposide (VM26). Our data also revealed that the overexpression of DNaseY alone was not sufficient to trigger cell death resulting, instead, in the generation of single-strand (ss) DNA breaks and activation of the DNA repair processes that enabled cellular survival.
Results
We have previously shown that DNaseY is constitutively present in mammalian cell nuclei in tight association with chromatin;8 therefore, it was expected that such a nuclease would be strictly regulated to protect the integrity of DNA. We used yeast two-hybrid screening to search for DNaseY regulatory factor(s). Yeast strain AH109 harbouring the two-hybrid construct (pGBKT7- DNaseY) expressing human DNaseY was used to screen a human brain cDNA library for genes encoding interacting proteins (Figure 1). From 5.3 × 106 transformants, only one clone, displaying Ade/His prototrophy and β-galactosidase activity, was obtained (Figure 1). Sequence analysis and GenBank searches revealed that this clone represented a cDNA encoding the C-terminal fragment of human actinin-α4 (amino acids 448–781), a non-muscle actin-binding protein.25 The interaction between these two proteins was reproducibly reconstructed in the yeast two-hybrid system and it passed all required specificity tests (Figure 1b).
Interaction between DNaseY and actinin-α4 in yeast two-hybrid system. Empty or DNaseY containing pGBKT7 bait vector and empty or actinin-α4 containing pACT2 library vector were cotransformed into yeast host cells AH109 and plated onto SD/-Trp-Leu-Ade-His+X-α-gal plate. (a) A standard positive test showing the interaction between p53 and T antigen; (b) a positive test showing the interaction between DNaseY and actinin-α4; (c) a negative test showing no interaction between lamin C and empty library vector; (d) a negative test of empty bait vector and actinin-α4; (e) a negative test of DNaseY bait vector plus empty library vector; (f) a negative test of empty bait vector plus empty library vector
We then proceeded to establish a cellular model to further characterize the functional relationship between actinin-α4 and DNaseY. Since the endogenous level of DNaseY is very low,8 we generated stable clones with conditional regulation of DNaseY expression. The DNaseY cDNA was inserted into a bidirectional biEGFP vector and transfected into the commercially available PC12 Tet-on cell line. The endogenous level of DNaseY in this cell line was indeed very low, as shown by both RT-PCR (Figure 2a, lane 1) and Western blotting (Figure 2b, lane 1), hence these cells were considered appropriate for the overexpression studies. Several stable clones were established and shown to produce very high levels of the DNaseY transcript (Figure 2a), particularly in the presence of Dox, but even in its absence (due to promoter leakiness). Similar differences were evident at the protein level (Figure 2b).
Dox-inducible overexpression of DNaseY in PC12 cells. The PC12 Tet-on cells were stably transfected with DNaseY/biEGFP or empty biEGFP vectors and cultured for 2 weeks in the presence or absence of 1?μg/ml of Dox. The cells were harvested, total cellular RNA and proteins were extracted and analysed by a semi-quantitative RT-PCR (a) and Western blotting (b) as described in Materials and Methods. (a) Ethidium bromide-stained agarose gel of RT-PCR products of DNaseY and GAPDH: lane 1 – cells transfected with empty biEGFP; lane 2 – cells transfected with DNaseY/biEGFP but not exposed to Dox; lane 3 – cells transfected with DNaseY/biEGFP and treated with Dox for 2 weeks (clone 7); M – size markers. (b) Western blot analysis of DNaseY gene product. Equal amounts of total cellular protein (150?μg/lane) were separated on 10% SDS-PAGE, electrotransferred onto a membrane and immunoblotted with anti-DNaseY antibody. Lanes 1–3 represent the same samples as in (a)
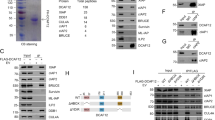
Nuclear colocalization of DNaseY and actinin-α4 was established based on immunofluorescence staining with specific antibodies (Figure 3). Confocal microscopy revealed that the DNaseY protein was localized mainly in the nuclei of overexpressing cells, although some cytoplasmic staining was also observed (Figure 3a). Actinin-α4, on the other hand, was localized predominantly in the cytoplasmic compartment, likely in the association with the cytoskeleton (Figure 3b). However, a punctuate nuclear immunostaining, indicative of the presence of actinin-α4-positive microdomains was also observed throughout the nucleus. The nuclear presence of actinin-α4 was further confirmed by Western blotting of nuclear protein extracts from the nuclei that were extensively washed and passed through a 2M-sucrose cushion. Based on both the fluorescence (Figure 3c) and phase-contrast (Figure 3d) images, the isolated nuclei were intact, free of cytoskeletal contamination and extracts from these nuclei clearly contained actinin-α4 (Figure 3e, lane 1). This established that DNaseY and a portion of actinin-α4 resided in the nucleus. The protein extract from DNaseY-overexpressing cells (Figure 3f, lane 1) was subsequently used to demonstrate an in vivo physical interaction between the two proteins. Immunoprecipitation was performed with anti-actinin-α4 antibody and the Western blot was developed with anti-DNaseY antibody. The blot clearly revealed the presence of DNaseY in the immunoprecipitates (Figure 3f, lane 2).
