Abstract
Endoscopic brush cytology is a promising surveillance technique for Barrett's esophagus. Ancillary markers are sought to increase the sensitivity of cytology and allow identification of patients at increased risk for disease progression. To determine if there are specific genetic changes in Barrett's esophagus with associated high-grade dysplasia/intramucosal adenocarcinoma compared to those without dysplasia, we performed fluorescence in situ hybridization (FISH) on cytologic specimens using probes to chromosomes and genomic regions previously described as altered in this disease. We studied archival brush cytology slides from 40 Barrett's esophagus patients: 21 with biopsy-proven high-grade dysplasia/carcinoma and 19 with no dysplasia and a minimum 5 years of negative follow-up. Centromeric enumeration probes (CEP) for chromosomes 6, 7, 11, and 12, and locus-specific probes (LSI) for 9p21 (p16 gene), and 17p13.1 (p53 gene) loci along with their corresponding CEP (9 and 17, respectively) were used in this study. A positive FISH result was defined as the presence of cells with >2 CEP signals or with a loss of the LSI signals relative to their corresponding CEP. p53 locus loss and/or aneusomy of chromosomes 6, 7, 11, and 12 abnormalities could be detected by FISH in routinely processed endoscopic brush cytology specimens from 95% of biopsy-positive cases with a specificity of 100%. Interestingly, all five cases with cytologic changes classified as indefinite for dysplasia from patients with a positive biopsy showed changes by FISH. Loss of the p16 locus was seen commonly in patients both with and without dysplasia/carcinoma. Selected biomarkers from this study merit further investigation to determine their potential to detect genetic changes in patients with Barrett's esophagus prior to the development of high-grade dysplasia.
Similar content being viewed by others
Main
Barrett's esophagus is a condition in which the normal squamous epithelium of the distal esophagus is replaced by an intestinal type columnar epithelium. Patients with this condition are at an increased risk for developing adenocarcinoma of the esophagus compared to the general population.1, 2, 3 The progression to malignancy in Barrett's esophagus is believed to occur through a multistep process, which is defined histologically as a metaplasia–dysplasia–carcinoma sequence, and is characterized by a progressive accumulation of genetic abnormalities in the epithelial cells.4, 5, 6
Current practice guidelines recommend endoscopic surveillance of patients with Barrett's esophagus in an attempt to detect malignancy at an early and potentially curable stage.7 Brush cytology may be complementary to endoscopic biopsy in surveillance programs and has a number of theoretic advantages—including the ability to sample a greater area of involved epithelium, preferential exfoliation of the less-cohesive dysplastic cells, technical simplicity, and a low cost. Cytology also provides an excellent source of material for the study of molecular genetic abnormalities using approaches such as fluorescence in situ hybridization (FISH). The potential for FISH in cytology can be illustrated by studies that demonstrate that compared to cytology, FISH detects recurrent urothelial cancer with the same high specificity but with a higher sensitivity. Furthermore, abnormalities detectable by FISH may precede the evidence of carcinoma on bladder biopsies.8, 9 FISH has also shown promise in the detection of carcinomas of the breast, lung, and colon.10, 11, 12, 13
Multiple genetic abnormalities have been reported in Barrett's esophagus-associated adenocarcinoma and its precursors. These changes include gene mutations and their frequently associated genomic losses of tumor suppressor genes including p53 (17p13.1 locus) and p16 (9p21 locus).14, 15, 16, 17, 18, 19, 20 In addition, frequent gains of chromosomes 6, 7, 11, and 12 have been previously reported in adenocarcinomas and adjacent metaplastic epithelium from esophagectomy specimens.4, 21, 22
By using endoscopic brushing cytology specimens, we evaluated the ability of FISH to detect molecular genetic changes previously reported to be associated with the malignant transformation in Barrett's esophagus. By analyzing these markers in the lesions at the extreme ends of the Barrett's-associated metaplasia–dysplasia sequence, we sought to assess the feasibility of this approach when applied to routinely processed cytologic material.
Materials and methods
Patients
Archival cytologic slides from 40 patients with biopsy-documented Barrett's esophagus obtained by conventional endoscopic brush cytology were studied. The cases studied included samples that were cytologically and histologically negative for dysplasia from patients with a minimum of 5 years of negative biopsy follow-up (n=19) as well as cytology specimens of biopsy-proven high-grade dysplasia (HGD; n=11) and intramucosal adenocarcinoma (n=10).
