Abstract
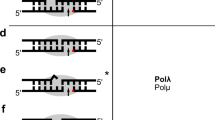
In most models of DNA replication, Watson–Crick hydrogen bonding drives the incorporation of nucleotides into the new strand of DNA and maintains the complementarity of bases with the template strand. Studies with nonpolar analogues of thymine and adenine, however, have shown that replication is still efficient in the absence of hydrogen bonds1,2,3,4. The replication of base pairs might also be influenced by steric exclusion, whereby inserted nucleotides need to be the correct size and shape to fit the active site against a template base5,6. A simple steric-exclusion model may not require Watson–Crick hydrogen bonding to explain the fidelity of replication, nor should canonical purine and pyrimidine shapes be necessary for enzymatic synthesis of a base pair if each can fit into the DNA double helix without steric strain6. Here we test this idea by using a pyrene nucleoside triphosphate (dPTP) in which the fluorescent ‘base’ is nearly as large as an entire Watson–Crick base pair. We show that the non-hydrogen-bonding dPTP is efficiently and specifically inserted by DNA polymerases opposite sites that lack DNA bases. The efficiency of this process approaches that of a natural base pair and the specificity is 102–104-fold. We use these properties to sequence abasic lesions in DNA, which are a common form of DNA damage in vivo7. In addition to their application in identifying such genetic lesions, our results show that neither hydrogen bonds nor purine and pyrimidine structures are required to form a base pair with high efficiency and selectivity. These findings confirm that steric complementarity is an important factor in the fidelity of DNA synthesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Moran, S., Ren, R. X.-F. & Kool, E. T. Athymidine triphosphate shape mimic lacking Watson–Crick pairing ability is replicated with high specificity. Proc. Natl Acad. Sci. USA 94, 10506–10511 (1997).
Goodman, M. F. Hydrogen bonding revisited: geometric selection as a principal determinant of DNA replication fidelity. Proc. Natl Acad. Sci. USA 94, 10493–10495 (1997).
Morales, J. C. & Kool, E. T. Efficient replication of a DNA base pair between non-hydrogen-bonded nucleoside analogues. Nature Struct. Biol. 5, 950–954 (1998).
Moran, S., Ren, R. X.-F., Rumney, S. & Kool, E. T. Difluorotoluene, a nonpolar isostere of thymine, codes specifically and efficiently for adenine in DNA replication. J. Am. Chem. Soc. 119, 2056–2057 (1997).
Goodman, M. F., Creighton, S., Bloom, L. B. & Petruska, J. Biochemical basis of DNA replication fidelity. Crit. Rev. Biochem. Mol. Biol. 28, 83–126 (1993).
Kool, E. T. Replication of non-hydrogen bonded bases by DNA polymerases: a mechanism for steric matching. Biopolymers (Nucleic Acid Sciences) 48, 3–17 (1998).
Loeb, L. A. & Preston, B. D. Mutagenesis by apurinic/apyrimidinic sites. Annu. Rev. Genet. 20, 201–230 (1986).
Ren, R. X.-F., Chaudhuri, N. C., Paris, P. L., Rumney, S. & Kool, E. T. Naphthalene, phenanthrene, and pyrene as DNA base analogues: synthesis, structure, and fluorescence in DNA. J. Am. Chem. Soc. 118, 7671–7678 (1996).
Matray, T. J. & Kool, E. T. Selective and stable DNA base pairing without hydrogen bonds. J. Am. Chem. Soc. 120, 6191–6192 (1998).
Mishra, M. C. & Broom, A. D. Anovel synthesis of nucleoside 5′ triphosphates. J. Chem. Soc., Chem. Commun. 1276–1277 (1991).
Hoard, D. E. & Ott, D. G. Conversion of mono- and oligodeoxyribonucleotides to 5′-triphosphates. J. Am. Chem. Soc. 87, 1785–1788 (1965).
Millican, T. A. et al. . Synthesis and biophysical studies of short oligodeoxynucleotides with novel modifications. Nucleic Acids Res. 12, 7435–7453 (1984).
Takeshita, M., Chang,,, C.,, Johnson, F., Will, S. & Grollman, A. P. Oligodeoxy-nucleotides containing synthetic abasic sites. Model substrates for DNA polymerases and apurinic/apyrimidinic endonucleases. J. Biol. Chem. 262, 10171–10179 (1987).
