Abstract
Mutation of p53 is rare in localized prostate carcinoma. The oncoprotein MDM2, whose gene has a response element for p53, promotes the degradation of p53 protein and inhibits its transcriptional activation of genes related to cell cycle arrest and apoptosis, constituting a negative feedback control. We studied p53 and MDM2 expression by immunohistochemistry and looked for mutations in p53 exons 5 to 8 by polymerase chain reaction-single strand conformational polymorphism in 118 patients submitted to radical prostatectomy for localized prostate cancer. In 28 cases, we studied cell proliferation by immunohistochemistry, using antibody for Ki-67, and apoptosis by the deoxynucleotidyl transferase mediated dUTP biotin nick end labeling technique. Although no p53 mutations were found, p53 protein was detected in 31.4% of the cases, and these cases had higher Gleason scores (P = .03) and more advanced tumor stages (P = .02). MDM2 was overexpressed in 40.7% of the cases, and these cases had greater tumor volumes (P = .001). Tumors that were positive for both p53 and MDM2 were larger (P = .003) and of more advanced stage (P = .03). Within the 28-case subset, the proliferative index was higher among MDM2-positive tumors (P = .046), and the apoptotic index was lower among p53-positive tumors (P = .01). We conclude that, although p53 mutation is a rare event in prostate carcinogenesis, the detection of p53 protein by immunohistochemistry is common and is associated with decreased apoptosis and increased histologic grade and tumor stage. We also conclude that the overexpression of MDM2 has a role in prostate carcinogenesis, being frequently detected and associated with increased cell proliferation and tumor volume. Finally, we propose that the MDM2-positive/p53-positive phenotype identifies prostate cancers with aggressive behavior.
Similar content being viewed by others
INTRODUCTION
The p53 protein, which activates genes involved in cell cycle arrest (1, 2) and apoptosis (3), is missing or inactive in more than 50% of human tumors as a result of point mutations or deletions (4, 5) or interactions with proteins (6, 7, 8). One such protein is MDM2, a cellular oncoprotein encoded by a gene located on chromosome 12q13-14 (9). MDM2 binds to p53, promoting its degradation via ubiquitin, masking its transactivation domain, and inhibiting its transcriptional activation of genes related to cell cycle arrest and apoptosis (9). The transcription of MDM2 is promoted by p53 (8, 9), creating a negative feedback control of p53 by MDM2 (9).
Amplification of the MDM2 gene or overexpression of the MDM2 protein have been implicated in the development of tumors (8, 10), and MDM2 overexpression has been related to more aggressive disease and poorer survival. Amplification of MDM2 has been found in 35% of soft tissue sarcomas (11), one third of malignant fibrous histiocytomas (12), 42% of liposarcomas (13), and 7% of osteosarcomas (14). Overexpression of MDM2 is common in lymphoblastic leukemia (15) and bronchogenic carcinoma (16), and has been related to more aggressive disease or poorer survival in these diseases as well as in lymphoma (17) and bladder cancer (18). A recent study showed MDM2 overexpression in all cases of salivary gland carcinomas, where p53 mutations are rare (19).
p53 mutations are also rare in prostate cancer. They occur only late in the development of the disease and have been related to metastasis and androgen-independent growth (20, 21, 22). In one series, p53 mutations were described in 4.4% of cases of localized adenocarcinoma, but in 45% of cases of bone marrow metastasis (20). However, even in the absence of p53 mutations, it could be that the function of p53 is abrogated by interactions with an oncogenic protein such as MDM2. The half-life of p53 is short, and it is not normally detected by immunohistochemistry (IHC), but p53 that has been inactivated by missense mutation or, sometimes, by interaction with other proteins is abnormally stable and, therefore, detectable (9, 23).
The aim of this study was to evaluate the roles played by p53 mutation, p53 expression, and MDM2 overexpression in 118 cases of localized prostatic adenocarcinoma. Mutations of p53 were detected by polymerase chain reaction-single strand conformational polymorphism analysis of exons 5, 6, 7, and 8 (24), and p53 and MDM2 were detected by IHC. The results were correlated with the stage, grade, and volume of the tumors. To further evaluate the roles of p53 and MDM2, cell proliferation and apoptosis were studied in a subset of 28 cases selected so that both normal and abnormal expression of p53 and MDM2 would be represented. A proliferative index was determined by IHC, using antibody to Ki-67, a protein that is expressed only by dividing cells (25). An apoptotic index was determined by the deoxynucleotidyl transferase mediated dUTP biotin nick end labeling technique (26).