Cytology
For each case, a Papanicolaou-stained thin-layer liquid-based cytology slide (ThinPrepTM; Cytyc, Boxborough, MA, USA) was examined. The cases included samples diagnosed as cytologically positive, negative, or indefinite for dysplasia/adenocarcinoma. All atypical cells in the cytologically positive/indefinite cases as well as the metaplastic columnar cells in the cytologically negative cases were marked on the coverslip. Subsequently, the marks were transferred to the back of the slides using a diamond-tip pen so that the cells would remain marked after the coverslip was removed. In addition, the microscope stage coordinates of each of the cells of interest were recorded, and the selected cells were photographed using high-resolution digital camera and capture software (AxioCam with Axiovison, Zeiss, Jena, Germany).
FISH
The Papanicolaou-stained slides were treated according to a previously described protocol.23 Briefly, the slides were immersed in xylene for 48–72 h to remove the coverslips, rinsed in 100 and 95% ethanols, washed, and decolorized in 0.5% acid alcohol (HCl and 70% alcohol).
Using a pretreatment kit (Vysis, Downers Grove, IL, USA), the decolorized slides were sequentially immersed in 2 × standard saline citrate (SSC)/0.1% nonionic detergent Nonindet (NP-40; Vysis) at 73°C for 5 min, protease digested for 15 min at 37°C, washed in 1 × phosphate-buffered saline (PBS), fixed in 1% formaldehyde and washed in 1 × PBS for 5 min each at room temperature. The slides were then dehydrated in 95% followed by 100% ethanol at room temperature.
Hybridization was performed first using a multicolor, multitarget FISH probe set consisting of directly labeled locus-specific identifier (LSI) probes for the 9p21 locus (p16 gene; Spectrum Gold), the 17p13.1 locus (p53 gene; Spectrum Red), and centromeric enumeration probes (CEP) to chromosome 9 (Spectrum Green) and chromosome 17 (Spectrum Aqua) (Vysis, Downers Grove, IL, USA). The FISH probe mix (8 μl) and the target DNA were codenatured at 75°C for 5 min, the slides were coverslipped, sealed with rubber cement and incubated at 37°C overnight in a humidified chamber. The slides were then washed in 1 × SSC/0.3% NP-40) at 73°C for 2 min and rinsed in 2 × SSC/0.1% NP-40 at room temperature and left to air-dry. After counterstaining with 4′,6-diamidino-2-phenyliondole (DAPI) dimethylsulfoxide, coverslips were applied and the cells were analyzed using a fluorescent microscope equipped with a multiband pass filter (Axioplan, Zeiss, Jena, Germany). Representative cells from each slide were captured to generate a permanent record for each case, and the stage coordinates of the abnormal cells were recorded.
In order to assess subsequently the cells on the same slides using a different set of probes, stripping of the previously hybridized probes was performed using the following protocol. The slides were soaked for 5 min in 2 × SSC to remove the coverslips, then washed in 0.1 × SSC/0.03% NP-40 at 85°C for 10 min, rinsed in 2 × SSC at room temperature and left to air-dry. The slides were stained with DAPI and examined using all fluorescence filters used in the analysis to ensure that the probes were completely stripped off. The DAPI counterstain was subsequently washed off by soaking the slides in 2 × SSC for 5 min at room temperature. The slides were then reprobed in the same manner as described above, but this time a second multicolor, multitarget FISH probe set was used. This second probe set included directly labeled CEP probes for chromosomes 6 (spectrum green), 7 (spectrum gold), 11 (spectrum red), and 12 (spectrum aqua) (Vysis, Downers Grove, IL, USA). The slides were then analyzed using a fluorescent microscope as described above.
Two reviewers experienced in FISH analysis (MF, MS) evaluated all cases without knowledge of the cytology or biopsy findings. The cells were identified using the markings on the slides and the stage coordinates in conjunction with their DAPI nuclear pattern. Only cells with clearly visible nuclear outlines were counted to avoid misinterpretation of overlapping cells. Damaged or smeared nuclei were excluded from the analysis. The slides were analyzed using a set of four probe-specific filters (spectrum red, green, aqua and gold; Vysis, Downer's Grove, IL, USA). An abnormal FISH finding was recorded when a gain of any of the CEP signals was found (>2 copies of a given chromosome signal per cell), or when a loss of the locus-specific probes was seen relative to their corresponding centromeric signals (9p21 vs CEP 9 and 17p13.1 vs CEP 17). An occasional finding of a cell with four signals of all four probes in the set was not considered a positive result, as such changes can be seen in occasional normal cells in G2–M phase.
