Abstract
In situ hybridisation technology provides a new tool for chromosome analysis of human spermatozoa. We have used dual-colour fluorescence in situ with probes specific for the X and Y chromosomes and chromosomes 1 and 12 to (a) identify the primary male gametic sex chromosome ratio; (b) assess the number of numerical sex chromosome abnormalities, and (c) quantify the incidence of diploid sperm. We have examined over 60,000 sperm from three normal males and found the primary sex ratio to be indistinguishable from unity. The frequency of hyperhaploid sperm was 0.8, 1.03 and 2.27 per thousand for XX, YY and XY respectively, whilst 1.67 per thousand sperm were diploid. A comparison of our results with estimates of sex chromosome aneuploidy in human populations suggests that sperm carrying two sex chromosomes may be at a selective disadvantage.
Similar content being viewed by others
Introduction
Information on the sex chromosome content of sperm has been obtained from studies using differential staining methods, sperm karyotyping, and in situ hybridisation. So far each of these techniques has met with serious limitations. Quinacrine fluorescent staining, for detection of the F-body, i.e., the Yqh region [1], produces some non-specific signals and has recently been confirmed as unreliable [2]. Sperm karyotyping, relying on pseudo-fertilisation of hamster eggs, allows detection of both X and Y sperm but is an expensive and laborious technique, impractical for the analysis of large numbers of gametes from anyone individual [3]. In situ hybridisation with chromosome-specific probes had proved difficult on ejaculate sperm in which the chromatin is highly condensed and protected by disulphide bridges [4]. However, this latter problem has been successfully overcome using various methods for sperm head decondensation which render the DNA more accessible to hybridisation [4–8]. These investigations have only illustrated the use of a single sex-chromosome-specific probe at a time. Data on the second chromosome has only been inferred from the proportion of sperm not hybridising for the other. Furthermore diploidy rates have only been assessed using nuclear size and not directly from chromosome content.
In order to determine the primary gametic sex chromosome ratio, the meiotic non-disjunction rates of X and Y, as well as the diploidy frequencies in human sperm, we have carried out dual-colour fluorescence in situ hybridisation with two sets of probes on individual spermatozoa from three males of proven fertility. Cohybridising with X-and Y-specific probes allowed us to count the number of gametes which were hypohaploid, haploid or apparently hyperhaploid for the sex chromosomes. This gave direct data on the ratio of Y-to X-bearing sperm. Knowledge of the diploidy rate, determined using autosomal probes, made it possible to calculate revised frequencies of sex-chromosome-aneuploid sperm.
Materials and Methods
Collection and Preparation of Semen Samples
Three men aged 28, 38, and 34 years, of proven fertility and with karyotypes indistinguishable from normal (known as D69, D117 and D218, respectively) were sperm donors at an in vitro fertilisation clinic. Their semen was frozen according to a modification of the method of Behrman and Sawada [9], each ejaculate was mixed with an equal volume of buffer (39.1 mM tri-sodium citrate, 83.3 mM glucose, 133.2 mM glycine, 15% glycerol, 20% egg yolk, pH 7.2–7.4) and loaded into labelled plastic straws. The straws were cooled in nitrogen vapour for 10 min before being transferred to liquid nitrogen for storage.
One straw from each man was warmed to room temperature and rinsed out by syringing through phosphate-buffered saline (PBS). The suspension was washed three times in excess PBS. Between washes the spermatozoa were collected by centrifugation at 1,500 g for 10 min. After the final wash as much buffer as possible was removed leaving only the pellet of spermatozoa. These were fixed slowly with an excess of ice-cold 3:1 methanol:acetic acid. After standing for 10 min at room temperature, centrifugation and fixation were repeated, finally providing a concentration of 40–60 sperm per field of view under × 400 magnification. The suspension was dropped onto standard glass microscope slides, air-dried and stored at −70°C for up to 4 months.