MATERIALS AND METHODS
Case Selection and Pathologic Examination
One hundred eighteen patients with prostate cancer, diagnosed by biopsy and clinically staged T1c-T2, were submitted to radical prostatectomy between August 1997 and June 1998 at the Hospital Sirio Libanes of Sao Paulo, Brazil. The surgical specimens were fixed in 10% buffered formalin for no longer than 4 hours. The entire surgical margin was stained with India ink, the left and right lobes were separated, 3-mm transverse serial sections were taken from each lobe, and the entire gland was submitted for histologic examination. Sections of the bladder neck, prostatic apex, seminal vesicles, and pelvic lymph nodes were also submitted. From 16 to 20 slides of each gland were examined. The Gleason score was used for histologic grading (27). The tumor volume was evaluated as described by Humphrey et al. (28). Briefly, a grid was placed below the slides, on which the area involved by the tumor had been previously sketched out. The percentage of tumor on a slide was determined by dividing the number of squares involved by tumor by the number of squares occupied by the whole section on the slide. Tumor volume was defined as the mean percentage of tumor in the prostate gland (the percentage of tumor on each slide divided by the number of slides from the prostate gland). Extraprostatic involvement was defined as tumor infiltration of the adipose tissue, the neurovascular plexus, or the parenchyma of the seminal vesicles. The TNM system was used for tumor staging (29).
p53 Mutation Analysis
DNA Extraction
DNA was extracted from three 10-μm sections of the paraffin block that best represented each tumor (previously selected from hematoxylin- and eosin-stained slides). Disposable microtome blades were used, and instruments were cleaned with xylene after each tissue section to avoid cross-contamination. Three baths in 500 μL of xylene at 95°C followed by three baths in 500 μL of 99% ethanol were used to dewax the tissue. Next, the tissue samples were centrifuged for 5 minutes at 13,000 rpm and 4°C. The resulting pellets were incubated overnight at 37°C in 500 μL of digestion buffer (10 mM Tris [pH 7.4], 100 mM NaCl, 25 mM disodium EDTA, 0.5% sodium dodecyl sulfate) containing 10 μL of proteinase K (200μg/mL final concentration). The enzyme was inactivated by heating the specimens for 10 minutes at 95°C, protein was removed by adding an equal volume of phenol:chloroform:isoamyl alcohol (25:24:1), and nucleic acid was precipitated from the aqueous phase by adding ¼ volume 8 M ammonium acetate and an equal volume of isopropanol. The nucleic acid samples were then incubated at −20°C for 1 hour and centrifuged for 10 minutes at 13,000 rpm and 4°C. The resulting pellets were washed in 70% ethanol to remove salt, air dried, and resuspended in 50 μL of TE/RNAse (30).
Polymerase Chain Reaction Amplification
The polymerase chain reaction (PCR) amplification and nonisotopic detection of exons 5 to 8 of the p53 gene were described by Soong et al. (31) and will be summarized here. The PCR reactions were carried out in 25 μL of 1 x Reaction Buffer containing 0.2 mM deoxynucleotide triphosphates, 2.5 mM magnesium chloride, each primer at a concentration of 0.4 μM, 0.25 units of Taq polymerase, and 1 μL of extracted DNA. Thirty-five thermal cycles of 30 seconds at 94°C, 1 minute at 60°C, and 1 minute at 72°C were completed, and the cycling was concluded by a 10-minute extension at 72°C, all in a thermocycle Gene Amp PCR System 2400 (Perkin Elmer, Foster City, CA). All procedures to prevent contamination were followed.
