Abstract
Human herpesvirus-8 (HHV-8) latency-associated nuclear antigen (LANA) is expressed in endothelial and spindle cells of nearly all Kaposi sarcomas, and the presence of this antigen in serum is strongly correlated with the risk of developing Kaposi sarcoma in immunocompromised individuals. Studies of vascular tumors occurring in the general population show LANA expression to be specific for Kaposi sarcoma. No study to date, however, has examined whether non-Kaposi sarcoma vascular tumors arising in immunocompromised patients may express LANA, possibly reflecting origin from an HHV-8-infected endothelial progenitor cell. The objective of this study was to evaluate the specificity of LANA expression for Kaposi sarcoma in immunocompromised patients by LANA immunohistochemistry and real-time polymerase chain reaction (PCR) for HHV-8. A total of 13 cases of non-Kaposi sarcoma vascular tumors (12 hemangiomas and one epithelioid hemangioendothelioma) and 24 cases of Kaposi sarcoma, all from known HIV-positive patients, were immunostained for LANA and evaluated for the presence of HHV-8 DNA by real-time PCR. LANA expression was seen in 22 of 24 (92%) of Kaposi sarcoma cases and in 0 of 13 non-Kaposi sarcoma cases. Real-time PCR detected HHV-8 in all of the Kaposi sarcoma cases and in four of the non-Kaposi sarcoma cases (all hemangiomas). LANA expression appears to be a highly sensitive and specific marker of Kaposi sarcoma in both the general population and in HIV-positive patients. This is in contrast to HHV-8 PCR, which is positive in a small subset of non-Kaposi sarcoma vascular tumors, most likely due to detection of HHV-8 within intratumoral blood mononuclear cells by the highly sensitive real-time PCR technique. For this reason, LANA immunohistochemistry is preferable to HHV-8 PCR for the evaluation of problematic vascular proliferations in HIV-positive individuals.
Similar content being viewed by others
Main
Human herpesvirus-8 (HHV-8) is a recently described gamma herpesvirus, that was first identified as an etiologic agent in Kaposi sarcoma in 1994 by Chang et al,1 who isolated DNA sequences of HHV-8 from endothelial cells in Kaposi sarcoma lesions from patients with the acquired immune deficiency syndrome (AIDS). The development of antibodies to the HHV-8 latency-associated nuclear antigen is considered the hallmark of latent HHV-8 infection, and seropositivity for antibodies to Latency-associated nuclear antigen (LANA) is found in over 85% of patients with Kaposi sarcoma as compared with only 1–4% of the general population.2, 3 The presence of LANA antibodies in HIV-positive patients strongly correlates with the risk of developing Kaposi sarcoma.4
Monoclonal antibodies to LANA have recently been developed for use in formalin-fixed and frozen tissues, and have been shown to be reliable markers of latent HHV-8 infection in Kaposi sarcoma as well as other HHV-8-associated processes, such as primary effusion lymphoma and multicentric Castleman disease.5, 6 A small number of studies have shown LANA expression to be specific for Kaposi sarcoma among vascular tumors occurring in the general population.6, 7, 8, 9 Evaluation of LANA expression has thus been proposed as an ancillary technique for the discrimination of Kaposi sarcoma from other vascular lesions with which it may be confused, such as stasis-associated vascular proliferations, spindle cell hemangioma, kaposiform hemangioendothelioma, and spindled variants of angiosarcoma. It is currently unknown, however, whether non-Kaposi sarcoma vascular lesions arising in immunocompromised patients might express LANA, possibly reflecting origin from an HHV-8-positive endothelial progenitor cell, or secondary HHV-8 infection. We therefore studied a group of Kaposi sarcoma and non-Kaposi sarcoma vascular tumors arising in known HIV-positive patients, for evidence of HHV-8 coinfection using LANA immunohistochemistry (IHC) and HHV-8 real-time PCR.
