Abstract
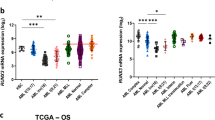
The t(8;21)(q22;q22) occurs frequently in acute myelogenous leukaemia and gives rise to the transcription factor fusion protein, RUNX1-RUNX1T1 (also known as AML1-ETO). To identify the genes dysregulated by the aberrant transcriptional activity of RUNX1-RUNX1T1, we used microarrays to determine the effect of this mutation on gene expression in human progenitor cells and during subsequent development. Gene signatures of these developmental subsets were very dissimilar indicating that effects of RUNX1-RUNX1T1 are highly context dependent. We focused on gene changes associated with the granulocytic lineage and identified a clinically relevant subset of these by comparison with 235 leukaemia patient transcriptional signatures. We confirmed the overexpression of a number of significant genes (Sox4, IL-17BR, CD200 and γ-catenin). Further, we show that overexpression of CD200 and γ-catenin is also associated with the inv(16) abnormality which like RUNX1-RUNX1T1 disrupts core binding factor activity. We investigated the functional significance of CD200 and γ-catenin overexpression in normal human progenitor cells. The effect of IL17 on growth was also assessed. Individually, none of these changes were sufficient to recapitulate the effects of RUNX1-RUNX1T1 on normal development. These data provide the most comprehensive and pertinent assessment of the effect of RUNX1-RUNX1T1 on gene expression and demonstrate the highly context-dependent effects of this fusion gene.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Peterson LF, Zhang DE . The 8;21 translocation in leukemogenesis. Oncogene 2004; 23: 4255–4262.
Miyoshi H, Kozu T, Shimizu K, Enomoto K, Maseki N, Kaneko Y et al. The t(8;21) translocation in acute myeloid leukemia results in production of an AML1-MTG8 fusion transcript. EMBO J 1993; 12: 2715–2721.
Wunderlich M, Krejci O, Wei J, Mulloy JC . Human CD34+ cells expressing the inv(16) fusion protein exhibit a myelomonocytic phenotype with greatly enhanced proliferative ability. Blood 2006; 108: 1690–1697.
Bennett JM, Catovsky D, Daniel MT, Flandrin G, Galton DA, Gralnick HR et al. Proposed revised criteria for the classification of acute myeloid leukemia. A report of the French-American-British Cooperative Group. Ann Intern Med 1985; 103: 620–625.
Reilly JT . Pathogenesis of acute myeloid leukaemia and inv(16)(p13;q22): a paradigm for understanding leukaemogenesis? Br J Haematol 2005; 128: 18–34.
Grimwade D, Walker H, Oliver F, Wheatley K, Harrison C, Harrison G et al. The importance of diagnostic cytogenetics on outcome in AML: analysis of 1612 patients entered into the MRC AML 10 trial. The Medical Research Council Adult and Children's Leukaemia Working Parties. Blood 1998; 92: 2322–2333.
Mulloy JC, Cammenga J, MacKenzie KL, Berguido FJ, Moore MA, Nimer SD . The AML1-ETO fusion protein promotes the expansion of human hematopoietic stem cells. Blood 2002; 99: 15–23.
Tonks A, Tonks AJ, Pearn L, Pearce L, Hoy T, Couzens S et al. Expression of AML1-ETO in human myelomonocytic cells selectively inhibits granulocytic differentiation and promotes their self-renewal. Leukemia 2004; 18: 1238–1245.
Lutterbach B, Westendorf JJ, Linggi B, Patten A, Moniwa M, Davie JR et al. ETO, a target of t(8;21) in acute leukemia, interacts with the N-CoR and mSin3 corepressors. Mol Cell Biol 1998; 18: 7176–7184.
Wang J, Hoshino T, Redner RL, Kajigaya S, Liu JM . ETO, fusion partner in t(8;21) acute myeloid leukemia, represses transcription by interaction with the human N-CoR/mSin3/HDAC1 complex. Proc Natl Acad Sci USA 1998; 95: 10860–10865.