SSCP Screening for p53 Mutations
Single-stranded DNA for SSCP analysis was produced by combining equal 5-μL volumes of PCR product and formamide loading buffer (95% formamide, 10 mM EDTA, 0.05% bromophenol blue, 0.05% xylene cyanol) and heating at 95°C for 10 minutes. The DNA was subjected to electrophoresis at 200 V for 2 hours at 4°C within a Mighty-small apparatus (Pharmacia, Uppsala, Sweden) containing a 15% polyacrylamide gel with 5% glycerol. Silver staining was done by the method of Bassam et al. (32). The gels were soaked for 5 minutes in 10% ethanol, 5 minutes in 1% nitric acid, and 10 minutes in an impregnating solution (0.1 g of silver nitrate and 150 μL of formaldehyde in 100 mL water); then soaked in a developing solution (3 g of sodium carbonate, 150 μL of formaldehyde, and 100 μL of sodium sulfate in 100 mL water) until bands were visible; and then fixed in 10% acetic acid for 5 minutes. The gels were analyzed under white light in a computerized imaging system (Gel Doc 1000, Biorad, North Ryde, Australia) and compared with samples of wild-type p53 from normal tissue extracted and amplified by the same methods to detect p53 mutations, which would appear as extra bands or mobility shifts.
Immunohistochemical Analysis
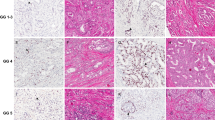
Three-micrometer sections from the paraffin block that best represented the tumor (the same block used for PCR) were placed on adhesive-coated slides. In a heat antigen retrieval process (33) the slides were placed in a citrate buffer (1 mM, pH 6.0) and heated three times for 8 minutes each time in a domestic microwave oven at high power. The slides were incubated overnight at 4°C with monoclonal antibodies to p53 (clone DO7, Dako, Glostrup, Denmark), MDM2 (clone 1B10, Novocastra, Newcastle upon Tyne, UK), or Ki-67 (clone MIB1, Immunotech, Marseille, France) at respective dilutions of 1:50, 1:50, and 1:100 in phosphate-buffered saline (PBS). Biotinylated antimouse immunoglobulin G was applied at 1:200 dilution for 60 minutes at room temperature. The slides were rinsed with PBS for 30 minutes, incubated with peroxidase-conjugated streptavidin (streptABC Kit, Dako) at 1:400 dilution in PBS for 45 minutes at room temperature, and then rinsed again with PBS for 30 minutes. Color was developed by incubating the slides in 0.06% diaminobenzidine in PBS for 15 minutes, and the slides were then rinsed in tap water, counterstained with Harris hematoxylin, dehydrated, coverslipped, and reviewed under a light microscope (34). For p53 and MDM2, any dark, brown nuclear staining was considered positive, indicating abnormal stabilization of p53 or overexpression of MDM2 (Figs. 1 and 2, respectively). For Ki-67, at least 500 cells were counted, and the percentage of cells with dark, brown nuclear staining (Fig. 3) was considered the proliferative index.
Detection of Apoptosis
The apoptotic index was assessed by the deoxynucleotidyl transferase mediated dUTP biotin nick end labeling technique using the TDT-FragEL kit (Oncogene, Cambridge, MA), as described by Gavrieli et al. (26). Briefly, 3-μm sections of the paraffin block that best represented the tumor (the same block used for PCR and IHC), were placed on adhesive-coated slides, deparaffinized in xylene, and rehydrated in alcohol. After cell membranes were made permeable by treating the slides with proteinase K diluted 1:10 in 10 mM Tris, pH 8.0, for 20 minutes at room temperature, endogenous peroxidase was inactivated by applying 3% H2O2 for 5 minutes at room temperature. Labeling was performed by incubating the slides in a humidified chamber at 37°C for 1.5 hours with the terminal deoxynucleotidyl transferase reaction mix from the kit, which contains labeled and unlabeled deoxynucleotides as well as terminal deoxynucleotidyl transferase enzyme. The reaction was stopped by applying 0.5 M EDTA, pH 8.0, for 5 minutes at 37°C and blocked by applying 4% bovine serum albumin in PBS for 10 minutes at room temperature. The peroxidase-conjugated streptavidin, diluted 1:50 in blocking buffer, was applied for 30 minutes at room temperature in a humidified chamber. The slides were then incubated with diaminobenzidine, counterstained with methyl green, and coverslipped. At least 200 cells were counted, and the percentage of stained cells (Fig. 4) was considered to be the apoptotic index.