Materials and methods
Samples
The Emory University and Grady Memorial Hospital Institutional Review Boards granted approval for this study. We retrospectively examined 37 formalin-fixed, paraffin-embedded samples obtained from 37 patients who had undergone resection or biopsy at Grady Memorial Hospital between 1993 and 2003. All patients were serologically positive for HIV by enzyme-linked immunoassay and Western blot. The patients ranged in age from 22 to 61 years, and included 35 male and two female patients. The non-Kaposi sarcoma lesions (n=13) included 12 capillary hemangiomas and one epithelioid hemangioendothelioma, and represented all of the available non-Kaposi sarcoma vascular tumors occurring in HIV-positive individuals in our institutions. The Kaposi sarcoma cases (n=24) ranged histologically from early patch to nodular stage. Both Kaposi sarcoma and non-Kaposi sarcoma lesions included a broad range of anatomic sites. The diagnoses for all cases were reconfirmed by an experienced soft-tissue pathologist (ALF) prior to inclusion in the study.
Immunohistochemistry
Sections, 5 μm thick, were cut and placed on charged glass slides. The slides were then immunostained with the LANA antibody (Clone 13B10, 1:50, Novocastra) using steam heat-induced epitope retrieval and the Dako Envision system according to the manufacturer's recommendations. Negative controls consisted of substitution of buffer for the primary antibody. The presence of any nuclear staining was scored as ‘positive’.
TaqMan Real-Time PCR
Two 5-μm paraffin-embedded sections were cut from each block and placed in Eppendorf tubes, dewaxed at 98°C for 15 min, and suspended in proteinase K digestion buffer for 1 h at 68°C. This was followed by DNA extraction using a QIAamp kit (Qiagen, Valencia, CA, USA). The samples were then centrifuged, and the supernatant was suspended in TE buffer (10 mM Tris-HCl, pH 8.0). The quality of the extracted DNA was confirmed by β-actin amplification. A body cavity B-cell lymphoma cell line (BCBL) was used as a positive control. Water was used as a negative control. An arbitrary fluorescence threshold value of 30 and cycles-to-threshold value of 50 was used. A value over the fluorescence threshold of 30 and within 50 cycles-to-threshold was considered a positive result.
The PCR reaction was performed on SmartCycler (Cepheid, Sunnyvale, CA, USA) at 50°C for 2 min, 95°C for 10 min. This was followed by 50 cycles at 95°C for 15 s and 60°C for 1 min. The following primers were used for HHV-8 detection: HHV8-1, 5′-AGCCGAAAGGATTCCACCATT-3′; HHV8-2, 5′-TCC GTGTTGTCTACGTCCAGA-3′. The HHV-8 TaqMan probe had the following sequence: 5F-TGCAGCAGC TGTTGGTGTACCACAT-T3′, where F=FAM and T=TAMRA. The following primers were used for β-actin PCR: 5′-TCACCCACACTGTGCCCATCTAC GA-3′; reverse, 5′-CAGCGGAACCGCTCATTGCCA ATGG-3′ and TaqMan probe: 5′F-ATGCCCTCCC CCA TGCCATCCTGCGT-3′, with the quencher TAMRA placed on base position 7 from the 5′ end.
Results
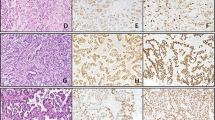
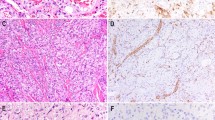
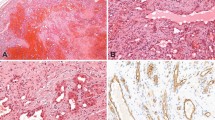
Table 1 summarizes the IHC and PCR data. LANA expression by IHC was present in 22 of 24 (92%) of the Kaposi sarcoma cases, and demonstrated a range of focal to diffuse, dot-like nuclear staining in endothelial or spindle cells (Figure 1a and b). No staining was detected in any of the 13 non-Kaposi sarcoma cases (Figures 2a, b and 3a, b).
