Abstract
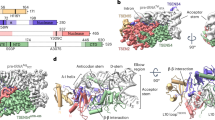
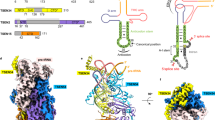
The Sex-lethal (Sxl) protein of Drosophila melanogaster regulates alternative splicing of the transformer (tra) messenger RNA precursor by binding to the tra polypyrimidine tract during the sex-determination process. The crystal structure has now been determined at 2.6 Å resolution of the complex formed between two tandemly arranged RNA-binding domains of the Sxl protein and a 12-nucleotide, single-stranded RNA derived from the tra polypyrimidine tract. The two RNA-binding domains have their β-sheet platforms facing each other to form a V-shaped cleft. The RNA is characteristically extended and bound in this cleft, where the UGUUUUUUU sequence is specifically recognized by the protein. This structure offers the first insight, to our knowledge, into how a protein binds specifically to a cognate RNA without any intramolecular base-pairing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Harford, J. B. & Morris, D. R. mRNA Metabolism and Post-transcriptional Gene Regulation (Wiley-Liss, New York, 1997).
Cline, T. W. & Meyer, B. J. Vive la difference: males vs females in flies vs worms. Annu. Rev. Genet. 30, 637–702 (1996).
Sosnowski, B. A., Belote, J. M. & McKeown, M. Sex-specific alternative splicing of RNA from the transformer gene results from sequence-dependent splice site blockage. Cell 58, 449–459 (1989).
Inoue, K., Hoshijima, K., Sakamoto, H. & Shimura, Y. Binding of the Drosophila Sex-lethal gene product to the alternative splice set of transformer primary transcript. Nature 344, 461–463 (1990).
Valcárcel, J., Singh, R., Zamore, P. D. & Green, M. R. The protein Sex-lethal antagonizes the splicing factor U2AF to regulate alternative splicing of transformer pre-mRNA. Nature 362, 171–175 (1993).
Sakashita, E. & Sakamoto, H. Characterization of RNA binding specificity of the Drosophila sex-lethal protein by in vitro ligand selection. Nucleic Acids Res. 22, 4082–4086 (1994).
Singh, R., Valcárcel, J. & Green, M. R. Distinct binding specificities and functions of higher eukaryotic polypyrimidine tract-binding proteins. Science 268, 1173–1176 (1995).
Burd, C. G. & Dreyfuss, G. Conserved structures and diversity of functions of RNA-binding proteins. Science 265, 615–621 (1994).
Birney, E., Kumar, S. & Krainer, A. R. Analysis of the RNA-recognition motif and RS and RGG domains: conservation in metazoan pre-mRNA splicing. Nucleic Acids Res. 21, 5803–5816 (1993).
Varani, G. & Nagai, K. RNA recognition by RNP proteins during RNA processing. Annu. Rev. Biopys. Biomol. Struct. 27, 407–445 (1998).
Nagai, K., Oubridge, C., Jessen, T. H., Li, J. & Evans, P. R. Crystal structure of the RNA-binding domain of the U1 small nuclear ribonucleoprotein A. Nature 348, 515–520 (1990).
Oubridge, C., Ito, N., Evans, P., Teo, C.-H. & Nagai, K. Crystal structure at 1.92 Å resolution of the RNA-binding domain of the U1A spliceosomal protein complexed with an RNA hairpin. Nature 372, 432–438 (1994).
Allain, F. H.-T. et al. Specificity of ribonucleoprotein interaction determined by RNA folding during complex formation. Nature 380, 646–650 (1996).
Price, S. R., Evans, P. R. & Nagai, K. Crystal structure of the spliceosomal U2B″-U2A′ protein complex bound to a fragment of U2 small nuclear RNA. Nature 394, 645–650 (1998).
Kanaar, R., Lee, A. L., Rudner, D. Z., Wemmer, D. E. & Rio, D. C. Interaction of the Sex-lethal RNA binding domains with RNA. EMBO J. 14, 4530–4539 (1995).
