Abstract
A total of 261 primary breast carcinomas were analyzed for amplification of the c-myc oncogene by fluorescence in situ hybridization performed on tumor tissue array samples. Results were compared with individual clinicopathologic and follow-up data. Thirty-eight (14.6%) of the tumors showed c-myc gene amplification (defined as two or more additional copies of c-myc gene in relation to the number of chromosome 8 centromere). The reproducibility of fluorescence in situ hybridization assay (defined by hybridization with two different myc probes) was good (kappa coefficient 0.402). Statistically significant associations were found between c-myc amplification and DNA aneuploidy (P = .0011), and progesterone receptor negativity (P = .0071), and c-myc amplification also tended to be associated with high histologic grade (P = .064), positive axillary nodal status (P = .080), and a high S-phase fraction (P = .052). c-myc amplification was not significantly associated with overall survival of patients with invasive cancer (P = .32). These data from a population-based tumor material suggest that c-myc amplification is a feature of aggressive breast cancers, but that it is unlikely to be a clinically useful prognostic factor.
Similar content being viewed by others
INTRODUCTION
Structural aberrations of chromosome 8 are common in breast cancer, as well as in various other solid tumors (1, 2). The copy number alterations of chromosome 8 detected by comparative genomic hybridization typically consist of gains on the q-arm, either entire 8q or only its telomeric parts (3, 4, 5). These aberrations have commonly been interpreted to reflect amplification of the c-myc oncogene, which is located at 8q24.1.
The central role of c-myc gene in physiologic proliferation and malignant transformation of human cells has been thoroughly described (6, 7, 8). c-myc has a role in most cellular functions, including replication, growth, metabolism, differentiation and apoptosis (9, 10, 11, 12, 13). In addition to these functions, the protein product of c-myc has been shown in cultured breast cancer cells to mimic the functions of the known breast cancer promoter, estrogen (8, 14). Many in vitro studies have also shown that expression of c-myc correlates positively to treatment with estrogens (15, 16), but negatively to the growth inhibitory effect of tamoxifen in vivo in estrogen receptor (ER)-positive breast tumors (17). Also the putative breast tumor suppressor gene BRCA1 has been shown to inhibit c-myc-mediated transcription and transformation (18). These findings implicate a specific role for c-myc gene in breast cancer pathogenesis.
Amplification of c-myc has been reported in breast cancer, as well as in other cancers in a large number of studies (19, 20, 21). Despite numerous studies the exact frequency of the amplification has remained obscure. The incidence of c-myc amplification ranges from 1 to 94% in different studies (22). Furthermore, unlike e.g., for the HER-2 oncogene amplification and its consequent protein overexpression, there is no clear consensus whether or not c-myc amplification is always associated with overexpression of its protein product (23). Wide variation has also been shown in the correlation between the c-myc oncogene amplification and clinicopathologic variables. These results may largely reflect the methodologic difficulties in detecting c-myc amplification when using Southern blotting. In particular, when evaluating low degree of amplification, the choice of the reference gene is critical. Fluorescence in situ hybridization (FISH), which is currently considered as the most accurate method to analyze gene amplification in human tumors, has been applied to c-myc only in a few studies including clinical breast tumor samples (20, 24, 25). In this study we applied the newly established tumor tissue array technology (26) and studied c-myc amplification by FISH using two different myc-specific probes. Our aim was to further clarify the role of c-myc amplification as a prognostic marker in breast cancer.
MATERIALS AND METHODS
Preparation of Tumor Tissue Arrays
A total of 351 archival paraffin blocks from primary invasive breast cancers were collected for this study. According to the data files of the Finnish Cancer Registry, these tumors represent 84.2% of all primary invasive cancers operated during 1991–1992 in the Tampere University Hospital district (population ∼400,000). The mean age of the patients was 61.6 years. The histologic types were available for 241 tumors including 198 (82%) ductal, 21 (9%) lobular, and 22 (9%) tumors of special histologic types. The previously collected clinicopathologic data (27) included patient age at the time of diagnosis, postoperative tumor size, axillary nodal status, and metastases, histologic grade, estrogen and progesterone receptor status (ER and PR, respectively), DNA ploidy, and S-phase fraction. The median follow-up time for the unrelapsed patients was 6.8 years (range 5.1 to 7.8 years), which enabled analysis of distant disease-free survival data 5 years from the diagnosis.