Nuclear colocalization and physical interaction between DNaseY and actinin-α4. (a, b) The DNaseY-overexpressing clone 7 was treated with Dox for 2 weeks. The cells were plated on glass coverslips, fixed and immunostained with anti-DNaseY (a) and anti-actinin-α4 (b) antibodies. The cells were examined and photographed under a Zeiss confocal microscope. (c, d) Nuclei were isolated from the Dox-treated clone 7, washed with cytoskeletal stripping buffer, passed through 2?M sucrose cushion and stained with Hoechst 33258 dye. Images, fluorescence (c) and phase contrast (d) were captured on an Olympus B max fluorescence microscope. (e) Proteins were extracted from Dox-treated cells (lane 2) and from isolated nuclei (lane 1), separated on 10% SDS-PAGE (150?μg/lane), electrotransferred onto a nitrocellulose membrane and analysed by Western blotting with anti-actinin-α4 antibody. (f) Total cellular proteins from Dox-treated clone 7 were extracted and immunoprecipitated with anti-actinin-α4 antibody as described in Materials and Methods. The precipitates were separated on 10% SDS-PAGE, electrotransferred onto a nitrocellulose membrane and analysed by Western blotting with anti-DNaseY antibody. Lane 1 – total cellular proteins from Dox-treated clone 7; lane 2 – proteins immunoprecipitated with anti-actinin-α4 antibody; lane 3 – mock immunoprecipitation in the absence of primary actinin-α4 antibody
To determine the biological significance of this interaction, we assessed the effects of actinin-α4 on DNaseY activity in vitro. The assay was performed using recombinant His-tag proteins and circular plasmid DNA as a substrate (Figure 4). We have shown previously that DNaseY is optimally active in the presence of Ca2+ and Mg2+ ions and is capable of single- and double-stranded DNA cleavage.8 In the reconstituted in vitro DNaseY activity assay, performed in the presence of both divalent cations, actinin-α4 significantly accelerated, in a concentration-dependent manner, the digestion of circular plasmid DNA (Figure 4a). This effect was actinin-α4-specific, since a recombinant Sox2 protein, produced and purified in the same fashion as actinin-α4, had no effect on the DNaseY activity (Figure 4b). The concentration-dependent actinin-α4 enhancement of nuclease activity was also seen in assays containing only one cation, that is, Mg2+ only (Figure 4c and d) or Ca2+ only (Figure 4e), although under such conditions the enzyme was far less active. This was especially true when Ca2+ ions were chelated by the addition of 2?mM EGTA (Figure 4d). However, in the absence of both cations actinin-α4 failed to activate DNaseY (Figure 4f). Taken together, these results showed that actinin-α4 could stimulate DNaseY activity at low concentrations of divalent cations, but not in their absence.
In vitro modulation of DNaseY activity by actinin-α4. Increasing amounts of recombinant His-tag actinin-α4 protein (0, 25, 125 and 500?ng, reconstituted in 20?mM Tris-HCl, pH 7.4) were combined with 1?μg of plasmid DNA and preincubated for 30?min at 37°C. An amount of 300?ng recombinant His-tag DNaseY was added to the reactions and incubated for the additional 60?min at 37°C. (a) Plasmid digestion was carried out in a 20?μl reaction buffer containing 20?mM Tris-HCl pH 7.4, 5?mM MgCl2 and 2?mM CaCl2; (b) the same as in (a) except that actinin-α4 was replaced with His-tag Sox2 protein; (c) the same as in (a) but without CaCl2; (d) the same as in (c) plus 2?mM EGTA; (e) the same as in (a) but without MgCl2; (f) plasmid digestion was carried out in 20?mM Tris-HCl pH 7.4 in the absence of both CaCl2 and MgCl2. Abbreviations: OC – open circle plasmid; Li – linear plasmid; SC – super coil plasmid. Ac – actinin α4; DY – DNaseY; Sx – Sox2
Recently, Shiokawa and Tanuma9 reported that overexpression of DNaseY/DNaseγ in HeLa S3 cells increased the proportion of apoptotic cells in response to C2-ceramide. This assessment was based on morphological changes. We also tested the sensitivity of DNaseY-overexpressing PC12 cells to different apoptosis inducing treatments, that is, the topoisomerase II inhibitor teniposide (VM26) and deprivation of growth factors (Figure 5). Several clones were selected, cultured for 2 weeks in the presence of Dox to achieve maximal induction of DNaseY expression (Figure 2) and subsequently treated for 24?h with 10?μM VM26 (Figure 5a). Approximately 40–50% of cells lost viability as a result of this treatment; however, the overexpression of DNaseY, clearly, had no effect on the rate of cell death in response to VM26. Similar behaviour was observed after a 24?h period of serum withdrawal (Figure 5b). None of the overexpressing clones showed higher sensitivity than control cells carrying empty vectors. The DNA degradation pattern in response to VM26 treatment was analysed by PFGE and is shown in Figure 6. These cells did not produce oligonucleosomal ladders; instead, the DNA was cleaved into a range of large fragments of 300?kb and above as well as 50?kb and below (Figure 6). Similar results were obtained with other treatments (data not shown). Taken together, these results indicated that the overexpression of DNaseY by itself neither triggered apoptotic cell death nor accelerated it in response to the well-known triggers, that is, VM26 and serum deprivation.
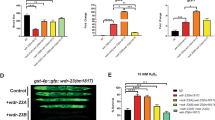
Induction of apoptosis in DNaseY-transfected clones. The PC12 Tet-on cells carrying empty biEGFP vector (control) and cells transfected with DNaseY/biEGFP (clones 5, 7 and 8) were cultured for 2 weeks in the presence of 1?μg/ml of Dox. Cell death was induced by a 24?h exposure to either 10?μM VM26 (a) or serum withdrawal (b). Cell viability was assessed by counting trypan blue positive cells using a haemocytometer
Analysis of DNA fragmentation by PFGE. Control PC12 Tet-on cells with empty biEGFP vector and DNAseY overexpression clone 7 were cultured for 2 weeks in the presence of 1?μg/ml of Dox. Apoptosis was induced by the 24?h treatment with 10?μM VM26. Equal numbers of cells were embedded in LMP agarose plugs and analysed by PFGE as described in Materials and Methods. The gel was stained with ethidium bromide and photographed on a transilluminator. Lane 1 – control untreated cells; lane 2 – clone 7 untreated; lane 3 – control cells treated with VM26; lane 4 – clone 7 treated with VM26. The size markers were: 123?bp DNA ladder; low range PFG marker (λ DNA - Hind III fragments); ladder PFG marker; yeast chromosome PFG marker (S. cerevisiae YPH 80)
The lack of sensitivity of the overexpressing cells contradicted the previously published reports9 and raised a question of whether the clones were actually expressing active DNaseY enzyme. To address this, we isolated the nuclei and examined in vitro chromatin digestion during a 60?min incubation period (Figure 7). After the incubation, the nuclei were embedded in agarose plugs and DNA integrity was assessed by PFGE.26 A comparison was made between the nuclei obtained from cells carrying empty biEGFP vector and from cells containing the vector encoding DNaseY. Prior to isolation of the nuclei, the cells were cultured for 2 weeks with Dox to achieve full induction of DNaseY expression. There was no DNA degradation in the absence of divalent cations, regardless of whether the cells expressed DNaseY or not (Figure 7, lanes 6–9). In the presence of Mg2+ ions, there was a significant difference between the control and overexpressing clones (Figure 7, compare lanes 10 and 11 to lanes 12 and 13), especially after the Dox treatment when the entire nuclear DNA content was converted to oligonucleosomal ladders (Figure 7, lane 13). More extensive chromatin digestion was observed during incubation with both Ca2+ and Mg2+ (Figure 7, lanes 14–17), especially in nuclei from the overexpressing cells in which DNA digestion neared completion (Figure 7, lanes 16 and 17). These results showed that DNaseY was active and capable of efficient chromatin fragmentation in isolated nuclei. Yet, in the context of the intact cell, its presence did not accelerate apoptosis.