Statistical Analysis
To verify significant relationships between variables, the non-parametric Mann-Whitney and Kruskal-Wallis tests were used for quantitative variables, and the χ2 test was used for qualitative variables. P values were two-sided and were considered significant when less than 0.05. The tests were performed with the computer program SPSS, version 8 (SPSS, Inc., Chicago, IL).
RESULTS
Patient and Tumor Characteristics
The median age of the patients was 63 years (range, 46 to 75 yr). The median Gleason score was 7 (range, 4 to 10) in the entire study group of 118 patients, and 8 (range, 6 to 10) in the 28-patient subset for the apoptosis and proliferation studies. The median tumor volume was 22% (range, 1 to 100; mean, 25.0; SD, 19.4) in the entire group and 26% (range, 1 to 90; mean, 33.2; SD, 24.4) in the subset for apoptosis and proliferation studies. In the entire study group, the pathologically determined tumor stage was pT2N0 in 69 cases (58.5%), pT3N0 in 44 cases (37.3%), and pT3N1 in five cases (4.2%). In the subset for apoptosis and proliferation studies, the stage was pT2N0 in 17 cases (61%), pT3N0 in nine cases (32%), and pT3N1 in two cases (7%).
p53 Mutation
No mutations were detected; electrophoresis of the amplified products of exons 5, 6, 7, and 8 of the p53 gene revealed no abnormalities of band migration in any of the 118 cases studied.
p53 Expression and Grade, Volume, and Stage of Tumors
Thirty-seven cases (31.4%) were positive for p53. Among the positive cases, brown staining was present in a median of 18% of cells. Characteristics of the p53-positive and p53-negative tumors are compared in Table 1. Positivity for p53 was related to the histologic grade of the tumor, with higher Gleason scores among the p53-positive cases than among the p53-negative cases. A relationship to tumor stage also was demonstrated: 23% (16/69) of the stage pT2 tumors were p53-positive, while 43% (21/49) of the stage pT3 tumors were p53-positive. No statistically significant relationship was demonstrated between p53 and tumor volume.
MDM2 Expression and Grade, Volume, and Stage of Tumors
MDM2 was overexpressed in 48 patients (40.7%). In most cases, only the nuclei of tumor cells were stained; cytoplasmic staining occurred rarely, mainly in Gleason 4 or 5 patterns. There was no background staining, and staining was often focal in the tumor. Characteristics of the MDM2-positive and MDM2-negative tumors are compared in Table 1. Overexpression of MDM2 was related to tumor volume: the median volume was 26% for MDM2-positive tumors and 17% for MDM2-negative tumors. A relationship of borderline significance was observed between MDM2 and tumor stage: 33% (23/69) of the stage pT2 tumors were MDM2-positive, while 51% (25/49) of the stage pT3 tumors were MDM2-positive. There was no relationship between MDM2 and the Gleason score.
Interaction of p53 and MDM2 Effects on Grade, Volume, and Stage of Tumors
When the cases were grouped according to their expression of both MDM2 and p53 (Table 1), there were statistically significant differences among the four groups with respect to tumor volume and tumor stage. The median tumor volume was 38% for tumors that were positive for both MDM2 and p53, but only 17% for tumors that were negative for both proteins. Extraprostatic invasion (stage pT3) was present in 62% (10/16) of tumors that were positive for both p53 and MDM2, but only 27% (13/49) of tumors that were negative for both proteins. Lymph node status did not appear to be related to p53 or MDM2 expression. There was no statistically significant variation in the Gleason score among the four groups, although the median Gleason score was 8 for tumors that were positive for both p53 and MDM2 and 6 for those that were negative for both proteins.
Apoptosis, Cell Proliferation, and p53 or MDM2 Expression
In the subset of 28 cases in which apoptosis and cell proliferation were studied, the median apoptotic index was 3.4% (range, 0.5 to 9.7; mean, 3.8; SD, 2.5), and the median proliferative index was 8% (range, 0.5 to 23.9; mean, 9.1; SD, 6.2). Table 2 compares the apoptotic and proliferative indexes for tumors grouped according to their expression of p53 and MDM2. Positivity for p53 was negatively associated with the apoptotic index. There was no statistically significant relationship between p53 and cell proliferation. Overexpression of MDM2 was positively associated with the proliferative index, but there was no statistically significant relationship between MDM2 and the apoptotic index.