The tumors were routinely fixed (overnight in 10% buffered formalin) and processed into paraffin blocks according to the established protocols. Representative tumor regions were defined from hematoxylin and eosin-stained sections. Tumor tissue array blocks were made as follows: one tissue cylinder with a diameter of 0.6 mm was punched through selected tumor areas from each “donor” tissue block. Tissue cylinders were then inserted into “recipient” tissue array paraffin blocks using a specific custom-made instrument as described elsewhere (26). The 351 tumors formed a set of four tissue array blocks. Sections of 5 μm from the resulting multitumor tissue array blocks were then transferred to glass slides, and baked in a 60° C oven for 2 to 4 hours before the hybridization.
Fluorescence In Situ Hybridization
Locus-specific probes for chromosome 8 centromere (pJM128, ATCC, Rockville, MD) and c-myc gene (RMC08p001) were labeled with FITC (DuPont, Boston, MA) and digoxigenin, respectively, using nicktranslation. A second set of tissue array slides was hybridized with a commercial myc probe labeled with Spectrum Orange (LSI c-myc, Vysis, Inc, Downers Grove, IL), using the pJM128 as a reference probe.
Before hybridization, tumor array tissue sections were deparaffinized, pretreated in a microwave oven (10 minutes at 92° C in Tissue Pretreatment Buffer, Zymed Inc., South San Francisco, CA) and digested with proteinase K (0.25 mg/mL in 2 × standard saline citrate solution, SSC) for 20 minutes. Slides were rinsed with 2 × SSC, dehydrated with graded ethanols and air dried before hybridization. Ten μl of probe cocktail (probes for c-myc gene and the centromere of chromosome 8 with human placental DNA and Cot-1 DNA, Roche Biochemicals, Mannheim, Germany) was applied onto slides that were coverslipped and sealed with rubber cement. Denaturation was carried out at for 94° C for 3 minutes on a thermal plate. After an overnight hybridization at 37° C, the slides were stringency washed with two successive incubations in 0.5 × SSC and 4 × SSC. The digoxigenin-labeled myc probe was detected with anti-digoxigenin rhodamine (diluted 1:300). The slides were counterstained with 0.4 μm 4′,6-diamino-2-phenylindole in antifade solution (Vectashield, Vector Laboratories, Burlingame, CA).
The analysis was performed using an Olympus epifluorescence microscope (with 60 × objective), equipped with a CCD camera (Photometrics, Tucson, AZ). At least 50 nonoverlapping nuclei in every tumor sample were scored to determine the number of test and reference probe hybridization signals. The scoring results were expressed as the actual copy numbers per cell in the majority of the cells in each sample. Amplification was defined, when at least two copies more of c-myc were detected in relation to the chromosome 8 centromere copy number in at least in 20% of the analyzed cells. Samples were considered as having c-myc amplification if either or both c-myc probes showed amplification. Alternatively, we also tested the criteria used by Schraml et al. (25).
Statistical Methods
Contingency tables of c-myc gene status and clinicopathologic variables were analyzed with Fisher's exact test. The χ2 test for trend was used to compare c-myc with histologic grade. The analysis of disease-free survival was performed using Kaplan-Maier survival analysis and log-rank test. All P-values are two-tailed.
RESULTS
In FISH analysis, 261 out of 351 samples were interpretable on with either c-myc probe (success rate 74.4%; 66.1% with RMC08p001 and 53.0% with Vysis' probe). Reasons for unsuccessful analyses included tissue damage, and a weak hybridization signal and high background equally with both myc probes. To confirm the results, we used two different myc probes in separate hybridizations. The results obtained with the two probes were highly concordant (kappa coefficient 0.402).