PFGE analysis of DNA fragmentation in isolated nuclei. Isolated nuclei were incubated for 60?min at 37°C in the presence or absence of divalent cations and embedded in LMP agarose plugs. Equal amounts of DNA were loaded onto a 0.8% agarose gel and analysed by PFGE as described in Materials and Methods. The gel was stained with ethidium bromide and photographed on a transilluminator. Lanes 6, 10 and 14 – nuclei from the empty vector control cells not treated with Dox; lanes 7, 11 and 15 – nuclei from control cells cultured for 2 weeks in the presence of 1?μg/ml of Dox; lanes 8, 12 and 16 – nuclei from clone 7 not treated with Dox; lanes 9, 13 and 17 – nuclei from clone 7 treated with Dox for 2 weeks. The molecular size markers used were: yeast chromosome PFG marker (S. cerevisiae YPH 80); λ ladder PFG marker; low range PFG marker (λ DNA - Hind III fragments); λ DNA Hind III marker; 123?bp DNA ladder
Thus far, we have established that actinin-α4 was capable of modulating DNaseY activity in vitro (Figure 4). To test their interaction in vivo, we transiently transfected a pcDNA 3.1/Myc-His plasmid harbouring full-length human actinin-α4 cDNA into the control and DNaseY-overexpressing cells and challenged the cells with either 10?μM VM26 or 1?μM staurosporine for 24?h (Figure 8). Again, in the absence of exogenous actinin-α4, the response of the DNaseY-overexpressing clone was the same as the empty vector control (approximately 40% of cells lost membrane integrity and became trypan blue positive). The expression of actinin-α4 in control cells also had no effect on their sensitivity to VM26. However, the picture was vastly different after the introduction of actinin-α4 into the DNaseY-overexpressing cells; the rate of cell death doubled and, within 24?h, approximately 70% of cells were dead (Figure 8). The cells were dying by apoptosis as evidenced by the characteristic chromatin condensation seen in Hoechst dye-stained nuclei (Figure 8, inset). Similar results were obtained in response to staurosporine (data not shown). These experiments confirmed that actinin-α4 was capable of DNaseY activation in vivo.
In vivo activation of DNaseY by actinin-α4. The empty vector control and DNAseY-overexpressing clone 7 were treated for 2 weeks with 1?μg/ml of Dox and were subsequently transiently transfected with a plasmid encoding the full length actinin-α4 cDNA. After 24?h, the transfected cells were challenged with 10?μM VM26 and cell viability was assessed by trypan blue exclusion assay after the additional 24?h. −DNaseY/−actinin – nontransfected empty vector control; +DNaseY/−actinin – nontransfected clone 7; −DNaseY/+actinin – control cells transfected with actinin-α4 cDNA; +DNaseY/+actinin – clone 7 transfected with actinin-α4. Inset: Cells (clone 7) were exposed to VM26 for 24?h, fixed and stained with Hoechst dye and examined under an Olympus B × 50 fluorescence microscope
The question still remained as to why prolonged overexpression of the active endonucleolytic enzyme had no effect on the sensitivity to apoptosis? For example, we have treated the clones with Dox for up to 5 weeks and established that these cells continued to produce very high levels of DNaseY (Figure 2) and, yet, they continued to grow and proliferate and did not change their susceptibility to the apoptotic inducers. However, while searching for evidence of nuclear DNA degradation, using the terminal deoxynucleotidyl transferase (TdT)-mediated nick end labelling (TUNEL) assay, we discovered that the entire population of DNaseY-overexpressing cells, especially after 2 weeks of induction with Dox, was brightly stained and TUNEL-positive (Figure 9a–c). Without Dox treatment, the intensity of TUNEL staining was much lower (Figure 9d–f). Moreover, the nuclear morphology of these cells was normal, without any evidence of apoptotic chromatin collapse, and the cells continued to grow and proliferate at the same rate as control cells with TUNEL-negative nuclei (Figure 9g–i). In the control cell population, TUNEL-positive nuclei with classical apoptotic morphology were only occasionally seen (Figure 9a–c). Clearly, the overexpression of active DNaseY leads to the generation of extensive DNA breaks with 3′-OH ends recognized and labelled by terminal deoxynucleotidyl transferase employed in the TUNEL assay. This observation raises concerns about the reliability of the TUNEL assay for the detection of early apoptotic cells, as every cell overexpressing DNaseY was TUNEL positive, but had normal nuclear morphology and continued to proliferate (Figure 9g and h).