Sakashita, E. & Sakamoto, H. Protein–RNA and protein–protein interactions of the Drosophila Sex-lethal mediated by its RNA-binding domains. J. Biochem. 120, 1028–1033 (1996).
Samuels, M., Deshpande, G. & Schedl, P. Activities of the Sex-lethal protein in RNA binding and protein:protein interactions. Nucleic Acids Res. 26, 2625–2637 (1998).
Inoue, M. et al. Acharacteristic arrangement of aromatic amino acid residues in the solution structure of the amino-terminal RNA-binding domain of Drosophila Sex-lethal. J. Mol. Biol. 272, 82–94 (1997).
Lee, A. L., Kanaar, R., Rio, D. C. & Wemmer, D. E. Resonance assignments and solution structure of the second RNA-binding domain of Sex-lethal determined by multidimensional heteronuclear magnetic resonance. Biochemistry 33, 13775–13786 (1994).
Shamoo, Y., Krueger, U., Rice, L. M., Williams, K. R. & Steitz, T. A. Crystal structure of the two RNA-binding domains of human hnRNP A1 at 1.75 Å resolution. Nature Struct. Biol. 4, 215–222 (1997).
Xu, R.-M., Jokhan, L., Cheng, X., Mayeda, A. & Krainer, A. R. Crystal structure of human UP1, the domain of hnRNP A1 that contains two RNA-recognition motifs. Structure 5, 559–570 (1997).
Simons, R. W. & Grunberg-Manago, M. RNA Structure and Function.(Cold Spring Harbor Laboratory Press, New York, 1998).
Valegárd, K., Murray, J. B., Stockley, P. G., Stonehouse, N. J. & Liljas, L. Crystal structure of an RNA bacteriophage coat protein–operator complex. Nature 371, 623–626 (1994).
Puglisi, J. D., Chen, L., Blanchard, S. & Frankel, A. D. Solution structure of a bovine immunodeficiency virus Tat-TAR peptide–RNA complex. Science 270, 1200–1203 (1995).
Battiste, J. L. et al. αHelix–RNA major groove recognition in an HIV-1 Rev peptide-RRE RNA complex. Science 273, 1547–1551 (1996).
De Guzman, R. N. et al. Structure of the HIV-1 nucleocapsid protein bound to the SL3 Ψ-RNA recognition element. Science 279, 384–388 (1998).
Cai, Z. et al. Solution structure of P22 transcriptional antitermination N peptide-box B RNA complex. Nature Struct. Biol. 5, 203–212 (1998).
Lee, A. L. et al. Chemical shift mapping of the RNA-binding interface of the multiple-RBD protein Sex-lethal. Biochemistry 36, 14306–14317 (1997).
Hall, K. B. Interaction of RNA hairpins with the human U1A N-terminal RNA binding domain. Biochemistry 33, 10076–10088 (1994).
Good, P. J. The role of elav -like genes, a conserved family encoding RNA-binding proteins, in growth and development. Semin. Cell Dev. Biol. 8, 577–584 (1997).
Fan, X. C. & Steitz, J. A. Overexpression of HuR, a nuclear-cytoplasmic shuttling protein, increases the in vivo stability of ARE-containing mRNAs. EMBO J. 17, 3448–3460 (1998).
Peng, S. S.-Y., Chen, C.-Y. A., Xu, N. & Shyu, A.-B. RNA stabilization by the AU-rich element binding protein, HuR, an ELAV protein. EMBO J. 17, 3461–3470 (1998).
Rould, M. A., Perona, J. J. & Steitz, T. A. Structural basis of anticodon loop recognition by glutaminyl-tRNA synthetase. Nature 352, 213–218 (1991).
Cavarelli, J., Rees, B., Ruff, M., Thierry, J.-C. & Moras, D. Yeast tRNAAsp recognition by its cognate class II aminoacyl-tRNA synthetase. Nature 362, 181–184 (1993).