Interaction of p53 and MDM2 Effects on Apoptosis and Cell Proliferation
When the cases were grouped according to their expression of both MDM2 and p53 (Table 2), there was statistically significant variation among the four groups in the apoptotic index, with lower indexes occurring in the two p53-positive groups, and the highest median apoptotic index in the p53-negative/MDM2-positive group. There was no statistically significant variation in proliferative index among the groups, although the median proliferative index was higher in the two MDM2-positive groups, with the highest median index in the group that was positive for both p53 and MDM2.
DISCUSSION
Although we detected no p53 mutations in our 118 cases of localized prostate carcinoma, IHC indicated that p53 was abnormally stabilized in 31% of the tumors. Among these tumors, the histologic grade was higher (P = .03), the tumor stage was more advanced (P = .02), and the apoptotic index was lower (P = .01) than among the p53-negative tumors. Overexpression of MDM2 was detected in 41% of the tumors and was positively related to the tumor volume (P = .001) and the proliferative index (P = .046).
Although the frequency of p53 detection was somewhat higher in our study than in previous reports, in which p53 was detected in 5% to 17% of prostate cancers, our results agreed with these reports that p53 expression is positively related to tumor stage and tumor grade (20, 22). This relationship of p53 to tumor grade was dramatic in another series, in which p53 was expressed in only 4.5% of well-differentiated prostatic carcinomas, but in 80% of poorly differentiated tumors (35). The frequency of MDM2 overexpression in our study was similar to the frequency in other reports that describe MDM2 overexpression in almost one third to almost one half of prostate cancers (36, 37). In a recent report (37), MDM2 overexpression was related to tumor stage, a relationship that was of marginal significance (P = .054) in our study.
Our results confirmed other reports that p53 mutations are rare in localized prostate cancer. In several studies, p53 mutations were found in only 2% to 6% of cases (20, 21, 38). A possible explanation for this low frequency of mutation is that in our study and most of the other studies with low p53 mutation rates (20, 38, 39), only exons 5 through 8 were examined. When primers to amplify the region extending from exon 4 through exon 11 were used to study a series of 48 cases of localized and metastatic prostate cancer (40), p53 mutations were found in 77% of the cases, suggesting that exons 4, 9, 10, and 11 ought to be evaluated in addition to exons 5 through 8.
To interpret our results, it is important to realize the situations that can cause a tumor to be p53-positive or p53-negative by IHC. As mentioned before, p53 normally has too short a half-life to be detected by IHC (24). Missense mutations are the most common p53 mutations, and because they stabilize the p53 protein, they are detected by IHC (24). Nonsense or frame-shift mutations, gene deletions, and mutations that cause truncation of the protein may not be detected by IHC. The IHC method can also detect p53 that has been temporarily stabilized by some mechanism other than mutation (41), such as the action of the product of the p19arf gene (42) or the formation of complexes with proteins such as MDM2 (9, 11). Stabilized p53 that is detected by IHC method is inactive and does not function normally in the control of the cell cycle and apoptosis induction (24).
MDM2-positivity by IHC, on the other hand, signifies that a tumor overexpresses MDM2, whether as a result of increased transcription (43), enhanced translation (44), or gene amplification. Amplification is the least common of these mechanisms (43, 45); in one prostate carcinoma series (46), no cases of MDM2 gene amplification were found.
The relationship we found between p53 and apoptosis appeared to be partially independent of MDM2 expression, suggesting that the ability of p53 to induce apoptosis cannot always be abrogated by MDM2.
The relationship we found between MDM2 overexpression and cell proliferation was interesting. It may explain the relationship between MDM2 overexpression and tumor volume, and it indicates to us that the overexpression of MDM2 has a role in prostate carcinogenesis.
Because MDM2 overexpression had a significant relationship to the proliferative index while p53 positivity did not, it would seem that MDM2 can affect cell proliferation not only by interfering with the p53’s control of the cell cycle, but also by some other mechanism. The cell cycle is negatively controlled by the Rb protein as well as by the p53 protein, and MDM2 may interfere with that control by interacting with the Rb protein directly (47) or by inhibiting the ability of transforming growth factor β to maintain the Rb protein in its active form (48). Also, a RING finger domain suggests that MDM2 could interact with rRNA, suggesting possible roles for MDM2 in translation control, RNA transport, or ribosome biogenesis (8).