To establish the background frequency of artefacts resulting from background hybridization variation, artefactual probe signal splitting, or unrecognized cellular overlap, the two probe sets utilized in the study were initially applied to a set of 10 cell brushings obtained from benign gastric and small intestinal mucosal surgical specimens. This type of material was selected since such samples closely resemble the metaplastic columnar cells found in the Barrett's esophagus specimens. Signals from 100 nonoverlapping interphase cell nuclei were counted in these negative control samples and the mean number of cells with the signal gain/loss displayed plus 3 standard deviations per slide was 2.6 for the centromeric probes 6, 7, 9, 11, 12, and 17, 2.8 cells for the p53 probe, and 2.7 cells for the p16 probe, respectively. Therefore, the presence of three or more cells per slide showing an abnormality for any of the analyzed probes was considered indicative of the presence of a significant abnormal population in the sample. This criterion was further reassessed in a receiver operator characteristics (ROC) analysis of the data obtained from the Barrett's esophagus samples (see the Statistics section). The scoring approach used in this study is analogous to that previously developed and validated for the detection of exfoliated malignant transitional cells in urine cytology specimens by multicolor FISH.23, 24 Using this approach, 25 cells are assessed initially focusing on the atypical/metaplastic cells marked by the cytologist in conjunction with their DAPI staining pattern (large, pale, and/or nonhomogeneous staining is typical for dysplastic/carcinoma cells, elongated nuclei with homogeneous DAPI staining are the usual appearance of metaplastic columnar cells). In cases where less than the diagnostic number of cells with FISH abnormalities were identified among the initial 25 cells, additional cells were analyzed and, unless sufficient cells to establish a positive FISH diagnosis were found, all cells on the entire slide were analyzed to ensure that no abnormal population was missed. As counting of 100 or 200 random cells is relatively inefficient, the above approach enables more effective identification of an abnormal cell population in the studied samples, and minimizes false negative findings resulting from randomly selecting cells for signal counting.
Statistical Analysis
The cutoff value for the number of cells with FISH abnormality that best distinguishes the dysplasia-negative samples from those of HGD or adenocarcinoma was established from ROC curves. Cutoffs were calculated for each of the analyzed probes. As the cutoff value increases from greater than or equal to zero (at which point the sensitivity is 100% and specificity is 0%) to greater than the largest number of abnormalities (at which point the sensitivity is 0% and specificity is 100%), each sensitivity/(1−specificity) pair is plotted. Ideally, there exists a cutoff such that both sensitivity and specificity are high (or 1−specificity is low). The ROC areas under the curve greater than 0.5 indicate that the test has diagnostic utility.25
Specificity of FISH was determined using the 19 patients with negative endoscopic biopsy findings and a negative 5-year surveillance outcome. Sensitivity was calculated using the 21 patients with a positive endoscopic biopsy obtained concurrently with the cytology sample subjected to FISH. Confidence intervals for sensitivity and specificity were established using the continuity correction.
Results
The ROC analysis revealed that the optimal cutoff, that is the cutoff yielding the optimal sensitivity and specificity for detecting HGD/carcinoma, was >2 cells with a gain of one or more of the CEP probes and with a loss of the p53 (Figure 1a and b). Therefore, for the current study, the finding of three or more cells with CEP gain or p53 loss was used as a criterion for scoring a case positive by FISH. No cutoff offering high sensitivity and specificity existed for the p16 probe (Figure 1c). All three curves in Figure 1 have (Area under the curve—AUC) AUC>0.5; however, the CEP and p53 probes have much higher AUC than the p16 probe.
ROC curves for the probes used in this study. (a, b) A cutoff of >2 cells with FISH abnormality is optimal for the CEP as well as for the p53 probe. This cutoff allows the best combination of sensitivity and specificity for the detection of HGD/carcinoma in the studied cases (90 and 100% for CEP probes, 76 and 100% for the p53 probe, respectively); (c) no cutoff offering high sensitivity and specificity exists for the p16 probe. AUC greater than 0.5 indicate the marker has diagnostic utility. All three curves have AUC>0.5; however, CEP probes and p53 probe have much higher AUC than the p16 probe.
The results for all of the eight analyzed probes are summarized in Table 1. Overall, the mean number of cells with abnormalities in the FISH-positive cases was 9.8 cells per slide showing a gain with the centromeric probes (range: 4–14, s.d.: 2.6), 9.0 cells showing a loss of the p53 probe (range: 4–12, s.d.: 2.0), and 10.5 cells showing a loss of the p16 probe (range: 6–18, s.d.: 2.8). In the FISH-negative cases, the mean number cells with FISH abnormality per slide was 0.2 cells with a chromosomal gain for the centromeric probes (range: 0–2, s.d.: 0.5), 0.5 cells with a loss of the p53 probe (range: 0–2, s.d.: 0.7), and 0.5 cells for the p16 probe (range: 0–2, s.d.: 0.7).