Randall, S. K., Eritja, R., Kaplan, B. E., Petruska, J. & Goodman, M. F. Nucleotide insertion kinetics opposite abasic lesions in DNA. J. Biol. Chem. 262, 6864–6870 (1987).
Sagher, D. & Strauss, B. Insertion of nucleotides opposite apurinic/apyrimidinic sites in deoxyribonucleic acid during in vitro synthesis: uniqueness of adenine nucleotides. Biochemistry 22, 4518–4526 (1983).
Schaaper, R. M., Kunkel, T. A. & Loeb, L. A. Infidelity of DNA synthesis associated with bypass of apurinic sites. Proc. Natl Acad. Sci. USA 80, 487–491 (1983).
Lawrence, C. W., Borden, A., Banerjee, S. K. & LeClerc, J. E. Mutation frequency and spectrum resulting from a single abasic site in a single-stranded vector. Nucleic Acids Res. 18, 2153–2157 (1990).
Paz-Elizur, T., Takeshita, M. & Livneh, Z. Mechanism of bypass synthesis through an abasic site analog by DNA polymerase I. Biochemistry 36, 1766–1773 (1997).
Doublié, S., Tabor, S., Long, A. M., Richardson, C. C. & Ellenberger, T. Crystal structure of a bacteriophage T7 DNA replication complex at 2.2 å resolution. Nature 391, 251–258 (1998).
Kiefer, J. R., Mao, C., Braman, J. C. & Beese, L. S. Visualizing DNA replication in a catalytically active Bacillus DNA polymerase crystal. Nature 391, 304–307 (1998).
Huang, H., Chopra, R., Verdine, G. L. & Harrison, S. C. Structure of a covalently trapped catalytic complex of HIV-1 reverse transcriptase: implications for drug resistance. Science 282, 1669–1675 (1998).
Diederichsen, U. Selectivity of DNA replication: the importance of geometry over hydrogen bonding. Angew. Chem. 37, 1655–1657 (1998).
Fygenson, D. K. & Goodman, M. F. Appendix. Gel kinetic analysis of polymerase fidelity in the presence of multiple enzyme DNA encounters. J. Biol. Chem. 272, 27931–27935 (1997).
Shibutani, S., Takeshita, M. & Grollman, A. P. Translesional synthesis on DNA templates containing a single abasic site. A mechanistic study of the “A rule”. J. Biol. Chem. 272, 13916–13921 (1997).
Lindahl, T. & Nyberg, B. Rate of depurination of native deoxyribonucleic acid. Biochemistry 11, 3610–3618 (1972).
Weiss, B. & Grossman, L. Phosphodiesterases involved in DNA repair. Adv. Enzymol. Rel. Areas Mol. Biol. 60, 1–34 (1972).
Ide, H. et al. . Synthesis and damage specificity of a novel probe for the detection of abasic sites in DNA. Biochemistry 32, 8276–8283 (1993).
Maulik, G. et al. . Novel non-isotopic detection of MutY enzyme-recognized mismatches in DNA via ultrasensitive detection of aldehydes. Nucleic Acids Res. 27, 1316–1322 (1999).
Denissenko, M. F., Pao, A., Tang, M. & Pfeifer, G. P. Preferential formation of benzo[a ]pyrene adducts at lung cancer mutational hotspots in p53. Science 274, 430–432 (1996).
Paris, P. L., Langenhan, J. & Kool, E. T. Probing DNA sequences in solution with a monomer–excimer fluorescence color change. Nucleic Acids Res. 26, 3789–3793 (1998).
Acknowledgements
This work was supported by grants from the NIH and from the US Army Research Office.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Matray, T., Kool, E. A specific partner for abasic damage in DNA. Nature 399, 704–708 (1999). https://doi.org/10.1038/21453
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/21453
This article is cited by
-
Fluorescent nucleobases as tools for studying DNA and RNA
Nature Chemistry (2017)
-
Cycloadditions for Studying Nucleic Acids
Topics in Current Chemistry (2016)
-
Identification of DNA lesions using a third base pair for amplification and nanopore sequencing
Nature Communications (2015)
-
Watson–Crick versus imidazopyridopyrimidine base pairs: theoretical study on differences in stability and hydrogen bonding strength
Structural Chemistry (2014)
-
Removal of uracil by uracil DNA glycosylase limits pemetrexed cytotoxicity: overriding the limit with methoxyamine to inhibit base excision repair
Cell Death & Disease (2012)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.