Identification of ss DNA breaks by TUNEL. The DNaseY-overexpressing clone 7 treated (a–c) or nontreated (d–f) for 2 weeks with Dox and empty vector control cells (g–i) were plated on glass coverslips, fixed and labelled with terminal deoxynucleotidyl transferase using biotin-16-dUTP and CY3-conjugated streptavidin (a, d and g). Nuclei were counterstained with DAPI (b, e and h). The same cells were also photographed with phase contrast optics (c, f and i)
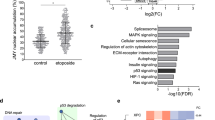
The presence of ss DNA breaks was further revealed by two-dimensional gel electrophoresis.27 Although DNA remained in the wells and seemed intact after separation in the first dimension by PFGE (Figure 10, top panel), alkaline denaturation prior to conventional electrophoresis in the second dimension liberated a tail of ss DNA fragments. This was especially evident in Dox-treated DNaseY overexpressing cells (Figure 10, bottom panel). Therefore, to grow, proliferate and faithfully replicate their genetic information, these cells must have an efficient DNA repair system to cope with the unusually high levels of ssb. To test this, we used Western blotting to determine the levels of several components of the DNA repair pathways (Figure 11). Firstly, we found that induction of DNaseY expression caused a significant upregulation of PARP-1, the ssb sensor protein (Figure 11a, lane 3). Secondly, there was also an induction of the Polβ–XRCC1–Lig3 pathway, as manifested by the increased expression of XRCC1 and Lig3 proteins. XRCC1 is the docking scaffold protein that interacts with PARP, PNK, Polβ, and Lig3. As shown in Figure 11b, its upregulation in the DNaseY-overexpressing cells was very significant, as was the elevation of both the 97 and 100?kDa forms of Lig3 (Figure 11c). The expression of Polβ, on the other hand, did not change (Figure 11d). Finally, the expression of two components of the Polδ/ɛ–PCNA–Lig1 complex, PCNA and Lig1, was also upregulated in the DNaseY-overexpressing cells (Figure 11e and f).
Western blotting analyses of DNA repair components. Nuclei were isolated from the empty vector control cells (lane 1) and from DNaseY-overexpressing clone 7, noninduced (lane 2) and induced by Dox (lane 3), and nuclear proteins were extracted as described in Materials and Methods. Amounts of 50?μg per lane of total nuclear proteins were separated on 10% SDS-PAGE, electrotransferred onto a nitrocellulose membrane and immunoblotted with anti- PARP-1 (a), anti-XRCC1 (b), anti-Lig3 (c), anti-Polβ (d), anti-PCNA (e) and anti-Lig1 (f) antibodies
Discussion
This study was undertaken to examine molecular mechanisms controlling the involvement of DNaseY in apoptosis. Previous studies have shown that this nuclease meets most of the biochemical and functional criteria (i.e., cation dependence, pH profile, ability to cleave ss and ds DNA with 3′-OH ends, nuclear localization) expected of an enzyme engaged in the apoptotic process.8 The constitutive presence of DNaseY in the nucleus suggests that its physiological role might be linked to the maintenance of DNA integrity and that its participation in cell death process must be tightly controlled. We have demonstrated that actinin-α4 protein could modulate DNaseY enzymatic activity both in vitro and in vivo (Figures 4 and 8). In the in vitro assay, actinin-α4 significantly stimulated it, particularly under less than optimal concentration of divalent cations, although not in their absence (Figure 4). This interaction in vivo resulted in a highly accelerated rate of cell death in response to apoptotic stimuli (i.e. VM26 and staurosporine, Figure 8).
The molecular mechanisms that govern the interaction and activation of DNaseY by actinin-α4 are at present unknown. Actinin-α4 is one of the four mammalian α-actinin genes, all of which encode highly homologous, approximately 100?kDa actin crosslinking or bundling proteins.28 These proteins contain a C-terminal EF hand calcium-binding domain and are believed to link the actin cytoskeleton to focal adhesions and control cell motility. Although actinin-α4 is a calcium-binding protein, it did not substitute for the requirements of divalent cations and DNaseY was completely inactive in their absence (Figure 4). However, the in vivo data showed that cell death was accelerated only when both proteins were overexpressed simultaneously, implying that the activation process might require formation of a physical complex, which is supported by the results of the coimmunoprecipitation experiments (Figure 3).
We have established that in addition to a prominent cytoplasmic localization, a fraction of actinin-α4 was also present in the nuclei of PC12 cells (Figure 3). These observations are consistent with those of Honda et al.,25 who reported recently that actinin-α4 exists in the nuclei of a certain population of breast cancer cell lines. Furthermore, these authors show that actinin-α4 translocates from the cytoplasm to the nucleus following treatments with wortmannin, the PI3 kinase inhibitor or cytochalasin D, the actin-depolymerizing drug. This translocation alters cellular motility and abolishes the cells' tumorigenic and metastatic potential.25,29 Although the precise mechanism of nuclear translocation has not been identified (actinin-α4 does not contain any apparent nuclear localization signals), the involvement of PI3 kinase is of great interest. PI3 kinase is one of the key molecules involved in the signalling pathways of activated receptor tyrosine kinases and regulates vital cellular processes (i.e., cell growth, motility, morphogenesis) as well as protecting the cells from apoptosis.30 Therefore, we may speculate that activation of DNaseY, at least in some cell types, could be brought about by the nuclear translocation of actinin-α4 facilitated by the inactivation of PI3 signalling. This sequence of events is supported by our results. For example, the active nuclease was capable of complete destruction of nuclear chromatin in the isolated nuclei (Figure 7), but not in intact cells (Figure 5). Thus, only after the removal of cytoplasmic control, or after the introduction of exogenous actinin-α4 (Figure 8), the effect of DNaseY overexpression became evident. It is also possible to speculate that the tumour suppressor function of actinin-α4 might be linked to its ability to stimulate DNaseY and consequently enhance apoptosis of cancer cells.25,29
The true physiological function of DNaseY is still not known, although its nuclear presence and close association with chromatin suggests that it plays a role in DNA metabolism. We have demonstrated here, for the first time, that DNaseY produced widespread ss DNA breaks. However, extensive accumulation of ssb (Figure 9) did not push the PC12 cells into apoptosis. This is in agreement with the previous studies, which reported that transfection of the same nuclease into HeLa cells or mouse skin fibroblasts increased cell death only in response to apoptotic stimuli, but not by itself.9,18 In our cell system, overexpression of DNaseY led to the activation of DNA repair pathways. The components of both repair systems, the Polβ–XRCC1–Lig3 pathway, which conducts both short and long-patch repair as well as the Polδ/ɛ–PCNA–Lig1 pathway, which provides an alternative or ‘backup’ mechanism for ssb repair and often conducts long-patch break repair,24 were clearly upregulated in the DNaseY-overexpressing cells (Figure 11e and f). The expression of Polβ, on the other hand, was not altered (Figure 11d) suggesting that DNaseY generated simple nicks on the DNA strands rather than gaps, which would require the activity of polymerase. Therefore, to fully engage in the execution of apoptosis, the cells must also degrade the key proteins involved in the DNA repair processes.31
It has been reported that, at early stages of apoptosis, DNaseY can be inactivated by PARP-1 through the poly-ADP-ribosylation mechanism.18,19 Here we found a significant upregulation of PARP-1 in the DNaseY-overexpressing cells (Figure 11a). Therefore, it is possible that PARP-1 acts both as a sensor to detect ssb and at the same time to synthesize negatively charged ADP-ribose polymers to inhibit the endonuclease that creates them.20,32 In the execution phase of apoptosis, PARP-1 is cleaved and inactivated by caspase-like proteases relieving inhibition of DNaseY;18,19 however, the inactivation of PARP-1 may not be sufficient to fully unleash the activity of DNaseY. The cytoplasmic changes, that is, alterations in signalling pathways and cytoskeletal changes, might also be required to accomplish this.