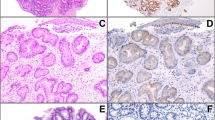
A cytology brush specimen is shown in Figure 2a. Simultaneous gains of chromosomes 6, 7, 11, and 12 were found in 19 of 21 (90%) of HGD/carcinoma cases. Although the gains of each of these four chromosomes occurred together in all positive cases, the numbers of signals for each of the particular probes were not the same and varied from 3 to 6 signals per cell (Figure 2b–d). All of the 19 cases with no dysplasia showed a diploid pattern for all four of these centromeric probes. Therefore, the sensitivity and specificity of this probe set alone for the detection of HGD/carcinoma was 90% (95% confidence interval (CI): 68–98%) and 100% (95% CI: 79–99.5%), respectively.
(a) An endoscopic brush cytology sample cytologically positive for adenocarcinoma shows hyperchromatic cells with a high nuclear–cytoplasmic ratio, irregular nuclear outlines, and prominent nucleoli; FISH analysis of the cells from this sample demonstrates gains of chromosomes 6 and 11 (b; spectrum green—five signals, and spectrum red—five to six signals), chromosome 7 (c; spectrum gold—three to five signals), and chromosome 12 (d; spectrum aqua—three signals); DAPI counterstain (b, c). Panel d was acquired without DAPI due to the poor visibility of the aqua signals against the DAPI background. Magnification: × 1000.
The assessment of the 17p13.1 (p53 gene)/chromosome 17 alterations showed a relative loss of p53 in 16/21 (76%) of the HGD/carcinoma cases. This loss was associated with a gain of chromosome 17 copies in 50% of the cases (three–five copies of chromosome 17 per cell; Figure 3a, b). Two additional HGD cases showed a synchronous gain of both the p53 and CEP 17 signals. This abnormal finding indicated a net gain of chromosome 17 without a concurrent deletion of the 17p13.1 loci and therefore, these two cases were scored as negative for the p53 loss. With the exception of one carcinoma case (case 6), which showed a loss of p53 (with a gain of chromosome 17) alone, all of the cases showing the p53/CEP17 alterations (including the two cases with a gain of chromosome 17) also showed concurrent gain of chromosomes 6, 7, 11, and 12. One carcinoma case (case 10) showed a normal pattern with the p53/CEP 17 probes, the cytology in this case was negative for dysplasia and the cells were negative for all of the remaining markers studied. Two HGD cases (both indefinite for dysplasia by cytology) showed a normal pattern with the p53/CEP17 probes; however, they showed chromosomal gains with the CEP 6, 7, 11, and 12 probes. All cases with no dysplasia included in the study showed a normal diploid pattern with the p53/CEP17 probes. Therefore, the sensitivity and specificity of p53 loss alone for the detection of HGD/carcinoma was 76% (95% CI: 52–91%) and 100% (95% CI: 52–91%), respectively.
A pair of cells from an endoscopic brush cytology sample from a patient with an intramucosal adenocarcinoma is shown. A malignant cell in the bottom portion of each of the photographs shows a relative loss of p53 signals (a; spectrum red, red arrows—two signals) compared with the corresponding centromeric signals of chromosome 17 (b; spectrum aqua, blue arrows—four signals), as well as a relative loss of the p16 signals (c; spectrum gold, yellow arrows—two signals) compared to the corresponding signals of chromosome 9 (d; spectrum green, green arrows—three signals). A nondysplastic cell in the upper portion of each of the photographs shows two copies of each of the depicted probes (a and d, DAPI counterstain). Panels (b and c) were acquired without the DAPI overlay in order to avoid decreased visibility of the signals against the blue background. In (c), the color spectrum was enhanced towards green to better visualize the cell nuclei in the absence of the DAPI overlay. Magnification: × 1000.
When combining the results of both FISH probe sets, aneusomy of chromosomes 6, 7, 11, and 12 or a p53 loss were identified in 20/21 (95%) of the HGD/carcinoma cases, including all five cases with cytologic changes classified as indefinite for dysplasia. The sensitivity and specificity for the detection of HGD/carcinoma using the combination of the above probes was 95 and 100%, respectively (95% CI: 74–99.8% and 79.1–100%; Table 2). The brushing from the above-mentioned biopsy-proven carcinoma case with a normal pattern by FISH was cytologically negative for dysplasia (case 10).