Hybridization and Detection
Excess fixative was removed from the slides by rinsing three times in 2 × SSC (pH 7) at room temperature (5 min each) followed by ethanol dehydration. Sperm head decondensation was carried out by incubating the slides at room temperature for 12 min in a solution of 25 mM dithiothreitol/0.1% trypsin [6]. This was followed by rinsing in 2 × SSC and alcohol dehydration.
The four probes pSVX (X centromere), pY10 (Y paracentromeric), pUC1.77 (1 paracentromeric) and pα12H8 (12 centromere) were variously labelled with the modified nucleotides: biotin-11-dUTP (bio-dUTP); digoxigenin-11-dUTP (dig-dUTP); rhodamine-6-dUTP (rhod-dUTP) (FluoroRed, Amersham) or aminomethylcoumarin-dUTP (AMCA-dUTP) (FluoroBlue, Amersham). This was done by a two-step nick translation reaction in which the probes were first DNAse I digested to a range of 0.1–5 kb, and 3 µg were then labelled in a 50-µl DNA polymerase I reaction similar to that which we have previously used for biotinylation [10]. The reaction concentration of bio-dUTP and dig-dUTP was 20 µM. Rhod-dUTP and AMCA-dUTP were both used at 8 µM in the presence of 12 µM dTTP.
In situ Hybridisation
The final hybridisation mixture consisted of 60% formamide, 2.2 × SSC, 11% dextran sulphate (MW = 500,000) and probe DNA at concentration of 0.1–2.3 ng/µl; no competitive DNA was included. Probe and target sequences were simultaneously denatured by heating the covered slides to 80–85°C for 5 min on a hot plate. Hybridisation was carried out for 2 h or overnight (according to convenience) in a moist chamber at 37°C.
Post-Hybridisation and Microscopy
Post-hybridisation washes were performed at 45°C as described in Hultén et al. [10]. Slides which had been hybridised with biotin- or digoxigenin-labelled probes were taken through the conjugate/antibody detection procedures previously described [10, 11]. Directly labelled probes required no further processing. Prior to analysis the slides were stained with propidium iodide (PI) at 0.3 µg/ml in Vector Shield mounting medium.
The slides were screened using a × 100 objective on Nikon fluorescent microscopes with filter sets B1A for fluorescein/PI, DM400 for AMCA, and DM580 for rhodamine and PI. Photographs were taken using Fujichrome 400 ASA film rated at 800 ASA for AMCA and 1,600 ASA for fluorescein and rhodamine/PI. The film was developed at 400 ASA.
Scoring
Scoring of signals was adapted to overcome the following difficulties: (a) ensuring that signals at different focal points in the same cell were detected; (b) determining whether or not apparently aneuploid/diploid cells were in fact two cells in close proximity, and (c) deciding whether ‘twin’ signals represented two chromosomes or a split hybridisation site originating from one chromosome [12]. To help solve these problems, approximately 1,000 photographs were taken, i.e., representing at least one from each relevant fluorescence filter of any suspected abnormal nucleus, as well as a random sample of normals. Often photographs of more than one focal point were taken including some focused on the sperm tails, under the AMCA filter, which helped to delineate single nuclei. All photographs were reviewed independently by four of the authors. Aneuploid and diploid nuclei were then defined according to the agreement of all reviewers. Those with signals which could not be clearly defined, even by examination of photographs, were discounted from the data (134 out of over 60,000 scored).
Double signals of the same colour presented in four categories for which the following scoring rules were applied: (a) apparently two signals in contact with each other — counted as one chromosome; (b) two small signals clearly separated but by a distance shorter than the signal size itself — these were also counted as one chromosome but their frequency was recorded; (c) two large signals separated by a distance shorter than the signal size itself — counted as two chromosomes, and (d) two signals (large or small) widely separated — counted as two chromosomes.