In conclusion, we have established that DNaseY, a Ca2+- and Mg2+-dependent endonuclease, was capable of producing ss DNA breaks and its activity was modulated by actinin-α4. Moreover, the ssb's produced by exogenous DNaseY were not detrimental to the cells as the cells activated the DNA repair processes to cope with this burden. Although it is still not clear whether, under normal physiological conditions, DNaseY plays a role in DNA repair and/or replication, the accumulated evidence suggests that it does also have a role in apoptosis. Other endonucleases implicated in apoptosis also have regular housekeeping functions. For example, EndoG is a mitochondrial enzyme that plays a role in DNA replication. It becomes involved in apoptotic process only when released from the mitochondria.11,33 L-DNaseII in its native form is a serine protease inhibitor (LEI); it transforms into a nuclease during apoptosis.34
Materials and Methods
Cell culture and experimental treatments
Rat adrenal phaeochromocytoma PC12 Tet-On cell line was purchased from Clontech (Palo Alto, CA, USA). The cells were cultured in Dulbecco's modified Eagle's medium (DMEM) supplemented with 10% horse serum, 5% fetal calf serum (Wisent Inc., St. Bruno, QC), 0.1?mg/ml gentamycin sulphate (Sigma Cell Culture, St. Louis, MO, USA), 230?μg/ml active G418 sulphate (Gibco BRL, Burlington, ON), and 25?μg/ml hygromycin B (Boehringer Mannheim, Montreal, QC). To induce the expression of DNase Y transcripts, cells were cultured in a medium containing 1?μg/ml Doxycycline (DOX, Sigma, Oakville, ON) for 2 weeks prior to the initiation of experiments. Cell death was triggered by either 24?h exposure to 10?μM VM26, 1?μM staurosporine or by serum withdrawal. Cell viability was assessed by the trypan blue exclusion assay using a haemocytometer.
Cell transfections
A fragment of DNase Y cDNA (lacking the N-terminal 25 amino acids encoding the signal peptide) encoding the active DNaseY protein was cloned into the biEGFP vector (Clontech, Palo Alto, CA, USA) and used to generate stable clones with conditional Tet-on regulation of DNaseY expression. The cells were transfected using the Calphos Mammalian transfection kit (Clontech, Palo Alto, CA, USA). Briefly, the PC12 Tet-on cells, cultured for 24?h in 10?cm Petri dishes (2 × 106 cells per dish), were placed in fresh medium 1?h prior to the transfection. Amounts of 10?μg of DNaseY/biEGFP or empty biEGFP vector control along with 2.5?μg of pHeBo were used per Petri dish. The transfection cocktail was prepared by adding the cDNA plasmids and calcium-Maximizer solution to 2 × PBS buffer drop by drop with constant bubbling and a subsequent 15?min incubation at room temperature. A volume of 700?μl of this transfection cocktail was added to each Petri dish. The cells were incubated for 5?h at 37°C, washed once with PBS and placed in 10?ml of fresh complete growth medium. At 72?h post transfection, the cells were trypsinized, split 1?:?5 and subsequently cultured in growth medium containing 25?μg/ml hygromycin B (Boehringer Mannheim, Germany). Positive colonies were selected using a Zeiss IM35 inverted fluorescence microscope.
Transient transfections of DNaseY-overexpressing cells with a plasmid encoding actinin-α4 were also performed. Full-length human actinin-α4 cDNA was cloned into pcDNA 3.1/Myc-His vector (Invitrogen, Burlington, ON). Cells were plated in 12-well plates (0.4 ×106 cells/well) 1 day before and were transfected with 5?μg/well of purified plasmid harbouring actinin-α4 cDNA and 15?μl of LipofectAmine 200 reagent (Invitrogen, Burlington, ON) according to the manufacturer's instruction.
RNA extraction and RT-PCR
Total RNA extraction, first-stand cDNA synthesis and RT-PCR analysis of DNaseY transcript were performed as previously described.8
Antibody production
Rat cDNA encoding an active DNaseY protein was cloned into pBAD/HisA vector (Invitrogen, Burlington, ON). Recombinant His-tag DNaseY protein was purified by His-Trip nickel column (Pharmacia Biotech, Baie d' urfe', QC). The proteins were freeze-dried and reconstituted in PBS for vaccination. White male New Zealand rabbits were first immunized with 500?μg of protein/inoculation according to standard operating protocol. Animals were boosted three times with 250?μg protein, within a 4-week interval. The titre of the antibody was tested at 10–14 days after each boosting. Polyclonal anti-DNaseY antibody was collected 14 days after the third boosting.