The analysis of the 9p21 (p16 gene) loss revealed that the cells from the majority of the HGD/carcinoma cases (17/21 or 81%) showed a loss of the 9p21 signals relative to the centromeric control (CEP 9). In addition, a net gain of chromosome 9 was seen in all of these cases (Figure 2c, d). One above-mentioned cytology-negative carcinoma case (case 10) showed a normal pattern for all of the studied markers including the 9p21/CEP 9 probes. Three HGD cases showed a synchronous gain of both 9p21 and CEP 9 signals, indicating a net gain of chromosome 9 without a concurrent loss of any of the 9p21 loci (all three cases also showed gains of chromosomes 6, 7, 11, and 12). Among the 19 biopsy-negative cases, nine (47%) showed a 9p21 loss. In three of these cases, this loss was associated with a gain of chromosome 9.
Discussion
Current endoscopic biopsy surveillance programs for patients with Barrett's esophagus are based on histopathological assessment of dysplasia, which is associated with problems such as intra- and inter-observer variability and sampling error.26, 27 Brush cytology may be complementary to endoscopic biopsies, and is recommended by some to be part of the routine endoscopic surveillance of patients with Barrett's esophagus.28 Cytology has a good sensitivity for the detection of adenocarcinoma and HGD, as well as a good specificity for intestinal metaplasia without dysplasia. However, its sensitivity for the detection of low-grade dysplasia is limited, which constitutes one of the reasons why cytology is currently only rarely performed as part of routine surveillance in the United States.29, 30 Ancillary techniques are sought to increase the sensitivity of cytology and to allow identification of patients at increased risk for disease progression. Since cytology specimens can provide a valuable substrate for biomarker testing, development and validation of cytology-based approaches for routine practice is highly desirable.
The FISH technique has been utilized in the analysis of cells from a variety of tumors.31 When compared with other cytogenetic techniques, FISH provides higher resolution and allows detection of chromosome alterations in small numbers of affected cells.32 Importantly, FISH can be performed on routine cytology preparations, and its results can be assessed in conjunction with cell morphology.23, 33 In this study, we have used FISH to study selected chromosome abnormalities (gains and losses) within interphase nuclei from routinely processed endoscopic brush cytology preparations. We have found that these genetic changes may be detected in cytology specimens from the majority of patients studied here with HGD or adenocarcinoma. Our results validate endoscopic brush cytology as a substrate for biomarker assessment in patients with Barrett's esophagus.
A widespread gain of DNA chromosomal material has been implicated in the progression of Barrett's esophagus-associated dysplasia, and the presence of aneuploidy, as assessed by flow cytometry, has been shown to predict the subsequent development of adenocarcinoma.26, 27 A variety of gains and losses of specific chromosomes and chromosome regions have also been detected in esophageal adenocarcinoma specimens and adjacent mucosa by both traditional cytogenetics and by comparative genomic hybridization.34, 35, 36, 37, 38, 39 The studies consistently show an accumulation of chromosomal abnormalities as the histologic changes progress from intestinal metaplasia to dysplasia and carcinoma.4, 40, 41, 42, 43 Previously reported changes detectable by FISH in esophagectomy specimens suggested that gains of chromosomes 6, 7, 11, and 12 might be among the numerical abnormalities occurring most frequently in the Barrett's esophagus-associated HGD and adenocarcinoma, findings which we have confirmed with the use of endoscopic cytology specimens.22
Only sparse data are available in the literature regarding the use of FISH on endoscopic brush cytology specimens for biomarker assessment in Barrett's esophagus. Recently, a group of investigators from Denmark applied FISH to cell pellets obtained from endoscopic brushings, and analyzed the aneusomic states of chromosomes 4, 8, 20, and Y, as well as genomic losses of p16, p53, and retinoblastoma (Rb) gene loci.44 This study showed that while aneusomy of chromosomes 4 and 8, and a p16 locus deletion are early changes present in the metaplastic Barrett's epithelium, p53 genomic loss (and rarely Rb loss) occurs predominantly in HGD and adenocarcinoma arising in Barrett's esophagus.44 A reliable distinction between benign metaplasia and dysplasia/carcinoma was, however, not possible using the markers selected by these authors. Similar findings were reported in a preliminary communication by Mayo Clinic investigators, who used FISH on brush cytology samples for the evaluation of changes affecting the p16 and p53 genes as well as chromosomes 9, 17, and Y. According to their preliminary data, the overall sensitivity and specificity of FISH for detecting HGD and adenocarcinoma was in the range of 88 and 75%, respectively.45
The results of our study indicate that a synchronous gains of chromosomes 6, 7, 11, and 12 are a highly prevalent abnormality in HGD and intramucosal adenocarcinoma (occurring in 91% of the cases). Since altered copy numbers of these four chromosomes were not found in the dysplasia-negative cases studied, this finding could be considered highly specific for the advanced lesions of the BE-associated dysplasia sequence.