Results
For the three donors D69, D117 and D218 the X and Y signals were counted in 10,240, 10,610 and 11,026 sperm, respectively (table 1). The ratio of single Y-:single X-bearing sperm displayed a slight excess of Xs in D69 (0.952), which was statistically significant (p < 0.05). In D117 and D218 the ratio was indistinguishable from unity (1,001 and 1,002, respectively). Overall the three donors revealed a small excess of X sperm with a Y:X ratio of 0.986, which was not statistically significant (p = 0.2). Including the aneuploid sperm with more than one X or Y (see below) does not significantly alter these ratios.
On average, 1/163 (6.12/1,000) sperm displayed more than one sex chromosome hybridisation signal (table 1). The majority of these (1/173 or 5.77/1,000) contained two target chromosomes representing XX, YY or XY sperm (fig. 1a–d). The remaining aneuploid sperm displayed three, (XXY, XYY) (fig. 1b) or four (XXYY) hybridisation signals at frequencies of 1/3,846 (0.26/1,000) and 1/11,111 (0.09/1,000), respectively. The frequency and type of sperm with ambiguous double sex chromosome signals are shown in table 2. These were comprised mainly of spurious XX and YY sperm, occurring at incidences of 1/2,857 (0.35/1,000) and 1/2,273 (0.44/1,000), respectively. Thus approximately 20% of the potentially XX and YY aneuploid sperm were excluded from the data. Also excluded were cells which were photographed but for which the hybridisation status was unclear. This was due mainly to diffuse signals, interfering crystals of fluorochrome or uncertainty regarding the overlapping of cells. There were 59 such cells excluded from a total of 31,876 between the three donors for slides hybridised for the sex chromosomes.
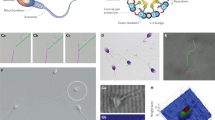
a–d Sperm hybridised with probes for both sex chromosomes seen through the AMCA fluorescence filter. The X probe (pSVX) was labelled with rhodamine-dUTP, the fluorescence of which can be seen in orange. The Y probe (pY10) was labelled with dig-dUTP and fluoresce blue after detection with AMCA-conjugated antibodies a Normal haploid sperm bearing either an X (arrow head) or a Y (short arrow) are seen in the same field as a sperm carrying both X and Y chromosomes (long arrow). b Two cells overlap making analysis difficult (arrow head) whilst one close by contains one X and two Y chromosomes (short arrow) and another has one of each chromosome (long arrow). c One cell has two X chromosomes (arrow heads). d A cell with two Y chromosomes is indicated (arrow head). e, f Sperm hybridised with probes for chromosomes 1 (pUC1.77), which was labelled with bio-dUTP detected with fluorescein, and chromosome 12 (pα12H8), which was labelled with AMCA-dUTP. The arrow heads indicate the same, diploid, cell with two fluorescent signals for both chromosomes 1 (e) and 12 (f). The haploid cell to the right of the field (short arrow) appears to be as large as the diploid.
Ploidy levels in the sperm were determined by dual hybridisations for chromosome 1 and 12 (fig. 1e, f). For the three donors D69, D117 and D218 signals for chromosomes 1 and 12 were counted in 12,005, 15,008 and 10,104 sperm, respectively. Each of the three individuals studied were found to have similar frequencies (p = 0.60) of diploid gametes with an average of 1/599 (1.67/1,000) (table 1). A total of 14 apparently diploid cells contained ambiguous twin signals for either chromosomes 1 or 12 (table 2). These represented 18% of the potentially diploid cells and were excluded from the calculations. A further 75 cells which were photographed could not be classified according to their ploidy due to the factors described above.
Two sex chromosome signals could represent either disomy or diploidy. The diploidy frequency, assessed by the analysis of chromosome 1 and 12 centromere probes, was used to calculate the proportion of sperm with two sex chromosome signals representing true disomics (table 1, 2). This was done by subtracting the diploid frequency from the sum of XX + YY + XY cells. Thus, on average we estimate 1/244 (4.10/1,000) sperm to be truly sex chromosome disomic (table 3).