Using real-time PCR, the BCBL-positive control was detectable at 25.63 cycles-to-threshold. HHV-8 DNA was detected in all of the Kaposi sarcoma lesions (Figure 1c), with an average cycles-to-threshold value of 45.48, and a range of 36.45–49.4. HHV-8 DNA was also detected in four of 13 (31%) of the non-Kaposi sarcoma lesions (Figure 2c), all of which were hemangiomas occurring in three males and one female patient. Repeat PCR of these four cases confirmed the presence of HHV-8 DNA at 44, 45, 46, and 47 cycles-to-threshold, respectively.
Discussion
Since Kaposi's seminal description in 1872 of the tumor bearing his name, the nature and etiology of Kaposi sarcoma has been the matter of significant dispute. Formerly a rare tumor, the incidence of Kaposi sarcoma exploded with the onset of the AIDS epidemic, and Kaposi sarcoma is now the most common tumor seen in AIDS patients, with a disproportionate incidence among homosexual and bisexual men.10, 11, 12 For this reason, an infectious etiology has long been suspected, and epidemiological, serologic, and molecular genetic studies over the past 10–15 years have identified a novel herpesvirus, HHV-8, as the likely causative agent of Kaposi sarcoma.10, 11, 12, 13, 14
HHV-8 is known to latently infect endothelial cells, as well as peripheral blood monocytes and B-lymphocytes in patients with Kaposi sarcoma. During latent infection, the virus exists in the nucleus as circular, episomal DNA that expresses a small number of viral genes.15 Although no significant difference in HHV-8 viral loads has been observed in biopsies from HIV-positive vs HIV-negative individuals with Kaposi sarcoma,16 HIV-infected individuals have a more aggressive course with frequent dissemination to skin and visceral organs.
LANA is one of the most highly expressed proteins during latent HHV-8 infection.17 The LANA protein is encoded by the open-reading frame 73 (ORF73) of the HHV-8 genome, where it tethers viral DNA to host heterochromatin and is thereby required for persistence of viral DNA in dividing cells.18, 19 LANA is also capable of interacting with pRb, which plays a regulatory role in the beginning of the S-phase of the cell cycle.20 Additionally, LANA mimics the E6 and E7 viral oncoproteins of human papillomavirus in its ability to inhibit the function of p53.21 LANA has an essential role in maintenance of the episomal DNA during latent infection and cell division, and also regulates gene expression in infected cells.22
A relatively small number of studies have examined LANA expression by IHC. Using a polyclonal antisera, Katano et al23 found LANA expression in 100% of Kaposi sarcoma cases studied. Kellam et al,6 Dupin et al5 and Courville et al,24 using the LNA53 monoclonal antibody, noted LANA expression in well over 90% of Kaposi sarcoma cases, as compared with 0% of non-Kaposi sarcoma controls. Schwartz et al,7 using the same 13B10 clone utilized in the present study, noted LANA expression in all classic and HIV-associated Kaposi sarcoma studied and in 69% of endemic (African) Kaposi sarcoma. All non-Kaposi sarcoma vascular tumors were LANA-negative. Most recently, three large studies, by Cheuk et al,8 Robin et al9 and Patel et al,25 have shown LANA expression in 71 of 72 cases, 50 of 50 cases and 21of 21 of Kaposi sarcoma, respectively, as compared with 0% of non-Kaposi sarcoma potential mimics. All of the non-Kaposi sarcoma vascular tumors in these three studies occurred in HIV-negative individuals, except for four cases of bacillary angiomatosis in the study of Cheuk et al;8 neither study examined HHV-8 positivity by PCR. The present study detected LANA expression in 92% of Kaposi sarcoma cases, a rate comparable to that seen in previous results. Our original speculation that a subset of vascular tumors from HIV-infected patients might express LANA proved not to be the case; as in the general population, we observed no LANA expression in any hemangioma occurring in HIV-positive patients, nor in the one epithelioid hemangioendothelioma. The number of such cases that we were able to accrue was, however, relatively small, despite our very extensive search of the archives of a large county hospital with a large number of HIV-positive patients. Conceivably, a study involving a larger number of cases might conceivably identify rare LANA-positive non-Kaposi sarcoma vascular tumors in immunocompromised patients.