Immunofluorescence staining
Cells were plated on poly-L-lysine-coated coverslip, fixed with paraformaldehyde (J.B. EM Services Inc., Point-Claire, PQ). Fixed cells were immunostained with one of the following primary antibodies diluted in PBS: anti-DNaseY (1?:?300 v/v8), or anti-actinin-α4 (1?:?10 v/v25). Cells were then incubated for 1?h in the corresponding secondary antibody: Cy3- conjugated goat anti-mouse Ig G(Fc) (1?:?200) or CY3-conjugated goat anti-mouse IgM(μ) (diluted 1?:?200, both from Jackson ImmunoResearch/BioCan Scientific, Mississauga, ON). Finally, the nuclei were counterstained with 0.2?μg/ml of 4′, 6-diamidino-2-phenylindole (DAPI) or Hoechst 33258 (Sigma, Oakville, ON) in PBS for 5?min and were mounted in Vectashield mounting medium (Vector Laboratories Inc., Burlingame, CA, USA). Optical sections were obtained on a Zeiss LSM-410 (Carl Zeiss, Thornwood, NY, USA) inverted laser scanning microscope equipped with a krypton/argon laser. CY3-labelled cells were analysed using a 530–585?nm-band pass filter and dichroic mirror (FT 488/568) for excitation, and a 575–640?nm-band pass filter to detect emission.
DNA breaks were detected by TUNEL essentially performed as described by Gavrieli et al.35 using biotin-16-dUTP (Boehringer Mannheim, Laval, PQ) and visualized with CY3-conjugated streptavidin (Jackson ImmunoResearch, West Grove, PA, USA). The cells were examined on an Olympus B max fluorescence microscope with Olympus × 40 N.A. 1.0 planapo oil immersion objective and images were processed using Northern Exposure software and Adobe Photoshop 5.5.
Protein extraction
Total cellular and nuclear proteins were extracted as described by Liu et al.17 The cytoskeletal structures were removed from the nuclear pellet by washing with cytoskeletal stripping buffer (BNB buffer containing a 2?:?1 ratio (v/v) of NP-40 and sodium deoxycholate in a final concentration of 1%?:?0.5%) for 15?min and centrifugation at 3000?r.p.m. for 5?min at 4°C. To ensure total removal of any cytoplasmic protein, the nuclei were resuspended in 300?μl of BNB, mixed with 3?ml sucrose cushion buffer (50?mM Tris-HCl, pH 7.4, 2?M sucrose, 5?mM MgCl2) and loaded on a top of 1?ml sucrose cushion buffer in a 5?ml ultracentrifuge tube. After centrifugation at 13?000?r.p.m. for 1?h, the nuclear pellet was further washed with fresh BNB buffer and a small sample was examined under the microscope to check for cytoplasmic protein contamination. The nuclei were finally collected by centrifugation at 3000?r.p.m. for 5?min at 4°C. The pellet was resuspended in a buffer containing 20?mM HEPES, pH 7.9, 420?mM NaCl, 25% glycerol, 0.2?mM EDTA, 1.5?mM MgCl2, 0.5?mM DTT and 0.5?mM PMSF and incubated on ice for 30?min. The nuclear proteins were collected either by ultracentrifugation at 55?000?r.p.m. at 4°C for 10?min, or by direct sonication on ice for 10 × 5?s.
Western blotting
Immunoblottings were performed as previously described.8 The blots were probed with the following primary antibodies: rabbit polyclonal anti-DNaseY (dilution 1?:?1000), mouse monoclonal anti-DNA ligase III (1?:?200 v/v, Genetex, San Diego, CA, USA), mouse monoclonal anti-XRCC1 (1?:?1000 v/v, Trevigen, Gaithersburg, MD, USA), mouse monoclonal anti-DNA ligase I (1?:?200 v/v, QED Biosciences Inc, San Diego, CA, USA), mouse monoclonal anti-PCNA (1?:?200 v/v, Calbiochem, Hornby, ON), mouse monoclonal anti-DNA polymerase β (1?:?100 v/v, NeoMarker, Fremont, CA, USA), mouse monoclonal anti-PARP (1?:?5000 v/v, Upstate Biotechnology, Charlottesville, VA, USA) and mouse monoclonal anti-actinin-α4 (1?:?10 v/v25). The antigens were detected using horseradish peroxidase-conjugated secondary antibodies?:?goat anti-mouse IgG (1?:?5000 v/v), donkey anti-rabbit IgG (1?:?5000 v/v, both from Promega Corporation, Madison, WI, USA), goat anti-mouse IgM (1?:?5000 v/v, Jackson ImmunoResearch, West Grove, PA, USA) and the complexes were revealed by enhanced chemiluminescence using the ECL kit (Amersham Biosciences, Baie d' urfe', PQ, Canada).
Coimmunoprecipitation
Magnetic Dynabeads (Dynal Biotech Inc.,. Lake Success, NY, USA) were prepared by resuspending 20?μl of beads in 500?μl of Co-IP buffer (20?mM Tris–HCl, pH 7.5, 150?mM NaCl, 0.1% Tween–20, 0.1% BSA, 1 × protease inhibitor cocktail), mixing on a rotator for 2?min, then recovering the beads with a magnetic stand. This step was repeated three times. The beads were then mixed with 500?μl Co-IP buffer and 2?μl anti-mouse IgM conjugated with biotin (Jackson ImmunoResearch, West Grove, PA, USA) for 30?min, recovered, washed five times with Co-IP buffer, then blocked in 500?μl Co-IP buffer with 0.5% BSA for 30?min on a rotator. The beads were again recovered and washed twice with Co-IP buffer. The sample protein was cleaned by mixing 500?μg to 1?mg with the beads in 100?μl Co-IP buffer for 1?h at 4°C. The beads were discarded; the protein was recovered and mixed with 5?μl of mouse monoclonal anti-actinin-α4 antibody25 for 2?h at 4°C. This mixture was added to a fresh set of beads, prepared as described above, and mixed overnight at 4°C. The beads were recovered, washed three times with Co-IP buffer without BSA. The bead-bound complexes were released by boiling in protein loading buffer for 5?min. The presence of DNaseY in the complex was revealed by Western blotting as described above.