It has been proposed that one of the possible mechanisms involved in the induction of chromosomal aneusomy in BE is a mutation of the p53 tumor suppressor gene. The loss of cell cycle check point control, which results from this alteration, can lead to asymmetrical chromosome segregation during cell division and subsequent aneuploidy.40 Mutations of p53 and 17p13.1 loss of heterozygosity have been reported in upto 92 and 100% of esophageal adenocarcinomas.41, 42, 43 Our study demonstrates that the genomic loss of the 17p13.1 region (p53 gene) occurs in the majority of the HGD/adenocarcinoma cases (76%). In conjunction with the presence of an aneusomy of chromosomes 6, 7, 11, and 12, the presence of the p53 gene deletion allowed us to correctly identify 20/21 (95%) of the malignant lesions, including all five cases with cytologic changes indefinite for dysplasia. A single case that was not detected by FISH was a sample obtained from a patient with an intramucosal carcinoma, which was negative by cytology, a finding that was likely attributed to an inadequate sampling of this patient's lesion.
Our findings of 9p21 (p16 gene) genomic loss in the majority of the HGD/carcinoma cases (86%) is consistent with the previous evidence that the deletion of the p16 gene locus constitutes a major genetic alteration accompanying the progression from Barrett's esophagus-related dysplasia to adenocarcinoma.14 The finding of this deletion in a substantial proportion of the nondysplastic Barrett's esophagus samples (47%) is in accordance with previous data, which indicated that 9p21 loss occurs early in the metaplasia–dysplasia sequence, and by itself may not be indicative of the presence of epithelial dysplasia.45 In addition, an increase of chromosome 9 copies as detected by the corresponding centromeric probe was found to coincide with this loss in a proportion of the cases included in our study.
In conclusion, multicolor, multitarget FISH with chromosome enumeration probes as well as locus-specific probes can be successfully applied to routinely obtained endoscopic brush cytology samples from patients with Barrett's esophagus for the assessment of potentially important genomic changes. Of the probes utilized in this pilot study, those detecting the genomic loss of 17p13.1 (p53 gene) in conjunction with the CEP 6, 7, 11, or 12 probes could detect the vast majority of HGD/adenocarcinoma cases with an excellent sensitivity and specificity. In addition, although studied to only a limited extent at this point, FISH showed the potential to become a valuable tool in the resolution of cases with changes cytologically indefinite for dysplasia. We have shown that endoscopic cytologic brushing from Barrett's esophagus represents a readily available and reliable approach for biomarker assessment by FISH. We now plan to extend these preliminary studies to the analysis of the entire spectrum of the Barrett's esophagus-associated dysplasia to evaluate the utility of this approach for the assessment of progression of these lesions to high-grade dysplasia and carcinoma. The future utilization of DNA-array technology as well as the introduction of automated microscopic FISH scanning systems will further advance this approach. These efforts may facilitate the development of FISH-based assays, which could serve as surveillance tools in patients with Barrett's esophagus in the near future.
References
Falk GW . Barrett's esophagus. Gastroenterology 2002;122:1569–1591 (Review).
Werner M, Mueller J, Walch A, et al. The molecular pathology of Barrett's esophagus. Histol Histopathol 1999;14:553–559 (Review).
Altorki NK, Oliveria S, Schrump DS . Epidemiology and molecular biology of Barrett's adenocarcinoma. Semin Surg Oncol 1997;13:270–280 (Review).
Walch AK, Zitzelsberger HF, Bruch J, et al. Chromosomal imbalances in Barrett's adenocarcinoma and the metaplasia–dysplasia–carcinoma sequence. Am J Pathol 2000;156:555–566.
Schmidt HG, Riddell RH, Walther B, et al. Dysplasia in Barrett's esophagus. J Cancer Res Clin Oncol 1985;110:145–152.
Nowell PC . The clonal evolution of tumor cell populations. Science 1976;194:23–28.
McArdle JE, Lewin KJ, Randall G, et al. Distribution of dysplasias and early invasive carcinoma in Barrett's esophagus. Hum Pathol 1992;23:479–482.
Halling KC, King W, Sokolova IA, et al. A comparison of cytology and fluorescence in situ hybridization for the detection of urothelial carcinoma. J Urol 2000;164:1768–1775.