Frank RC, Sun X, Berguido FJ, Jakubowiak A, Nimer SD . The t(8;21) fusion protein, AML1/ETO, transforms NIH3T3 cells and activates AP-1. Oncogene 1999; 18: 1701–1710.
Rhoades KL, Hetherington CJ, Rowley JD, Hiebert SW, Nucifora G, Tenen DG et al. Synergistic up-regulation of the myeloid-specific promoter for the macrophage colony-stimulating factor receptor by AML1 and the t(8;21) fusion protein may contribute to leukemogenesis. Proc Natl Acad Sci USA 1996; 93: 11895–11900.
Alcalay M, Meani N, Gelmetti V, Fantozzi A, Fagioli M, Orleth A et al. Acute myeloid leukemia fusion proteins deregulate genes involved in stem cell maintenance and DNA repair. J Clin Invest 2003; 112: 1751–1761.
Muller-Tidow C, Steffen B, Cauvet T, Tickenbrock L, Ji P, Diederichs S et al. Translocation products in acute myeloid leukemia activate the Wnt signaling pathway in hematopoietic cells. Mol Cell Biol 2004; 24: 2890–2904.
Shimada H, Ichikawa H, Nakamura S, Katsu R, Iwasa M, Kitabayashi I et al. Analysis of genes under the downstream control of the t(8;21) fusion protein AML1-MTG8: overexpression of the TIS11b (ERF-1, cMG1) gene induces myeloid cell proliferation in response to G-CSF. Blood 2000; 96: 655–663.
Morin PJ, Sparks AB, Korinek V, Barker N, Clevers H, Vogelstein B et al. Activation of beta-catenin-Tcf signaling in colon cancer by mutations in beta-catenin or APC. Science 1997; 275: 1787–1790.
Zheng X, Beissert T, Kukoc-Zivojnov N, Puccetti E, Altschmied J, Strolz C et al. Gamma-catenin contributes to leukemogenesis induced by AML-associated translocation products by increasing the self-renewal of very primitive progenitor cells. Blood 2004; 103: 3535–3543.
Lennon G, Auffray C, Polymeropoulos M, Soares MB . The I.M.A.G.E. Consortium: an integrated molecular analysis of genomes and their expression. Genomics 1996; 33: 151–152.
Tonks A, Pearn L, Tonks AJ, Pearce L, Hoy T, Phillips S et al. The AML1-ETO fusion gene promotes extensive self-renewal of human primary erythroid cells. Blood 2003; 101: 624–632.
Shimada H, Ichikawa H, Ohki M . Potential involvement of the AML1-MTG8 fusion protein in the granulocytic maturation characteristic of the t(8;21) acute myelogenous leukemia revealed by microarray analysis. Leukemia 2002; 16: 874–885.
Wildonger J, Mann RS . The t(8;21) translocation converts AML1 into a constitutive transcriptional repressor. Development 2005; 132: 2263–2272.
Hellstrom I, Beaumier PL, Hellstrom KE . Antitumor effects of L6, an IgG2a antibody that reacts with most human carcinomas. Proc Natl Acad Sci USA 1986; 83: 7059–7063.
Dey A, Westphal H, Nebert DW . Cell-specific induction of mouse Cyp1a1 mRNA during development. Proc Natl Acad Sci USA 1989; 86: 7446–7450.
Bullinger L, Dohner K, Bair E, Frohling S, Schlenk RF, Tibshirani R et al. Use of gene-expression profiling to identify prognostic subclasses in adult acute myeloid leukemia. N Engl J Med 2004; 350: 1605–1616.
Boyd KE, Xiao YY, Fan K, Poholek A, Copeland NG, Jenkins NA et al. Sox4 cooperates with Evi1 in AKXD-23 myeloid tumors via transactivation of proviral LTR. Blood 2006; 107: 733–741.
Ross ME, Mahfouz R, Onciu M, Liu HC, Zhou X, Song G et al. Gene expression profiling of pediatric acute myelogenous leukemia. Blood 2004; 104: 3679–3687.