On the assumption that each of the categories XX, YY and XY are equally likely to be diploid, the rate of disomy for each type was calculated from their relative frequencies in the initial data and the overall disomic rate (table 3). For example (taking the average results), XX represented 1.13/5.77 of the sperm with two sex chromosome signals, the overall rate of sex chromosome disomy was 4.10/1,000 so the adjusted disomic rate for XX was calculated to be 1.13/5.77 × 4.10, i.e., 0.80/1,000 or 1/1,250. The adjusted disomy frequencies calculated for YY and XY are 1.03/1,000 (1/971) and 2.27/1,000 (1/441), respectively.
The sperm which were found to have no hybridisation signal could represent either non-disjunctional events or failed hybridisations. The total number of hyperhaploid nuclei (after adjusting for diploidy) should correspond to the number of sex chromosome nullisomic gametes. Using the average data, the sum of XX + YY + XY + 2 × (XXY + XYY) is 4.62/1,000 (1/216) (from tables 1 and 3). The number of sperm with no hybridisation signal was 5.93/1,000 (1/169) (table 1).
Subtracting the expected frequency of null-signal sperm from the observed frequency gives an excess of 1.31/1,000 (1/763). Taking these cells to represent failed hybridisations, this is indicative of a hybridisation efficiency in excess of 99.8%.
Discussion
This study, using fluorescent chromosome-specific DNA markers in ejaculate sperm from three human male donors, had three main aims: (a) to identify the primary male gametic sex chromosome ratio; (b) to assess the occurrence of numerical sex chromosome abnormality, i.e., aneuploidy, and (c) to quantify the incidence of sperm having double the normal chromosome number, being diploid rather than haploid.
Sex Chromosome Ratio
We have found the Y:X ratio in more than 30,000 sperm from the three control subjects to be 0.986, revealing a slight excess of X-over Y-bearing sperm. A more significant excess of X sperm (with ratios as low as 0.8) has frequently been noted by sperm karyotyping following pseudo-fertilisation of hamster oocytes [13, 14]. The reason for this discrepancy remains unknown, however the statistical significance found for the deviation from unity in D69 indicates that interindividual variation may be a factor. Future studies will be required to determine if the close to unity ratios we have found arise in all situations for 46, XY males, as variation in sex ratio at birth indicates that this may not be the case [15–21].
Sex Chromosome Aneuploidy
We have determined the total incidence of sex chromosomal aneuploidy to be 1/112 (0.89%) which is a conservative estimate due to our stringent scoring protocol.
Disomy
Taking into consideration a diploidy rate of 1.67/1,000 (1/599) we have estimated the incidence of sperm truly disomic for the sex chromosomes to be 4.10/1,000 (1/244). This value is much higher than that either observed by sperm karyotyping [22] or postulated by Jacobs [23] based on knowledge of parental origin and the frequency of sex chromosome trisomy in clinically recognised pregnancies (fig. 2, 3).
Estimates of sex chromosomal trisomy rate based on our sperm data (black) and population studies (white). The expected frequencies were calculated from the current study assuming fetal loss rates of 30% for 47, XXX, 45% for 47, XXY, and 0% for 47, XYY [22]. The live-born data [36] have been adjusted on the assumption that 7% of 47, XXX and 49% of 47, XXY arise due to paternal error [37, 38].
Possible reasons for these discrepancies include selection operating against aneuploid sperm, either at fertilisation/pseudofertilisation [24] or early post-zygotically before clinical recognition of pregnancy. The presence of selection is indicated by our high numbers of YY sperm compared to the frequency expected assuming that 47, XYY fetuses are fully viable (fig. 3). The largest discrepancy with respect to Jacobs’ predictions concerns XX sperm (fig. 2). The uniform frequencies of XX and YY sperm found by karyotyping [22] indicate their equal ability to fertilise, at least in the human-hamster in vitro system. We therefore suggest that paternal X chromosomal disomy may be less viable, post-zygotically, than paternal Y disomy. Extra selection against two paternal X chromosomes may reflect either expression and/or imprinting of X-linked genes prior to inactivation [25–27].