Molecular genetic techniques, such as traditional gel-based PCR, in situ PCR, and in situ hybridization, are more sensitive than LANA IHC, and have traditionally been accepted as the ‘gold standard’ for the detection of latent HHV-8 infection. Real-time PCR, as used in this study, offers several advantages over conventional PCR. It is a quantitative assay that generates a signal based on the starting copy number only if there is amplification of the target sequence. Real-time PCR has also been shown to have greater sensitivity than conventional PCR, and is less time-consuming and cumbersome because it obviates the need for postamplification gel electrophoresis.26
Our finding of uniform HHV-8 expression in Kaposi sarcoma, by PCR, is in agreement with numerous prior studies, all of which have found >90% of Kaposi sarcoma of all types to be HHV-8 positive, using both traditional and in situ PCR.16, 27, 28, 29, 30, 31, 32, 33 The 100% detection rate we observed may in part be due to our use of a high cycles-to-threshold limit of 50. We chose to carry reactions out to 50 cycles-to-threshold for the purpose of detecting potentially very low viral loads in paraffin-embedded tissue blocks that had been in storage for up to 10 years. Although one could argue that this is the upper limit of sensitivity for the TaqMan technique, it is our experience that a value over a fluorescence threshold of 30 is evidence of a true infection despite a slightly higher cycle limit.
Our finding of HHV-8 positivity in 4 LANA-negative capillary hemangiomas is intriguing, and in agreement with a small number of studies which have documented HHV-8 positivity in occasional non-Kaposi sarcoma vascular tumors. HHV-8 DNA sequences have been identified by both PCR and Southern blot analysis in a subset of angiosarcomas by both McDonagh et al30 and Gyulai et al,34 and in a single case of pyogenic granuloma.29 None of these cases were reported to have occurred in an immunocompromised individual, and none was studied for LANA expression. The great majority of non-Kaposi sarcoma vascular tumors previously studied have, however, been negative for HHV-8 by PCR.16, 27, 28, 29, 30, 31, 32, 33 Given that the positive cases in the present study all showed levels above our preset high threshold level, we believe our results do indicate the true presence of HHV-8, rather than representing ‘false positives’. Regrettably, we did not have access to frozen tissue from these positive cases, and thus were unable to perform Southern blot confirmation. There are several possible explanations for our higher rate of detection of HHV-8 DNA in non-Kaposi sarcoma vascular tumors. We believe the most likely explanation is that the real-time PCR technique was able to detect HHV-8 within intratumoral blood mononuclear cells. A certain subset of these HIV-positive patients were likely to be coinfected with HHV-8, and probably harbored latent DNA in blood mononuclear cells. Further, the endothelial cells in capillary hemangiomas from these same patients failed to show staining with the LANA antibody. Although there is a small possibility that latent DNA is present in endothelial cells at a level that is undetectable by LANA staining, the similar PCR cycles-to-threshold values seen in the Kaposi sarcoma and non-Kaposi sarcoma cases make this explanation less likely. Lastly, although it is difficult to exclude completely the possibility of a ‘false positive’ due to amplification of a nonspecific PCR reaction product, in the absence of Southern blot confirmation, repeated PCR study of these four cases showed identical results.
We conclude that the IHC demonstration of LANA expression is a highly sensitive and specific marker of Kaposi sarcoma in both the general population, as shown previously, and in HIV-positive patients. Given the importance of distinguishing Kaposi sarcoma from its many potential mimics, it is noteworthy that no non-Kaposi sarcoma vascular tumor to date has been reported to express LANA. This is in stark contrast to both our own and previous HHV-8 PCR studies, which have documented HHV-8 positivity in a small but significant subset of non-Kaposi sarcoma vascular lesions. These observations have important clinical implications inasmuch as they support the use of LANA immunohistochemistry, rather than HHV-8 PCR, as a histomorphologic adjunct in the evaluation of problematic vascular proliferations that include Kaposi sarcoma in the differential diagnosis.