Out of the 351 samples, 90 (25.6%) could not be analyzed. Only tumors that consisted of at least 50 nuclei were scored. However, the most common reason for uninterpretable results was weak hybridization, which is not related to the tissue array technology. According to our experience, weak hybridizations occur in 10 to 15% of hybridizations on ordinary tumor tissue sections as well. Tissue array technique itself induced few additional technical problems: the small samples are sometimes detached from the slides during the procedure, and the pretreatments needed by each sample vary greatly. The same pretreatment gives perfect signals on one sample, and another may be either totally damaged or totally intact showing no signals at all. In 64 cases the reason was the same with both probes used: 21 tissue samples (6.0%) were not representative for the tumor, and 32 samples (9.1%) were detached from the slide during the FISH procedure. Cell damage due to the strong treatments required by the FISH procedure was responsible for eight lost samples (2.3%), and in three cases (0.9%) no signals were seen. In 25 cases (7.1%) the reasons for a failure to analyze c-myc copy number status varied between the two sets of hybridization: missing samples (18 cases), cell damage (19 cases) and presence of no signals (15 cases).
Amplifications of c-myc by FISH
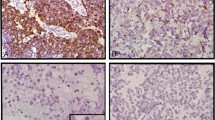
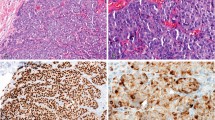
c-myc was amplified in 38 (14.6%) out of the 261 informative cases. The distribution of extra copy numbers varied from 2 (12 cases) to more than 10 (3 cases, Fig. 1, A–B), the median being 4 additional copies relative to chromosome 8 centromere. The proportion of cells showing the amplification varied from 20 to 100%.
Association of c-myc with the Clinicopathologic Data and Patient Survival
Association of c-myc amplification with the clinicopathologic data is shown in Table 1. The statistically most significant correlation was found in aneuploidy (P = .0011, odds ratio 4.8). The inverse association of c-myc amplification with PR expression was also very significant (P = .0071, odds ratio 0.34 PR-negativity being associated with amplifications. The S-phase fraction size tended to be associated with c-myc amplification with a P-value of .052 (odds ratio 2.4), and presence of axillary nodal metastases with a P-value of .080 (odds ratio 2.0). The other parameters investigated (age, tumor size, presence of metastases, ER expression, and distant disease-free survival) were not significantly associated with c-myc amplification.
DISCUSSION
In the present study, we used the newly described tumor tissue array technology to examine c-myc oncogene amplification as a prognostic biomarker in breast cancer. The tissue array technology was found powerful and fast to perform once the tedious preparation of the multitissue array blocks was completed. FISH of 261 tumors could be done on four microscope slides only. The success rate of our FISH assay was 74.4%, which is 10 to 15% units lower than in our similar assays on ordinary tumor sections. According to our experience, the main reason for weak hybridizations is the difficulty to find pretreatment protocols that could uncover the DNA sequences of single copy genes. In our experience, the most efficient pretreatment is the combination of microwave oven boiling with mild protease digestion, as shown in our recent study using chromogenic in situ hybridization of the HER-2 oncogene (28). The other source for inaccuracy was the tumor array technology itself. A small fraction of the tissue samples on array slides repeatedly detached from the slides during the FISH protocol, which includes protease treatment and incubations at high temperatures. In spite of these technical shortcomings and a failure to analyze c-myc expression in a part of the original series, we believe that the present series is highly representative of breast cancer in general, because we were able to collect tumor tissue from nearly all breast cancer patients diagnosed the disease within a well-defined geographical area and a defined time period.