Nullisomy
Based on the frequencies of sperm found to be sex chromosome hyperhaploid, we calculated the respective nullisomy rate to be 1/216. This is less than half the frequency of all clinically recognised pregnancies, suggested to have a 45.Xmat karyotype [28], lending support to the idea that a large proportion of 45, Xmat fetuses arise from post-zygotic exclusion of the paternal sex chromosome [29].
If preferential exclusion of paternal chromosomes affects the Xpat more often than the Y, then this, along with the 99% loss rate of 45, X fetuses [30], could be responsible for the slight excess of male births in the normal population [15]. If this is true, the frequently found 45, Xmat mosaics should more often have a second X chromosome than a Y. Selection bias associated with phenotypic differences would complicate the testing of this hypothesis.
Non-Disjunction
The frequency of XY gametes, determined to be 1/441, corresponds to a first meiotic non-disjunction rate of 1/220 meioses. Hassold et al. [31] have shown that the majority (33/39) of XXPatY offspring inherit non-recombinant Xpat and Y, providing evidence that first meiotic non-disjunction of X and Y arises due to failure of chiasma formation. On the other hand, the frequency of XY univalents at first meiosis is normally much higher than 1/220, being found in up to one third of first meiosis [32, 33]. We therefore believe that the XY bivalent regularly segregates earlier than the rest of the genome, so producing the majority of the XY univalents seen at first meiosis.
The second meiotic non-disjunction products XX and YY were found at the slightly different frequencies of 1/1,246 and 1/971, respectively. The total of 1/550 corresponds to rates of second meiotic non-disjunction of either 1/275 secondary spermatocytes or 1/138 complete meiotic divisions. When compared to the first meiotic non-disjunction rate of 1/220 this means that the sex chromosome bivalents in a primary spermatocyte non-disjoin approximately 25% more frequently than either X or Y in a secondary spermatocyte. On the other hand, in total, the second meiotic non-disjunction rate is more than 50% more common than the first.
On the basis of these figures, and assuming that the two segregational errors are independent, we calculate that approximately 8 per million sperm should be products of double non-disjunctional events. The frequency of such sperm (XXY + XYY) actually found was, however, more than 30-fold this value at 260 per million (0.26/1,000). Such an increase of observed over the expected frequencies of double meiotic non-disjunctions is consistent with the suggestion of Hassold et al. [34] that, for the sex chromosomes, a first meiotic non-disjunction event can be associated with an increased risk for a second meiotic non-disjunction. The mechanism for this remains unknown, but may well be related to an upset of the normal timing of meiosis affecting both first and second divisions.
Diploidy
The frequency of diploid gametes produced by the three subjects was determined by cohybridising with probes specific for chromosomes 1 and 12. On average we found 1/599 (1.67/1,000) sperm to yield two clearly separate signals from each probe with an interindividual range of 1.25–1.93/1,000. This is the first time that sperm diploidy has been assessed using dual hybridisations with autosomal probes. In previous studies, cell size has been used as an indicator of ploidy [8, 35]. Han et al. [8] recently reported an average diploidy rate of approximately 4/1,000 in 20,000 decondensed sperm from 13 men, with an individual range of 0–8.9/1,000. The discrepancy between the two reports may simply reflect normal variation. However, the size criterion may not be accurate, particularly since the decondensation protocol does not modify the size of all cells equally (fig. 1e, f) [4–6].