References
Chang Y, Cesarman E, Pessin MS, et al. Identification of herpesvirus-like DNA sequences in AIDS-associated Kaposi's sarcoma. Science 1994;266:1865–1869.
Simpson GR, Schulz TF, Whitby D, et al. Prevalence of Kaposi's sarcoma associated herpesvirus infection measured by antibodies to recombinant capsid protein and latent immunofluorescence antigen. Lancet 1996; 348:1133–1138.
Kedes DH, Operskalski E, Busch M, et al. The seroepidemiology of human herpesvirus 8 (Kaposi's sarcoma-associated herpesvirus): distribution of infection in KS risk groups and evidence for sexual transmission. Nat Med 1996;2:918–924.
Martin JN, Ganem DE, Osmond DH, et al. Sexual transmission and the natural history of human herpesvirus 8 infection. N Engl J Med 1998;338:948–954.
Dupin N, Fisher C, Kellam P, et al. Distribution of human herpesvirus-8 latently infected cells in Kaposi's sarcoma, multicentric Castleman's disease, and primary effusion lymphoma. Proc Natl Acad Sci USA 1999;96:4546–4551.
Kellam P, Bourboulia D, Dupin N, et al. Characterization of monoclonal antibodies raised against the latent nuclear antigen of human herpesvirus 8. J Virol 1999; 73:5149–5155.
Schwartz EJ, Dorfman RF, Kohler S . Human herpesvirus-8 latent nuclear antigen-1 expression in endemic Kaposi sarcoma: an immunohistochemical study of 16 cases. Am J Surg Pathol 2003;27:1546–1550.
Cheuk W, Wong KOY, Wong CSC, et al. Immunostaining for human herpesvirus 8 latent nuclear antigen-1 helps distinguish Kaposi sarcoma from its mimickers. Am J Clin Pathol 2004;121:335–342.
Robin Y-M, Guillou L, Michels J-J, et al. Human herpesvirus 8 immunostaining. Am J Clin Pathol 2004; 121:330–334.
Babal P, Pec J . Kaposi's sarcoma—still an enigma. J Eur Acad Dermatol Venereol 2003;17:377–380.
Mbulaiteye SM, Parkin DM, Rabkin CS . Epidemiology of AIDS-related malignancies: an international perspective. Hematol Oncol Clin North Am 2003;17:673–696 v.
Dukers NH, Rezza G . Human herpesvirus 8 epidemiology: what we do and do not know. AIDS 2003;17:1717–1730.
Stebbing J, Portsmouth S, Bower M . Insights into the molecular biology and sero-epidemiology of Kaposi's sarcoma. Curr Opin Infect Dis 2003;16:25–31.
Gandhi M, Greenblatt RM . Human herpesvirus 8, Kaposi's sarcoma, and associated conditions. Clin Lab Med 2002;22:883–910.
Boshoff C, Schulz TF, Kennedy MM, et al. Kaposi's sarcoma-associated herpesvirus infects endothelial and spindle cells. Nat Med 1995;1: 1274–1278.
Bezold G, Messer G, Peter R, et al. Quantitation of human herpesvirus 8 DNA in paraffin-embedded biopsies of HIV-associated and classical Kaposi's sarcoma by PCR. J Cutan Pathol 2001;28:127–130.
Kedes DH, Lagunoff M, Renne R, et al. Identification of the gene encoding the major latency-associated nuclear antigen of the Kaposi's sarcoma-associated herpesvirus. J Clin Invest 1997;100:2606–2610.
Komatsu T, Ballestas ME, Barbera AJ, et al. The KSHV latency-associated nuclear antigen: a multifunctional protein. Front Biosci 2002;7:d726–d730.
Szekely L, Kiss C, Mattsson K, et al. Human herpesvirus-8-encoded LNA-1 accumulates in heterochromatin- associated nuclear bodies. J Gen Virol 1999;80 (Part 11):2889–2900.