The c-myc oncogene was found to be amplified in 14.6% of the primary breast tumors. In a recent meta-analysis (22), c-myc gene was amplified threefold or more in breast cancer biopsies in 1 to 94% of the cases, with an average of 15.5%. The large variation has been explained by methodologic differences and at selection bias of the tumors studied, especially when small numbers have been studied. Our patient cohort chosen to be analyzed is based on the data files of the Finnish Cancer Registry, which has a coverage close to 100%. The majority of the cancers were diagnosed at an early stage (63% of tumors had size less than 2 cm and negative axillary lymph nodes) as in many modern series where a relatively large proportion of the cases are detected in mammography. Because c-myc is known to be amplified more frequently in more advanced stage cancers, this may be one reason for our relatively low percentage of amplifications detected as compared with some other recent studies. The method we used, two-color fluorescence in situ hybridization, is currently regarded as the most specific and sensitive method to detect gene amplification in human tumor samples (29). In this method, the amplification of c-myc can readily be detected as an excess of gene copies as compared with the number of chromosome 8 centromeres, detected in the same hybridization with another fluorescent color. However, despite the straightforwardness of the FISH assay itself, the criteria for scoring gene amplification has remained more or less arbitrary. In addition to classical bridge-fusion-breakage type oncogene amplification, the presence of supernumerary copies of c-myc could be due to cytogenetic aberrations such as isochromosome formation, which are particularly common in chromosome 8 (1, 30). In these cases it remains uncertain whether or not the extra copies of c-myc lead to its functional oncogenic activation.
In general, c-myc amplification clearly correlated with adverse biological features of the tumors. DNA ploidy, low tumor histologic grade, presence of axillary nodal metastases, a high S-phase fraction size, and negative PR status tended to occur concomitantly with the amplification of c-myc. Statistically the most significant correlation was found with DNA aneuploidy of the tumor. These findings are in line with data suggesting that c-myc amplifications manifest in a later stage of tumor progression (22). The positive association with the S-phase fraction size and c-myc amplification fits with the wide experimental data supporting the central role of c-myc gene in DNA replication, and in tumor growth in general (6, 9, 11, 12, 13).
We found c-myc amplification to be more common in PR-negative tumors. This is also in concordance with the recent meta-analysis performed by Deming et al. (22), which suggested that PR-negativity is the only statistically significant association with c-myc amplification. For reasons not known, c-myc amplification is more strongly linked with progesterone rather than ER expression. The mechanisms by which progesterone affects differentiation, proliferation and other functions of breast tissue are complicated and still very poorly understood (31). The expression of c-myc mRNA is rapidly but transiently induced by progestin treatment, whereas relatively long-term treatment results in suppression of its expression (32, 33). The suppressive role of PR may explain at least partly the correlation between c-myc amplification and PR-negativity in breast tumors.
In our set of tumors, ER status was not significant in relation to c-myc amplification. According to the meta-analysis, there is great variability in the detected association between c-myc gene amplification and ER-status in different studies (22). The correlation between high estrogen levels and c-myc amplification and/or overexpression has nevertheless been found in in vitro and in vivo studies. For example, it has been shown that estrogen treatment induces the expression of c-myc mRNA directly (15, 16). Furthermore, ER-positive breast tumors excised from patients who have received tamoxifen treatment show a decreased level of c-myc mRNA as compared tumors from patients who have not been treated with tamoxifen (17). Because c-myc gene does not contain the canonical estrogen-responsive-element, the transcriptional activation of c-myc is induced through indirect mechanisms, possibly via estrogen-ER signaling.
c-Myc protein is also able to mimic estrogen inducing cyclinE/cdk2 activity in breast cancer cells in culture (14), and it induces also directly expression of cyclin E (8). Despite these data, it still remains unknown how estrogen-ER signaling regulates c-myc expression in human breast cancer, because several reports show that overexpression and/or amplification of c-myc occurs preferentially in ER-negative tumors.
These data from a population-based tumor material suggest that c-myc amplification is a feature of aggressive breast cancers, but that it is unlikely to be a clinically useful prognostic factor. The low percentage of c-myc amplification seen implicates that there may be other unknown mechanisms that connect c-myc with the breast cancer pathogenesis. It would be interesting to resolve other ways of c-myc oncogene activation, e.g., specific translocations. c-myc oncogene is able to direct cells to different, partly opposite directions; proliferation, differentiation or apoptosis in physiologic situations, and the tumor formatting ability of the different isoforms of c-Myc protein product may also be of future interest. This challenges us to study further the possible mediating proteins and coexisting oncogenic factors required for the tumorigenic functions c-myc.