Skacel M, Fahmy M, Brainard JA, et al. Multicolor fluorescence in situ hybridization assay UroVysion detect transitional cell carcinoma in the majority of patients with atypical and negative urinary cytology. J Urol 2003;169:2101–2105.
Heselmeyer-Haddad K, Chaudhri N, Stoltzfus P, et al. Detection of chromosomal aneuploidies and gene copy number changes in fine needle aspirates is a specific, sensitive, and objective genetic test for the diagnosis of breast cancer. Cancer Res 2002;62:2365–2369.
King BL, Tsai SC, Gryga ME, et al. Detection of chromosomal instability in paired breast surgery and ductal lavage specimens by interphase fluorescence in situ hybridization. Clin Cancer Res 2003;9:1509–1516.
Sokolova IA, Bubendorf L, O'Hare A, et al. A fluorescence in situ hybridization-based assay for improved detection of lung cancer cells in bronchial washing specimens. Cancer 2002;96:306–315.
O'Sullivan J, Bronner MP, Brentnall TA, et al. Chromosomal instability is related to telomere shortening in a human preneoplastic disease (ulcerative colitis). Nat Genet 2002;32:280–284.
Barrett MT, Sanchez CA, Galipeau PC, et al. Allelic loss of 9p21 and mutation of the CDKN2/p16 gene develop as early lesions during neoplastic progression in Barrett's esophagus. Oncogene 1996;13:1867–1873.
Huang Y, Boynton RF, Blount PL, et al. Loss of heterozygosity involves multiple tumor suppressor genes in human esophageal cancers. Cancer Res 1992;52:6525–6530.
Montesano R, Hollstein M, Hainaut P . Genetic alterations in esophageal cancer and their relevance to etiology and pathogenesis. Int J Cancer 1996;69:225–235 (Review).
Neshat K, Sanchez CA, Galipeau PC, et al. Barrett's esophagus: the biology of neoplastic progression. Gastroenterol Clin Biol 1994;18:D71–D76.
Neshat K, Sanchez CA, Galipeau PC, et al. p53 mutations in Barrett's adenocarcinoma and high-grade dysplasia. Gastroenterology 1994;106:1589–1595.
Schneider PM, Casson AG, Levin B, et al. Mutations of p53 in Barrett's esophagus and Barrett's cancer: a prospective study of ninety-eight cases. J Thorac Cardiovasc Surg 1996;111:323–331.
Younes M, Lebovitz RM, Lechago LV, et al. p53 protein accumulation in Barrett's metaplasia, dysplasia, and carcinoma: a follow-up study. Gastroenterology 1993;105:1637–1642.
Gleeson CM, Sloan JM, McGuigan JA, et al. Barrett's oesophagus: microsatellite analysis provides evidence to support the proposed metaplasia–dysplasia–carcinoma sequence. Genes Chromosomes Cancer 1998;21:49–60.
Persons DL, Croughan WS, Borelli KA, et al. Interphase cytogenetics of esophageal adenocarcinoma and precursor lesions. Cancer Genet Cytogenet 1998;106: 11–17.
Skacel M, Pettay JD, Tsiftsakis EK, et al. Validation of a multicolor interphase fluorescence in situ hybridization assay for detection of transitional cell carcinoma on fresh and archival thin-layer, liquid-based cytology slides. Anal Quant Cytol Histol 2001;23:381–387.
Sokolova IA, Halling KC, Jenkins RB, et al. The development of a multitarget, multicolor fluorescence in situ hybridization assay for the detection of urothelial carcinoma in urine. J Mol Diagn 2000;2: 116–123.
Zhou X, McClich DK, Obuchowski NA . Statistical Methods in Diagnostic Medicine. John Wiley & Sons: New York, 2002.
Reid BJ, Levine DS, Longton G, et al. Predictors of progression to cancer in Barrett's esophagus: baseline histology and flow cytometry identify low- and high-risk patient subsets. Am J Gastroenterol 2000;95:1669–1676.
Reid BJ, Blount PL, Rubin CE, et al. Flow cytometric and histological progression to malignancy in Barrett's esophagus: prospective endoscopic surveillance of a cohort. Gastroenterology 1992;102:1212–1219.
Geisinger KR . Endoscopic biopsies and cytologic brushings of the esophagus are diagnostically complementary. Am J Clin Pathol 1995;103:295–299.
Falk GW, Chittajallu R, Goldblum JR, et al. Surveillance of Barrett's esophagus for dysplasia and cancer with balloon cytology. Gastroenterology 1997;112:1787–1797.