McWhirter JR, Kretz-Rommel A, Saven A, Maruyama T, Potter KN, Mockridge CI et al. Antibodies selected from combinatorial libraries block a tumor antigen that plays a key role in immunomodulation. Proc Natl Acad Sci USA 2006; 103: 1041–1046.
Moreaux J, Hose D, Reme T, Jourdan E, Hundemer M, Legouffe E et al. CD200 is a new prognostic factor in multiple myeloma. Blood 2006; 108: 4194–4197.
Schwarzenberger P, Kolls JK . Interleukin 17: an example for gene therapy as a tool to study cytokine mediated regulation of hematopoiesis. J Cell Biochem Suppl 2002; 38: 88–95.
Barclay AN, Wright GJ, Brooke G, Brown MH . CD200 and membrane protein interactions in the control of myeloid cells. Trends Immunol 2002; 23: 285–290.
Tonks A, Hills R, White P, Rosie B, Mills KI, Burnett AK et al. CD200 as a prognostic factor in acute myeloid leukaemia. Leukemia 2007; 21: 566–568.
Maeda O, Usami N, Kondo M, Takahashi M, Goto H, Shimokata K et al. Plakoglobin (gamma-catenin) has TCF/LEF family-dependent transcriptional activity in beta-catenin-deficient cell line. Oncogene 2004; 23: 964–972.
Nimer SD, Moore MA . Effects of the leukemia-associated AML1-ETO protein on hematopoietic stem and progenitor cells. Oncogene 2004; 23: 4249–4254.
Westendorf JJ, Yamamoto CM, Lenny N, Downing JR, Selsted ME, Hiebert SW . The t(8;21) fusion product, AML-1-ETO, associates with C/EBP-alpha, inhibits C/EBP-alpha-dependent transcription, and blocks granulocytic differentiation. Mol Cell Biol 1998; 18: 322–333.
Pabst T, Mueller BU, Harakawa N, Schoch C, Haferlach T, Behre G et al. AML1-ETO downregulates the granulocytic differentiation factor C/EBPalpha in t(8;21) myeloid leukemia. Nat Med 2001; 7: 444–451.
Mulloy JC, Jankovic V, Wunderlich M, Delwel R, Cammenga J, Krejci O et al. AML1-ETO fusion protein up-regulates TRKA mRNA expression in human CD34+ cells, allowing nerve growth factor-induced expansion. Proc Natl Acad Sci USA 2005; 102: 4016–4021.
Moseley TA, Haudenschild DR, Rose L, Reddi AH . Interleukin-17 family and IL-17 receptors. Cytokine Growth Factor Rev 2003; 14: 155–174.
Li J, Shen H, Himmel KL, Dupuy AJ, Largaespada DA, Nakamura T et al. Leukaemia disease genes: large-scale cloning and pathway predictions. Nat Genet 1999; 23: 348–353.
Reichling T, Goss KH, Carson DJ, Holdcraft RW, Ley-Ebert C, Witte D et al. Transcriptional profiles of intestinal tumors in Apc(Min) mice are unique from those of embryonic intestine and identify novel gene targets dysregulated in human colorectal tumors. Cancer Res 2005; 65: 166–176.
Wright GJ, Cherwinski H, Foster-Cuevas M, Brooke G, Puklavec MJ, Bigler M et al. Characterization of the CD200 receptor family in mice and humans and their interactions with CD200. J Immunol 2003; 171: 3034–3046.
Polakis P . Wnt signaling and cancer. Genes Dev 2000; 14: 1837–1851.
Charpentier E, Lavker RM, Acquista E, Cowin P . Plakoglobin suppresses epithelial proliferation and hair growth in vivo. J Cell Biol 2000; 149: 503–520.
Simcha I, Geiger B, Yehuda-Levenberg S, Salomon D, Ben-Ze’ev A . Suppression of tumorigenicity by plakoglobin: an augmenting effect of N-cadherin. J Cell Biol 1996; 133: 199–209.