References
Rummukainen JK, Kytölä S, Karhu R, Farnebo F, Larsson C, Isola J . Aberrations of chromosome 8 in sixteen breast cancer cell lines by comparative genomic hybridization, fluorescence in situ hybridization, and spectral karyotyping. Cancer Genet Cytogenet 2001; 126: 1–7.
Forozan F, Karhu R, Kononen J, Kallioniemi A, Kallioniemi O-P . Genome screening by comparative genomic hybridization. Trends Genet 1997; 13: 405–409.
Isola JJ, Kallioniemi O-P, Chu LW, Fuqua SAW, Hilsenbeck SG, Osborne CK, et al. Genetic aberrations detected by comparative genomic hybridization predict outcome in node-negative breast cancer. Am J Pathol 1995; 147: 905–911.
Nishizaki T, DeVries S, Chew K, Goodson WH III, Ljung BM, Thor A, et al. Genetic alterations in primary breast cancers and their metastases: direct comparison using modified comparative genomic hybridization. Genes Chromosom Cancer 1997; 19: 267–272.
Tirkkonen M, Tanner M, Karhu R, Kallioniemi A, Isola J, Kallioniemi O-P . Molecular cytogenetics of primary breast cancer by CGH. Genes Chromosom Cancer 1998; 21: 177–184.
Dang CV . c-Myc target genes involved in cell growth, apoptosis, and metabolism. Mol Cell Biol 1999; 19: 1–11.
Bouchard C, Staller P, Eilers M . Control of cell proliferation by Myc. Trends Cell Biol 1998; 8: 202–206.
Amati B, Alevizopoulos K, Vlach J . Myc and the cell cycle. Front Biosci 1998; 3: D250–D268.
Hoffman B, Liebermann DA . The proto-oncogene c-myc and apoptosis. Oncogene 1998; 17: 3351–3357.
Packham G, Cleveland JL . c-Myc and apoptosis. Biochim Biophys Acta, 1995; 1242: 11–28.
Dang CV, Resar LMS, Emison E, Kim S, Li Q, Prescott JE, et al. Function of the c-Myc oncogenic transcription factor. Exp Cell Res 1999; 253: 63–77.
Elend M, Eilers M . Cell growth: downstream of Myc—to grow or to cycle? Curr Biol 1999; 9: R936–R938.
Prendergast GC . Mechanisms of apoptosis by c-Myc. Oncogene 1999; 18: 2967–2987.
Prall OWJ, Rogan EM, Musgrove EA, Watts CKW, Sutherland RL . c-Myc or cyclin D1 mimics estrogen effects on cyclin E-Cdk2 activation and cell cycle reentry. Mol Cell Biol 1998; 18: 4499–4508.
Shiu RP, Watson PH, Dubik D . c-myc oncogene expression in estrogen-dependent and -independent breast cancer. Clin Chem 1993; 39: 353–355.
Hyder SM, Stancel GM, Loose-Mitchell DS . Steroid hormone-induced expression of oncogene encoded nuclear proteins. Crit Rev Eukaryot Gene Expr 1994; 4: 55–116.
Le Roy X, Escot C, Brouillet JP, Theillet C, Maudelonde T, Simony-Lafontaine J, et al. Decrease of c-erbB-2 and c-myc RNA levels in tamoxifen-treated breast cancer. Oncogene 1991; 6: 431–437.
Wang Q, Zhang H, Kajino K, Greene MI . BRCA1 binds c-Myc and inhibits its transcriptional and transforming activity in cells. Oncogene 1998; 17: 1939–1948.
Escot C, Theillet C, Lidereau R, Spyratos F, Champeme MH, Gest J, et al. Genetic alteration of the c-myc protooncogene (MYC) in human primary breast carcinomas. Proc Natl Acad Sci USA 1986; 83: 4834–4838.
Visscher DW, Wallis T, Awussah S, Mohamed A, Crissman JD . Evaluation of MYC and chromosome 8 copy number in breast carcinoma by interphase cytogenetics. Genes Chromosom Cancer 1997; 18: 1–7.
Nesbit CE, Tersak JM, Prochownik EV . MYC oncogenes and human neoplastic disease. Oncogene 1999; 18: 3004–3016.
Deleted in proof.