References
Barlow P, Vosa CG: The Y chromosome in human spermatozoa. Nature 1970;226:961–962
van Kooij RF, van Oost BA: Determination of sex ratio of spermatozoa with a deoxyribonucleic acid-probe and quinacrine staining: A comparison. Fertil Steril 1992;58:384–386
Martin RH, Balkan W, Burns K, Rademaker AW, Lin CC, Rudd NL: The chromosome consitution of 1,000 human spermatozoa. Hum Genet 1983;63:305–309
Wyrobeck AJ, Alhborn T, Balhorn R, Stanker L, Pinkel D: Fluorescence in situ hybridization to Y chromosomes in decondensed human sperm nuclei. Mol Reprod Dev 1990;27:200–208
Guttenbach M, Schmid M: Determination of Y chromosome aneuploidy in human sperm nuclei by nonradioactive in situ hybridization. Am J Hum Genet 1990;46:553–558
Coonen E, Pieters MHEC, Dumoulin JCM, Meyer H, Evers JLH, Radmaekers FCS, Geraedts JPS: Non-isotopic in situ hybridisation as a method for nondisjunction studies in human spermatazoa. Mol Reprod Dev 1991;28:18–22
Holmes JM, Martin RH: Aneuploidy detection in human sperm nuclei using fluorescence in situ hybridization. Hum Genet 1993;91:20–24
Han TL, Webb GC, Flaherty SP, Correll A, Matthews CD, Ford JD: Detection of chromosome 17- and X-bearing human spermatozoa using fluorescence in situ hybridization. Mol Reprod Dev 1992;33:189–194
Behrman SJ, Sawada Y: Heterologous and homologous inseminations with human semen frozen and stored in a liquid-nitrogen refrigerator. Fertil Steril 1966;17:457–466
Hultén MA, Goldman ASH, Saadallah N, Wallace BNM, Creasy MR: Meiotic investigations; in Rooney DE, Czepuldouski RH (eds): A Practical Approach. Oxford, IRL Press, 1992.
Goldman ASH, Hultén MA: Chromosome in situ suppression hybridisation in human male meiosis. J Med Genet 1992;29:101–105
Guttenbach M, Schmid M: Non-isotopic detection of chromosome 1 in human meiosis and demonstration of disomic sperm nuclei. Hum Genet 1991;87:261–265
Bean B: Progenitive sex ratio among functioning sperm cells. Am J Hum Genet 1990;47:352–353
Kamiguchi Y, Mikamo K: An improved, efficient method for analyzing human sperm chromosomes using zona-free hamster ova. Am J Hum Genet 1986;38:724–740
James WH: The human sex ratio. Part 1: A review of the literature. Hum Biol 1987;59:721–752
Astolfi P, Martinetti M, Gilgi-Berzolari F, Cuccua M: The effect of parental and maternal-fetal histocompatibility at MHC on sex ratio in offspring. Tissue Antigens 1990;35:172–177
Drickamer LC: Seasonal variation m fertility, fecundity and litter sex ratio in laboratory and wild stocks of house mice (Mus domesticus). Lab Anim Sci 1990;40:284–288
James WH: The hypothesized hormonal control of human sex ratio at birth — an update. J Theor Biol 1990;143:555–564
Hedricks C, McClintock MK: Timing of insemination is correlated with the secondary sex ratio of Norway rats. Physiol Behav 1990;48:625–632
Sapp MC, Martin-DeLeon PA: Sperm age, sex ratio, and hyperhaploidy frequency in mice. Cytogenet Cell Genet 1992;61:61–66
Bernstein ME: Variation of primary and secondary mammalian sex ratio (letter). Hum Biol 1990;62:437–439
Martin RH, Rademaker A: The frequency of aneuploidy among individual chromosomes in 6,821 human sperm chromosome complements. Cytogenet Cell Genet 1990;53:103–107
Jacobs PA: The role of chromosome abnormalities in reproductive failure. Reprod Nutr Dev 1990;supp11:63s–74s
Zackowski JL, Martin-DeLeon PA: Segregation products of male mice doubly heterozygous for the RB(6.16) and RB(16.17) translocations: Influence of sperm karyotype on fertilizing competence under varying mating frequencies. Gamete Res 1989;22:93–107
Ashworth A, Rastan S, Lovell-Badge R, Graham K: X-chromosome inactivation may explain the difference in viability of X0 humans and mice. Nature 1991;351:406–408