Breault JE, Shiina H, Igawa M, Ribeiro-Filho LA, Deguchi M, Enokida H et al. Methylation of the gamma-catenin gene is associated with poor prognosis of renal cell carcinoma. Clin Cancer Res 2005; 11 (Part 1): 557–564.
Polychronopoulou S, Tsatsopoulou A, Papadhimitriou SI, Panagiotou JP, Anastasakis A, Paterakis G et al. Myelodysplasia and Naxos disease: a novel pathogenetic association? Leukemia 2002; 16: 2335–2337.
Austin TW, Solar GP, Ziegler FC, Liem L, Matthews W . A role for the Wnt gene family in hematopoiesis: expansion of multilineage progenitor cells. Blood 1997; 89: 3624–3635.
He TC, Sparks AB, Rago C, Hermeking H, Zawel L, da Costa LT et al. Identification of c-MYC as a target of the APC pathway. Science 1998; 281: 1509–1512.
Shtutman M, Zhurinsky J, Simcha I, Albanese C, D’Amico M, Pestell R et al. The cyclin D1 gene is a target of the beta-catenin/LEF-1 pathway. Proc Natl Acad Sci USA 1999; 96: 5522–5527.
Simon M, Grandage VL, Linch DC, Khwaja A . Constitutive activation of the Wnt/beta-catenin signalling pathway in acute myeloid leukaemia. Oncogene 2005; 24: 2410–2420.
Jamieson CH, Ailles LE, Dylla SJ, Muijtjens M, Jones C, Zehnder JL et al. Granulocyte-macrophage progenitors as candidate leukemic stem cells in blast-crisis CML. N Engl J Med 2004; 351: 657–667.
Zhurinsky J, Shtutman M, Ben-Ze’ev A . Differential mechanisms of LEF/TCF family-dependent transcriptional activation by beta-catenin and plakoglobin. Mol Cell Biol 2000; 20: 4238–4252.
Zhurinsky J, Shtutman M, Ben-Ze’ev A . Plakoglobin and beta-catenin: protein interactions, regulation and biological roles. J Cell Sci 2000; 113 (Part 18): 3127–3139.
Wilson A, Murphy MJ, Oskarsson T, Kaloulis K, Bettess MD, Oser GM et al. c-Myc controls the balance between hematopoietic stem cell self-renewal and differentiation. Genes Dev 2004; 18: 2747–2763.
Zhang J, Niu C, Ye L, Huang H, He X, Tong WG et al. Identification of the haematopoietic stem cell niche and control of the niche size. Nature 2003; 425: 836–841.
Acknowledgements
We would like to thank Xiaomin Zheng and Martin Ruthardt (Johann Wolfgang Goethe-Universität, Frankfurt, Germany) for providing the γ-catenin-PINCO plasmid. We also acknowledge the MRC for access to patient sample material enrolled in the NCRI clinical trials (MRC-AML 12, 14 and 15). This work was supported by Leukaemia Research, UK.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Leukemia website (http://www.nature.com/leu)
Supplementary information
Rights and permissions
About this article
Cite this article
Tonks, A., Pearn, L., Musson, M. et al. Transcriptional dysregulation mediated by RUNX1-RUNX1T1 in normal human progenitor cells and in acute myeloid leukaemia. Leukemia 21, 2495–2505 (2007). https://doi.org/10.1038/sj.leu.2404961
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2404961
Keywords
This article is cited by
-
CD200 expression in T cell neoplasms
Annals of Hematology (2023)
-
Validation and refinement of a RUNX1 mutation-associated gene expression signature in blast crisis chronic myeloid leukemia
Leukemia (2022)
-
Increased expression of RUNX3 inhibits normal human myeloid development
Leukemia (2022)
-
AML1/ETO and its function as a regulator of gene transcription via epigenetic mechanisms
Oncogene (2021)
-
The RUNX1-ETO target gene RASSF2 suppresses t(8;21) AML development and regulates Rac GTPase signaling
Blood Cancer Journal (2020)