Hardwick RH, Morgan RJ, Warren BF, et al. Brush cytology in the diagnosis of neoplasia in Barrett's esophagus. Dis Esophagus 1997;10:233–237.
King W, Proffitt J, Morrison L, et al. The role of fluorescence in situ hybridization technologies in molecular diagnostics and disease management. Mol Diagn 2000;5:309–319 (Review).
Zimmermann W, Naumann T, Bauschke E, et al. Comparison of different in situ hybridization techniques for the detection of human papillomavirus DNA in cervical smears. Acta Virol 1991;35:322–331.
Cajulis RS, Frias-Hidvegi D, Yu GH, et al. Detection of numerical chromosomal abnormalities by fluorescence in situ hybridization of interphase cell nuclei with chromosome-specific probes on archival cytologic samples. Diagn Cytopathol 1996;14:178–181.
Menke-Pluymers MB, van Drunen E, Vissers KJ, et al. Cytogenetic analysis of Barrett's mucosa and adenocarcinoma of the distal esophagus and cardia. Cancer Genet Cytogenet 1996;90:109–117.
Garewal HS, Sampliner R, Liu Y, et al. Chromosomal rearrangements in Barrett's esophagus. A premalignant lesion of esophageal adenocarcinoma. Cancer Genet Cytogenet 1989;42:281–296.
Moskaluk CA, Hu J, Perlman EJ . Comparative genomic hybridization of esophageal and gastroesophageal adenocarcinomas shows consensus areas of DNA gain and loss. Genes Chromosomes Cancer 1998;22:305–311.
Van Dekken H, Geelen E, Dinjens WN, et al. Comparative genomic hybridization of cancer of the gastroesophageal junction: deletion of 14Q31–32.1 discriminates between esophageal (Barrett's) and gastric cardia adenocarcinomas. Cancer Res 1999;59:748–752.
Varis A, Puolakkainen P, Savolainen H, et al. DNA copy number profiling in esophageal Barrett's adenocarcinoma: comparison with gastric adenocarcinoma and esophageal squamous cell carcinoma. Cancer Genet Cytogenet 2001;127:53–58.
El-Rifai W, Frierson Jr HF, Moskaluk CA, et al. Genetic differences between adenocarcinomas arising in Barrett's esophagus and gastric mucosa. Gastroenterology 2001;121:592–598.
Krishnadath KK, Tilanus HW, Alers JC, et al. Detection of genetic changes in Barrett's adencoarcinoma and Barrett's esophagus by DNA in situ hybridization and immunohistochemistry. Cytometry 1994;15:176–184.
Krishnadath KK, Tilanus HW, van Blankenstein M, et al. Accumulation of genetic abnormalities during neoplastic progression in Barrett's esophagus. Cancer Res 1995;55:1971–1976.
Riegman PH, Vissers KJ, Alers JC, et al. Genomic alterations in malignant transformation of Barrett's eosphagus. Cancer Res 2001;61:3164–3170.
Wu TT, Watanabe T, Heitmiller R, et al. Genetic alterations in Barrett esophagus and adenocarcinomas of the esophagus and esophagogastric junction region. Am J Pathol 1998;153:287–294.
Doak SH, Jenkins GJ, Parry EM, et al. Chromosome 4 hyperploidy represents an early genetic aberration in premalignant Barrett's oesophagus. Gut 2003;52:623–628.
Krishnadath S, Wang K, Halling K, et al. Detection of cytogenetic abnormalities by multi-color FISH in brush cytology specimens of Barrett's esophagus. Gastroenterology 2002;122:M1375A.
Acknowledgements
This work was supported in part by the American Society for Gastrointestinal Endoscopy Research Award. Discounted reagents were provided by Vysis Inc. (Downers Grove, IL, USA), a wholly owned subsidiary of Abbott Laboratories (Abbott Park, IL, USA).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fahmy, M., Skacel, M., Gramlich, T. et al. Chromosomal gains and genomic loss of p53 and p16 genes in Barrett's esophagus detected by fluorescence in situ hybridization of cytology specimens. Mod Pathol 17, 588–596 (2004). https://doi.org/10.1038/modpathol.3800088
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3800088
Keywords
This article is cited by
-
Point–Counterpoint: Screening and Surveillance for Barrett’s Esophagus, Is It Worthwhile?
Digestive Diseases and Sciences (2018)
-
Sponge Sampling with Fluorescent In Situ Hybridization as a Screening Tool for the Early Detection of Esophageal Cancer
Journal of Gastrointestinal Surgery (2017)
-
Clinical Implications of Molecular Changes in Pediatric Barrett’s Esophagus
Current Gastroenterology Reports (2012)