Whatever the additional mechanism of MDM2 control over the cell cycle may be, it may add to the effect of p53 inactivation. In the four-group analysis, the group in which MDM2 and p53 were positive had a higher median proliferative index than the other groups, although we were not able to show this difference to be statistically significant. This group also had greater tumor volumes (P = .003) and more advanced tumor stages (P = .03) than the other groups. It may be that the MDM2-positive/p53-positive phenotype identifies an aggressive subset of prostate adenocarcinoma. It has been suggested that the results of IHC investigation for both proteins, rather than just one, would better reflect the behavior of tumors (49), and we believe that assessment of both p53 and MDM2 by IHC should be part of the evaluation of prostate cancer.
References
Kastan M, Zhan Q, El-Deiry W, Carrier F, Jacks T, Walsh W, et al. A mammalian cell cycle checkpoint pathway utilizing p53 and GADD45 is defective in ataxia-telangectasia. Cell 1992; 71: 587–597.
El-Deiry W, Harper J, O’Connor P, Velculescu V, Canman C, Jackman J, et al. WAF1/CIP1 is induced in p53-mediated G1 arrest and apoptosis. Cancer Res 1994; 54: 1169–1174.
Evan G, Littlewood T . A matter of life and death. Science 1998; 281: 1317–1322.
Greenblatt M, Bennett W, Hollstein M, Harris C . Mutations in p53 tumor suppressor gene: clues to cancer etiology and molecular pathogenesis. Cancer Res 1994; 54: 4855–4878.
Hollstein M, Shomer B, Greenblat M, Soussi T, Hovig E, Montesano R, et al. Somatic point mutations in the p53 gene of human tumors and cell lines: update compilation. Nucleic Acids Res 1996; 24: 141–146.
Scheffner MB, Werness BA, Huibregtse JM, Levine AJ, Howley PM . The E6 oncoprotein encoded by human papilloma virus types 16 and 18 promotes the degradation of p53. Cell 1990; 63: 1129–1136.
Querido E, Marcellus RC, Lai A, Charbonneau R, Teodor JG, Ketner G, et al. Regulation of p53 levels by the E1B 55kD protein and E4 or E6 in adenovirus-infected cells. J Virol 1997; 71: 3788–3798.
Freedman DA, Wu L, Levine AJ . Functions of the MDM2 oncoprotein. Cell Mol Life Sci 1999; 55: 96–107.
Momand J, Zambetti GP . MDM2: big brother of p53. J Cell Biochem 1997; 64: 343–352.
Farkhazadeh SS, Trusko SP, George DL . Tumorigenic potential associated with enhanced expression of a gene that is amplified in a mouse tumor cell line. EMBO J 1991; 10: 1565–1569.
Oliner JE, Kinzler KW, Meltzer PS, George DL, Vogelstein B . Amplification of a gene encoding a p53-associated protein in human sarcomas. Nature 1992; 358: 80–83.
Reid AH, Tsai MM, Venzon DJ, Wright CF, Lack EE, O’Leary TJ . MDM2 amplification, p53 mutation, and accumulation of the p53 gene product in malignant fibrous histiocytoma. Diagn Mol Pathol 1996; 5: 65–73.
Scheider-Stock T, Walter H, Radig K, Rys J, Bosse A, Kuhnen C, et al. MDM2 amplification and loss of heterozygosity at Rb and p53 genes: no simultaneous alterations in the oncogenesis of liposarcomas. J Cancer Res Clin Oncol 1998; 124: 532–540.
Lonardo F, Ueda T, Huvos AG, Healey J, Ladanui M . p53 and MDM2 alterations in osteosarcomas—correlation with clinicopathologic features and proliferative rate. Cancer 1997; 79: 1541–1547.
Gustafsson B, Stal O, Gustaffson B . Overexpression of MDM2 in acute childhood lymphoblastic leukemia. Pediatr Hematol Oncol 1998; 15: 519–526.