Driscoll DJ, Migeon BR: Sex difference in methylation of single-copy genes in human meiotic germ cells: Implications for X chromosome inactivation, parental imprinting, and origin of CpG mutations. Somat Cell Mol Genet 1990;16:267–282
Handel MA, Hunt PA: Genomic imprinting: Male mice with uniparentally derived sex chromosomes. Mol Reprod Dev 1991;28:337–340
Hassold T, Benham F, Leppert M: Cytogenetic and molecular analysis of sex chromosome monosomy. Am J Hum Genet 1988;42:534–541
Hassold T, Pettay D, Robinson A, Uchida I: Molecular studies of parental origin and mosaicism in 45, X conceptuses. Hum Genet 1992;89:647–652
Hook EB, Warburton D: The distribution of chromosomal genotypes associated with Turner’s syndrome: Live birth prevalence rates and evidence for diminished fetal mortality and severity in genotypes associated with structural X abnormalities. Hum Genet 1983;64:24–27
Hassold TJ, Sherman SL, Pettay D, Page DC, Jacobs PA: XY chromosome nondisjunction in man is associated with diminished recombination in the pseudoautosomal region. Am J Hum Genet 1991;49:252–260
Hultén M: Chiasma distribution at diakinesis in the normal human male. Hereditas 1974;76:55–78
Goldman ASH, Martin RH, Johannisson R, Gould CP, Davison V, Burn J, Hultén MA: Meiotic and sperm chromosome analysis in a male carrier of an inverted insertion (3;10)(q13.2;p14p13). J Med Genet 1992;29:460–464
Hassold T, Pettay D, May K, Robinson A: Analysis of non-disjunction in sex chromosome tetrasomy and pentasomy. Hum Genet 1990;85:648–650
Joseph AM, Gosden RJ, Chandley AC: Estimation of the aneuploidy levels in human spermatozoa using chromosome specific probes and in situ hybridization. Hum Genet 1984;66:234–238
Hook EB, Hamerton JL: The frequency of chromosome abnormalities detected in consecutive newborn studies: Differences between studies — results by sex and by severity of phenotypic involvement; in Hook EB, Porter IH (eds): Population Cytogenetics, Studies in Humans. New York, Academic Press, 1977, pp 73–79.
May KM, Jacobs PA, Lee M, Ratcliff S, Robinson A, Nielsen J, Hassold TJ: The parental origin of the extra X chromosome in 47, XXX females. Am J Hum Genet 1990;46:754–761
Harvey J, Jacobs P, Hassold T, Pettey D: The parental origin of 47, XXY males; in Evans JA, Hamerton JL (eds): Children and Young Adults with Sex Chromosome Aneuploidy. March of Dimes Birth Defects Orig Art Ser. New York, Wiley-Liss, 1990, vol 26, pp 297–304.
Acknowledgements
We would like to thank Dr. F. Granath, University of Stockholm, for carrying out the statistical analyses and advising us on their significance, and Prof. Letten F. Saugstad for providing a travel fellowship to ZF. We are also grateful to the staff of the Regional Genetics Services at East Birmingham Hospital for their general support. This project is supported by the UK Health and Safety Excutive.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Goldman, A.S.H., Fomina, Z., Knights, P.A. et al. Analysis of the Primary Sex Ratio, Sex Chromosome Aneuploidy and Diploidy in Human Sperm Using Dual-Colour Fluorescence in situ Hybridisation. Eur J Hum Genet 1, 325–334 (1993). https://doi.org/10.1159/000472431
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1159/000472431
Key Words
This article is cited by
-
A new approach of successful denaturation of human sperm chromatin without undergoing decondensation treatment
Genes & Genomics (2013)
-
Interphase cytogenetics in estimation of genomic mutations in somatic cells
Russian Journal of Genetics (2005)
-
Interphase cytogenetics in estimation of genomic mutations in somatic cells
Russian Journal of Genetics (2005)
-
Increased incidence of hyperhaploid 24,XY spermatozoa detected by three-colour FISH in a 46,XY/47,XXY male
Human Genetics (1996)
-
FISHing chromosomes in endocrinology
Endocrine (1996)