Analysis of DNA fragmentation
Nuclei auto-digestion was carried out as described in Pandey et al.26 Conventional, pulsed field and two-dimensional gel electrophoresis of DNA was performed exactly as previously described.27
Yeast two-hybrid screening
Human DNaseY cDNA encoding the active DNaseY protein was cloned into pGBKT7 vector (Clontech, Palo Alto, CA, USA) to generate a chimaeric open reading frame encoding Gal4 DNA binding domain and DNaseY protein. This construct was introduced into yeast Saccharomyces cerevisiae strain AH109. A single colony containing cells harbouring the DNaseY/pGBKT7 plasmid was then used as host cells for screening a human cDNA expression library constructed in pACT2 vector (Clotech, Palo Alto, CA, USA). The protein–protein interaction was first screened by plating the transformants onto SD/-Trp-Leu-His-Ade selection plates. Positive clones were then rescreened for the presence of β-galactosidase activity to eliminate false-positive interaction. Library plasmids harbouring DNaseY interacting proteins were rescued and reintroduced into the DNaseY/pGBKT7 containing host cell to further eliminate false interaction. The identity of the cDNA encoding DNaseY-interacting protein was revealed by DNA sequencing and database searches.
DNase Y activity assay
The activity assay was performed by preincubation of 1?μg of plasmid DNA with variable amounts of purified His-tag human actinin-α4 protein in 20?μl buffer containing 20?mM Tris-HCl, pH 7.4, 5?mM MgCl2 and 2?mM CaCl2 for 30?min at 37°C. A volume of 5?μl (0.3?μg) purified His-tag rat DNaseY was then added to the reaction and incubated for 60?min at 37°C. The pattern of plasmid DNA digestion was analysed by electrophoresis on 0.8% agarose gels in 40?mM Tris-acetate buffer pH 8.5 and 2?mM EDTA at 20?V overnight. Gels were stained with ethidium bromide and photographed on a UV transilluminator, and analysed by agarose gel electrophoresis in TAE buffer.
Abbreviations
- MARs:
-
matrix attachment regions
- ssb:
-
single-strand break
- BUR:
-
base-unpairing region
- ds:
-
double strand
- ss:
-
single strand
- XRCC1:
-
X-ray repair cross complementing 1
- Polβ:
-
DNA polymerase β
- Lig3:
-
DNA ligase 3
- Polδ/ɛ:
-
DNA polymerase δ/ɛ
- PCNA:
-
proliferating cell nuclear antigen
- Lig1:
-
DNA ligase1
References
Lagarkova MA, Iarovaia OV and Razin SV (1995) Large-scale fragmentation of mammalian DNA in the course of apoptosis proceeds via excision of chromosomal DNA loops and their oligomers. J. Biol. Chem. 270: 20239–20241
Walker PR and Sikorska M (1997) New aspects of the mechanism of DNA fragmentation in apoptosis. Biochem. Cell Biol. 75: 287–299
Sakahira H, Enari M and Nagata S (1998) Cleavage of CAD inhibitor in CAD activation and DNA degradation during apoptosis. Nature 391: 96–99
Walisser JA and Thies RL (1999) Poly(ADP-ribose) polymerase inhibition in oxidant-stressed endothelial cells prevents oncosis and permits caspase activation and apoptosis. Exp. Cell Res. 251: 401–413
Boulares AH, Zoltoski AJ, Sherif ZA, Yakovlev AG and Smulson ME (2002) The Poly(ADP-ribose) polymerase-1-regulated endonuclease DNAS1L3 is required for etoposide-induced internucleosomal DNA fragmentation and increases etoposide cytotoxicity in transfected osteosarcoma cells. Cancer Res. 62: 4439–4444
Enari M, Sakahira H, Yokoyama H, Okawa K, Iwamatsu A and Nagata S (1998) A caspase-activated DNase that degrades DNA during apoptosis, and its inhibitor ICAD. Nature 391: 43–50
Baron WF, Pan CQ, Spencer SA, Ryan AM, Lazarus RA and Baker KP (1998) Cloning and characterization of an actin-resistant DNase I-like endonuclease secreted by macrophages. Gene 215: 291–301
Liu QY, Pandey S, Singh RK, Lin W, Ribecco M, Borowy-Borowski H, Smith B, LeBlanc J, Walker PR and Sikorska M (1998) DNaseY: a rat DNaseI-like gene coding for a constitutively expressed chromatin-bound endonuclease. Biochemistry 37: 10134–10143
Shiokawa D and Tanuma S (1998) Molecular cloning and expression of a cDNA encoding an apoptotic endonuclease DNase gamma. Biochem. J. 332: 713–720
Yakovlev AG, Wang G, Stoica BA, Simbulan-Rosenthal CM, Yoshihara K and Smulson ME (1999) Role of DNAS1L3 in Ca2+- and Mg2+-dependent cleavage of DNA into oligonucleosomal and high molecular mass fragments. Nucleic Acids Res. 27: 1999–2005
Li LY, Luo X and Wang X (2001) Endonuclease G is an apoptotic DNase when released from mitochondria. Nature 412: 95–99
Peitsch MC, Polzar B, Stephan H, Crompton T, MacDonald HR, Mannherz HG and Tschopp J (1993) Characterization of the endogenous deoxyribonuclease involved in nuclear DNA degradation during apoptosis (programmed cell death). EMBO J. 12: 371–377
Torriglia A, Perani P, Brossas JY, Chaudun E, Treton J, Courtois Y and Counis MF (1998) L-DNase II, a molecule that links proteases and endonucleases in apoptosis, derives from the ubiquitous serpin leukocyte elastase inhibitor. Mol. Cell. Biol. 18: 3612–3619
Kovacsovics M, Martinon F, Micheau O, Bodmer J, Hofmann K and Tschopp J (2002) Overexpression of helicard, a CARD-containing helicase cleaved during apoptosis, accelerates DNA degradation. Curr. Biol. 12: 1633