Radkov SA, Kellam P, Boshoff C . The latent nuclear antigen of Kaposi sarcoma-associated herpesvirus targets the retinoblastoma-E2F pathway and with the oncogene Hras transforms primary rat cells. Nat Med 2000;6:1121–1127.
Friborg Jr J, Kong W, Hottiger MO, et al. p53 inhibition by the LANA protein of KSHV protects against cell death. Nature 1999;402:889–894.
Verma SC, Robertson ES . Molecular biology and pathogenesis of Kaposi sarcoma-associated herpesvirus. FEMS Microbiol Lett 2003;222:155–163.
Katano H, Sato Y, Kurata T, et al. High expression of HHV-8-encoded ORF73 protein in spindle-shaped cells of Kaposi's sarcoma. Am J Pathol 1999;155:47–52.
Courville P, Simon F, Le Pessot F, et al. Detection of HHV8 latent nuclear antigen by immunohistochemistry. A new tool for differentiating Kaposi's sarcoma from its mimics. Ann Pathol 2002;22:267–276.
Patel RM, Goldblum JR, Hsi ED . Immunohistochemical detection of human herpesvirus-8 latent nuclear antigen-1 is useful in the diagnosis of Kaposi sarcoma. Mod Pathol 2004;17:456–460.
Wilhelm J, Pingoud A . Real-time polymerase chain reaction. Chembiochem 2003;4:1120–1128.
Nuovo M, Nuovo G . Utility of HHV8 RNA detection for differentiating Kaposi's sarcoma from its mimics. J Cutan Pathol 2001;28:248–255.
Kennedy MM, O'Leary JJ, Oates JL, et al. Human herpesvirus 8 (HHV-8) in Kaposi's sarcoma: lack of association with Bcl-2 and p53 protein expression. Mol Pathol 1998;51:155–159.
Hisaoka M, Hashimoto H, Iwamasa T . Diagnostic implication of Kaposi's sarcoma-associated herpesvirus with special reference to the distinction between spindle cell hemangioendothelioma and Kaposi's sarcoma. Arch Pathol Lab Med 1998;122:72–76.
McDonagh DP, Liu J, Gaffey MJ, et al. Detection of Kaposi's sarcoma-associated herpesvirus-like DNA sequence in angiosarcoma. Am J Pathol 1996;149: 1363–1368.
Kang GH, Kwon GY, Kim CW . Human herpesvirus 8 in Kaposi's sarcoma and Kaposi's sarcoma-mimicking vascular tumors. J Korean Med Sci 1998;13:54–59.
Ryan P, Aarons S, Murray D, et al. Human herpesvirus 8 (HHV-8) detected in two patients with Kaposi's sarcoma-like pyogenic granuloma. J Clin Pathol 2002; 55:619–622.
Lebbe C, Pellet C, Flageul B, et al. Sequences of human herpesvirus 8 are not detected in various non-Kaposi sarcoma vascular lesions. Arch Dermatol 1997;133: 919–920.
Gyulai R, Kemeny L, Kiss M, et al. Herpesvirus-like DNA sequence in angiosarcoma in a patient without HIV infection. N Engl J Med 1996;334:540–541.
Acknowledgements
We thank Dr Margaret Offermann and her laboratory staff from the Emory University Department of Medicine, Division of Hematology and Oncology, for providing us with a BCBL cell line.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hammock, L., Reisenauer, A., Wang, W. et al. Latency-associated nuclear antigen expression and human herpesvirus-8 polymerase chain reaction in the evaluation of Kaposi sarcoma and other vascular tumors in HIV-positive patients. Mod Pathol 18, 463–468 (2005). https://doi.org/10.1038/modpathol.3800221
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3800221
Keywords
This article is cited by
-
Gastrointestinal tract spindle cell lesions—just like real estate, it's all about location
Modern Pathology (2015)
-
Entzündung, Malignität und Immunologie gastrointestinaler Spindelzelltumoren
Der Pathologe (2014)
-
Detection of human herpesvirus 8 by quantitative polymerase chain reaction: development and standardisation of methods
BMC Infectious Diseases (2012)