Deming SL, Nass SJ, Dickson RB, Trock BJ . C-myc amplification in breast cancer: a meta-analysis of its occurrence and prognostic relevance. Br J Cancer 2000; 83: 1688–1695.
Biéche I, Lidereau R . Genetic alterations in breast cancer. Genes Chromosom Cancer 1995; 14: 227–251.
Persons DL, Borelli KA, Hsu PH . Quantitation of HER-2/neu and c-myc gene amplification in breast carcinoma using fluorescence in situ hybridization. Mod Pathol 1997; 10: 720–727.
Schraml P, Kononen J, Bubendorf L, Moch H, Bissig H, Nocito A, et al. Tissue microarrays for gene amplification surveys in many different tumor types. Clin Cancer Res 1999; 5: 1966–1975.
Kononen J, Bubendorf L, Kallioniemi A, Bärlund M, Schraml P, Leighton S, et al. Tissue microarrays for high-throughput molecular profiling of tumor specimens. Nat Med 1998; 4: 844–847.
Lundin J, Lundin M, Holli K, Kataja V, Elomaa L, Pylkkanen L, et al. Omission of histologic grading from clinical decision making may result in overuse of adjuvant therapies in breast cancer: results from a nationwide study. J Clin Oncol 2001; 19: 28–36.
Tanner M, Gancberg D, Di Leo A, Larsimont D, Rouas G, Piccart MJ, et al. Chromogenic in situ hybridization: a practical alternative for fluorescence in situ hybridi-zation to detect HER-2/neu oncogene amplification in archival breast cancer samples. Am J Pathol 2000; 157: 1467–1472.
Kytölä S, Rummukainen J, Nordgren A, Karhu R, Farnebo F, Isola J, et al. Chromosomal alterations in 15 breast cancer cell lines by comparative genomic hybridization and spectral karyotyping. Genes Chromosom Cancer 2000; 28: 308–317.
Pauletti G, Godolphin W, Press MF, Slamon DJ . Detection and quantitation of HER-2/neu gene amplification in human breast cancer archival material using fluorescence in situ hybridization. Oncogene 1996; 13: 63–72.
Clarke CL, Sutherland RL . Progestin regulation of cellular proliferation. Endocr Rev 1990; 11: 266–301.
Musgrove EA, Lee CS, Sutherland RL . Progestins both stimulate and inhibit breast cancer cell cycle progression while increasing expression of transforming growth factor alpha, epidermal growth factor receptor, c-fos, and c-myc genes. Mol Cell Biol 1991; 11: 5032–5043.
Wong MS, Murphy LC . Differential regulation of c-myc by progestins and antiestrogens in T-47D human breast cancer cells. J Steroid Biochem Mol Biol 1991; 39: 39–44.
Acknowledgements
The authors thank Mrs. Sari Toivola, Ms. Päivi Järvinen, and Mrs. Anne Luuri for their skillful technical assistance. This work was supported by the Tampere Hospital Foundation, and the Foundation for Finnish Cancer Society.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rummukainen, J., Salminen, T., Lundin, J. et al. Amplification of c-myc by Fluorescence In Situ Hybridization in a Population-Based Breast Cancer Tissue Array. Mod Pathol 14, 1030–1035 (2001). https://doi.org/10.1038/modpathol.3880431
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3880431
Keywords
This article is cited by
-
C-myc as a predictive marker for chemotherapy in metastatic breast cancer
Clinical and Experimental Medicine (2012)
-
Insulin-like growth factor-1 receptor gene expression is associated with survival in breast cancer: a comprehensive analysis of gene copy number, mRNA and protein expression
Breast Cancer Research and Treatment (2011)
-
The Ras oncogene signals centrosome amplification in mammary epithelial cells through cyclin D1/Cdk4 and Nek2
Oncogene (2010)
-
Gene amplification in ductal carcinoma in situ of the breast
Breast Cancer Research and Treatment (2010)
-
Frequency, prognostic impact, and subtype association of 8p12, 8q24, 11q13, 12p13, 17q12, and 20q13 amplifications in breast cancers
BMC Cancer (2006)