Chen J, Jin K, Chen M, Pei W, Kawaguchi K, Greenberg DA and Simon RP (1997) Early detection of DNA strand breaks in the brain after transient focal ischemia: implications for the role of DNA damage in apoptosis and neuronal cell death. J. Neurochem. 69: 232–245
Walker PR, LeBlanc J and Sikorska M (1997) Evidence that DNA fragmentation in apoptosis is initiated and propagated by single-strand breaks. Cell Death Differ. 4: 506–515
Liu QY, Ribecco-Lutkiewicz M, Carson C, Testolin L, Bergeron D, Kohwi-Shigematsu T, Walker PR and Sikorska M (2003) Mapping the initial DNA breaks in apoptotic Jurkat cells using ligation-mediated PCR. Cell Death Differ. 10: 278–289
Yakovlev AG, Wang G, Stoica BA, Boulares HA, Spoonde AY, Yoshihara K and Smulson ME (2000) A role of the Ca2+/Mg2+-dependent endonuclease in apoptosis and its inhibition by Poly(ADP-ribose) polymerase. J. Biol. Chem. 275: 21302–21308
Boulares AH, Zoltoski AJ, Contreras FJ, Yakovlev AG, Yoshihara K and Smulson ME (2002) Regulation of DNAS1L3 endonuclease activity by poly(ADP-ribosyl)ation during etoposide-induced apoptosis. Role of poly(ADP-ribose) polymerase-1 cleavage in endonuclease activation. J. Biol. Chem. 277: 372–378
de Murcia G and Menissier dM (1994) Poly(ADP-ribose) polymerase: a molecular nick-sensor. Trends Biochem. Sci. 19: 172–176
Jeggo PA (1998) DNA repair: PARP–another guardian angel? Curr. Biol. 8: R49–R51
Shall S and de Murcia G (2000) Poly(ADP-ribose) polymerase-1: what have we learned from the deficient mouse model? Mutat Res 460: 1–15
Masson M, Niedergang C, Schreiber V, Muller S, Menissier-de Murcia J and de Murcia G (1998) XRCC1 is specifically associated with poly(ADP-ribose) polymerase and negatively regulates its activity following DNA damage. Mol. Cell. Biol. 18: 3563–3571
Caldecott KW (2001) Mammalian DNA single-strand break repair: an X-ra(y)ted affair. Bioessays 23: 447–455
Honda K, Yamada T, Endo R, Ino Y, Gotoh M, Tsuda H, Yamada Y, Chiba H and Hirohashi S (1998) Actinin-4, a novel actin-bundling protein associated with cell motility and cancer invasion. J. Cell Biol. 140: 1383–1393
Pandey S, Walker PR and Sikorska M (1997) Identification of a novel 97?kDa endonuclease capable of internucleosomal DNA cleavage. Biochemistry 36: 711–720
Walker PR, LeBlanc J, Smith B, Pandey S and Sikorska M (1999) Detection of DNA fragmentation and endonucleases in apoptosis. Methods 17: 329–338
Takada F, Vander Woude DL, Tong HQ, Thompson TG, Watkins SC, Kunkel LM and Beggs AH (2001) Myozenin: an alpha-actinin- and gamma-filamin-binding protein of skeletal muscle Z lines. Proc. Natl. Acad. Sci. USA 98: 1595–1600
Nikolopoulos SN, Spengler BA, Kisselbach K, Evans AE, Biedler JL and Ross RA (2000) The human non-muscle alpha-actinin protein encoded by the ACTN4 gene suppresses tumorigenicity of human neuroblastoma cells. Oncogene 19: 380–386
Suhara T, Mano T, Oliveira BE and Walsh K (2001) Phosphatidylinositol 3-kinase/Akt signaling controls endothelial cell sensitivity to Fas-mediated apoptosis via regulation of FLICE-inhibitory protein (FLIP). Circ. Res. 89: 13–19
Bernstein C, Bernstein H, Payne CM and Garewal H (2002) DNA repair/pro-apoptotic dual-role proteins in five major DNA repair pathways: fail-safe protection against carcinogenesis. Mutat. Res. 511: 145–178
Trucco C, Oliver FJ, de Murcia G and Menissier-de Murcia J (1998) DNA repair defect in poly(ADP-ribose) polymerase-deficient cell lines. Nucleic Acids Res. 26: 2644–2649
Cote J and Ruiz-Carrillo A (1993) Primers for mitochondrial DNA replication generated by endonuclease G. Science 261: 765–769
Counis MF and Torriglia A (2000) DNases and apoptosis. Biochem. Cell Biol. 78: 405–414
Gavrieli Y, Sherman Y and Ben Sasson SA (1992) Identification of programmed cell death in situ via specific labeling of nuclear DNA fragmentation. J. Cell Biol. 119: 493–501
Acknowledgements
We thank Joanne Chartier and Min Wang for their technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by RA Lockshin
Rights and permissions
About this article
Cite this article
Liu, Q., Lei, J., LeBlanc, J. et al. Regulation of DNaseY activity by actinin-α4 during apoptosis. Cell Death Differ 11, 645–654 (2004). https://doi.org/10.1038/sj.cdd.4401401
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401401
Keywords
This article is cited by
-
Müller glial cells located in the peripheral retina are more susceptible to high pressure: implications for glaucoma
Cell & Bioscience (2024)
-
Gene model-related m6A expression levels predict the risk of preeclampsia
BMC Medical Genomics (2022)
-
Dysregulated expression of ACTN4 contributes to endothelial cell injury via the activation of the p38-MAPK/p53 apoptosis pathway in preeclampsia
Journal of Physiology and Biochemistry (2019)
-
Serum level of DNase1l3 in patients with dermatomyositis/polymyositis, systemic lupus erythematosus and rheumatoid arthritis, and its association with disease activity
Clinical and Experimental Medicine (2017)
-
A novel neuron-enriched protein SDIM1 is down regulated in Alzheimer's brains and attenuates cell death induced by DNAJB4 over-expression in neuro-progenitor cells
Molecular Neurodegeneration (2011)