Gorgooulis VG, Zoumpourlis V, Rassidakis GZ, Karameris A, Rassidakis AN, Spandidos DA, et al. A molecular and immunohistochemical study of the MDM2 protein isoforms and p53 gene product in bronchogenic carcinoma. J Pathol 1996; 180: 129–137.
Moeller MB, Nielsen O, Pedersen NT . Oncoprotein MDM2 overexpression is associated with poor prognosis in distinct non-Hodgkin’s lymphoma entities. Mod Pathol 1999; 12: 1010–1016.
Özdemir E, Kakehi Y, Okuno H, Habuchi T, Okada Y, Yoshida O . Strong correlation of basement membrane degradation with p53 inactivation and/or MDM2 overexpression in superficial urothelial carcinomas. J Urol 1997; 158: 206–211.
Araujo VC, Martins MT, Leite KRM, Gomez RS, Araujo NS . Immunohistochemical MDM2 expression in minor salivary gland tumours and its relationship to p53 gene status. Oral Oncology 2000; 36: 67–69.
Navone N, Troncoso P, Pisters LL, Goodrow TL, Palmer JL, Nichols WW, et al. p53 protein accumulation and gene mutation in the progression of human prostate carcinoma. J Natl Cancer Inst 1993; 85: 1657–1669.
Bookstein R, MacGrogan D, Hilsenbeck SG, Sharkey F, Allred C . p53 is mutated in a subset of advanced-stage prostate cancers. Cancer Res 1993; 53: 3369–3373.
Visakorpi T, Kallioniemi OP, Heikkinen A, Koivukla T, Isola J . Small subgroup of aggressive, highly proliferative prostatic carcinoma defined by p53 accumulation. J Natl Cancer Inst 1992; 84: 883–887.
Bodner SM, Minna JD, Jensen SM, D’amico D, Carbone D, Mitsudomi T, et al. Expression of mutant p53 proteins in lung cancer correlates with the class of p53 mutations. Oncogene 1992; 7: 743–749.
Bosari S, Viale G . The clinical significance of p53 aberrations in human tumors. Virch Arch 1995; 427: 229–241.
Cattoretti G, Becker MH, Key G, Duchrow M, Schwter C, Galle J, et al. Monoclonal antibodies against recombinant parts of the Ki-67 antigen (Mib-1 and Mib-6) detect proliferating cells in microwave processed formalin-fixed paraffin sections. J Pathol 1992; 168: 357–363.
Gavrielli Y, Sherman Y, Ben-Sasson AS . Identification of programmed cell death in situ via specific labeling of nuclear DNA fragmentation. J Cell Biol 1992; 119: 493–501.
Gleason DF . Histologic grading of prostate cancer: a perspective. Hum Pathol 1992; 23: 273–279.
Humphrey PA, Vollmer RT . Intraglandular tumor extent and prognosis in prostatic carcinoma: application of a grid method to prostatectomy specimens. Hum Pathol 1990; 21: 799–804.
Schroder FH, Hermanek P, Denis L, Fair WR, Gospodarowicz MK, Pavone-Macaluso M . The TNM classification of prostate carcinoma. Prostate 1992; 4: (Suppl.) 129–138.
Wright DK, Manos MM . Sample preparation from paraffin-embedded tissues. In: Innis M, editor. PCR protocols: a guide to methods and applications. San Diego: Academic Press; 1990. p. 153.
Soong R, Iacopetta BJ . A rapid and nonisotopic method for the screening and sequencing of p53 gene mutations in formalin-fixed, paraffin-embedded tumors. Mod Pathol 1997; 10: 252–258.
Bassam BJ, Caetano-Anolles G, Gresshoff PM . Fast and sensitive silver staining of DNA polyacrylamide gels. Anal Biochem 1991; 196: 80–83.
Battifora HB, Alsabeh R, Jenkins KA, Gown A . Epitope retrieval (unmasking) in immunohistochemistry. Adv Pathol Lab Med 1995; 8: 101–118.
Taylor CR, Cote RJ . Immunocytomicroscopy: a diagnostic tool for the surgical pathologist. In: Taylor CR, Cote RJ, editors. Major problems in pathology. Vol 19. 2nd ed. Philadelphia: WB Saunders; 1994. p. 421–423.