Biou, V., Yaremchuk, A., Tukalo, M. & Cusack, S. The 2.9 Å crystal structure of T. thermophilus seryl-tRNA synthetase complexed with tRNASer. Science 263, 1404–1410 (1994).
Cusack, S., Yaremchuk, A. & Tukalo, M. The crystal structures of T. thermophilus lysyl-tRNA synthetase complexed with E. coli tRNALys and a T. thermophilus tRNALys transcript: anticodon recognition and conformational changes upon binding of a lysyl-adenylate analogue. EMBO J. 15, 6321–6334 (1996).
Cusack, S., Yaremchuk, A., Krikliviy, I. & Tukalo, M. tRNAPro anticodon recognition by Thermus thermophilus prolyl-tRNA synthetase. Structure 6, 101–108 (1998).
Shamoo, Y., Abdul-Manan, N. & Williams, K. R. Multiple RNA binding domains (RBDs) just don't add up. Nucleic Acids Res. 23, 725–728 (1995).
McKay, S. J. & Cooke, H. hn RNP A2/B1 binds specifically to single stranded vertebrate telomeric repeat TTAGGGn. Nucleic Acids Res. 20, 6461–6464 (1992).
Kajita, Y., Nakayama, J., Aizawa, M. & Ishikawa, F. The UUAG-specific RNA binding protein, heterogenous nuclear ribonucleoprotein D0. J. Biol. Chem. 270, 22167–22175 (1995).
Sarig, G., Weisman-Shomer, P., Erlitzki, R. & Fry, M. Purification and characterization of qTBP42, a new single-stranded and quadruplex telomeric DNA-binding protein from rat hepatocytes. J. Biol. Chem. 272, 4474–4482 (1997).
Kim, I. et al. NMR analysis of the hydrogen bonding interactions of the RNA-binding domains of the Drosophila Sex-lethal protein with target RNA fragments with site-specific [3-15N]uridine substitutions. Nucleic Acids Res. 25, 1565–1569 (1997).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
CCP4: Collaborative Computational Project No. 4. Acta Crystallogr. D 50, 760–763 (1994).
Jones, T. A., Zhou, J.-Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for binding protein models in electron density maps and the location of errors in these models. Acta. Crystallogr. A 47, 110–119 (1991).
Brünger, A. T. X-PLOR: A System for X-Ray Crystallography and NMR(Yale Univ. Press, New Haven, 1992).
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Kraulis, P. Molscript: a program to produce both detailed and schematic plots of protein structures. J.Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E. A. & Bacon, D. J. Raster3D photorealistic molecular graphics. Methods Enzymol. 277, 505–524 (1997).
Koradi, R., Billeter, M. & Wüthrich, K. MOLMOL: a program for display and analysis of macromolecular structures. J. Mol. Graph. 14, 51–55, 29–32 (1996).
Acknowledgements
We thank D. G. Vassylyev for helpful discussions, and M. Horikoshi for data collection. This work was supported in part by Grants-in-Aid for Scientific Research on Priority Areas to S.Y. from the Ministry of Education, Science, Culture and Sports of Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Handa, N., Nureki, O., Kurimoto, K. et al. Structural basis for recognition of the tra mRNA precursor by the Sex-lethal protein. Nature 398, 579–585 (1999). https://doi.org/10.1038/19242
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/19242
This article is cited by
-
The solution structure of Dead End bound to AU-rich RNA reveals an unusual mode of tandem RRM-RNA recognition required for mRNA regulation
Nature Communications (2022)
-
De novo computational RNA modeling into cryo-EM maps of large ribonucleoprotein complexes
Nature Methods (2018)
-
The role of small-angle scattering in structure-based screening applications
Biophysical Reviews (2018)
-
DND1 maintains germline stem cells via recruitment of the CCR4–NOT complex to target mRNAs
Nature (2017)
-
Structural basis for the assembly of the Sxl–Unr translation regulatory complex
Nature (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.