Matsushima H, Kitamura T, Goto T, Hosaka Y, Homma Y, Kawabe K . Combined analysis with Bcl2 and p53 immunostaining predicts poorer prognosis in prostatic carcinoma. J Urol 1997; 158: 2278–2283.
Chen YQ, Gao X, Grignon D, Sakr W, Honn KV, Borders JS, et al. Multiple mechanisms of p53 inactivation in human prostate cancer. Cancer Mol Biol 1994; 1: 357–367.
Osman I, Drobnjak M, Fazzari M, Ferrara J, Scher HI, Cordon-Cardo C . Inactivation of the p53 pathway in prostate cancer: impact on tumor progression. Clin Cancer Res 1999; 5: 2082–2088.
Voeller HJ, Sugars LY, Pretlow T, Gelmann EP . p53 oncogene mutations in human prostate cancer specimens. J Urol 1994; 151: 492–495.
Mottaz AE, Markwalder R, Fey MF, Klina I, Merz VW, Thalmann GN, et al. Abnormal p53 expression is rare in clinically localized human prostate cancer: comparison between immunohistochemical and molecular detection of p53 mutations. Prostate 1997; 31: 209–215.
Chi S-G, White RWV, Meyers FJ, Siders DB, Lee F, Gumerlock PH . p53 in prostate cancer: frequent expressed transition mutations. J Natl Cancer Inst 1994; 86: 926–933.
Harris CC, Hollstein M . Clinical implications of the p53 tumor-suppressor gene. N Engl J Med 1993; 28: 1318–1327.
Sherr CJ, Weber JD . The ARF/p53 pathway. Curr Opin Genet Dev 2000; 10: 94–99.
Bueso-Ramos CE, Manshouri T, Haidar MA, Yang Y, McCown NP, Ordonez N, et al. Abnormal expression of MDM2 in breast carcinomas. Breast Cancer Res Treat 1996; 37: 179–188.
Landers JE, Cassel SL, George DL . Translational enhancement of MDM2 oncogene expression in human tumour cells containing a stabilized wild-type p53 protein. Cancer Res 1997; 57: 3562–3568.
Watanabe T, Hootta T, Ichikawa A, Kinoshita T, Nagai H, Uchida T . The MDM2 oncogene overexpression in chronic lymphocytic leukemia and low grade lymphoma of B cell origin. Blood 1994; 44: 3158–3165.
Ittmann M, Wieczorek R, Heller P, Dave A, Provet J, Krolewski J . Alterations in the p53 and MDM2 genes are infrequent in clinically localized, stage B prostate adenocarcinomas. Am J Pathol 1994; 1994: 287–293.
Xiao Z, Cheen J, Levine AJ, Modjtajedi N, Xing JS, Sellers WR, et al. Interaction between the retinoblastoma protein and the oncoprotein MDM2. Nature 1995; 375: 694–698.
Sunn P, Dong P, Dal K, Dannon GJ, Beach D . p53 independent role of MDM2 in TGF-β1 resistance. Science 1998; 282: 2270–2272.
Marchetti A, Buttitta F, Merlo G, Diella F, Pellegrini S, Pepe S, et al. p53 alterations in non-small cell lung cancers correlate with metastatic involvement of hilar and mediastinal lymph nodes. Cancer Res 1993; 53: 2846–2851.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Leite, K., Franco, M., Srougi, M. et al. Abnormal Expression of MDM2 in Prostate Carcinoma. Mod Pathol 14, 428–436 (2001). https://doi.org/10.1038/modpathol.3880330
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3880330
Keywords
This article is cited by
-
Biologically informed deep neural network for prostate cancer discovery
Nature (2021)
-
Biomarker expression patterns that correlate with high grade features in treatment naive, organ-confined prostate cancer
BMC Medical Genomics (2008)
-
Mdm2-SNP309 polymorphism in prostate cancer: no evidence for association with increased risk or histopathological tumour characteristics
British Journal of Cancer (2008)
-
Ubiquitin E3 ligase WWP1 as an oncogenic factor in human prostate cancer
Oncogene (2007)
-
The use of whole genome amplification to study chromosomal changes in prostate cancer: insights into genome-wide signature of preneoplasia associated with cancer progression
BMC Genomics (2006)