Featured
-
-
Article
| Open Access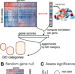 Overcoming false-positive gene-category enrichment in the analysis of spatially resolved transcriptomic brain atlas data
Overcoming false-positive gene-category enrichment in the analysis of spatially resolved transcriptomic brain atlas data
Identifying enriched gene sets in transcriptomic data is routine analysis. Here, the authors show that conventional gene category enrichment analysis (GCEA) applied to brain-wide atlas data yields biased results and develop a flexible ensemble-based null model framework to enable appropriate inference in GCEA.
- Ben D. Fulcher
- , Aurina Arnatkeviciute
- & Alex Fornito
-
Article
| Open Access Co-occupancy identifies transcription factor co-operation for axon growth
Co-occupancy identifies transcription factor co-operation for axon growth
After injury to the nervous system, many neurons fail to initiate transcriptional programs needed for axon growth. Here the authors examine co-operative binding of factors to regulatory DNA to predict combinations that improve axon growth when ectopically co-expressed.
- Ishwariya Venkatesh
- , Vatsal Mehra
- & Murray G. Blackmore
-
Article
| Open Access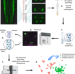 Single nucleus RNA-sequencing defines unexpected diversity of cholinergic neuron types in the adult mouse spinal cord
Single nucleus RNA-sequencing defines unexpected diversity of cholinergic neuron types in the adult mouse spinal cord
The full heterogeneity and different functional roles of cholinergic neurons in the adult spinal cord remain to be defined. Here the authors develop a targeted single nuclear RNA sequencing approach and use it to identify an array of cholinergic interneurons, as well as visceral and skeletal motor neurons.
- Mor R. Alkaslasi
- , Zoe E. Piccus
- & Claire E. Le Pichon
-
Article
| Open Access Learning cis-regulatory principles of ADAR-based RNA editing from CRISPR-mediated mutagenesis
Learning cis-regulatory principles of ADAR-based RNA editing from CRISPR-mediated mutagenesis
The RNA sequence and secondary structure regulate RNA editing by ADAR. Here the authors employ a CRISPR/Cas9-mediated saturation mutagenesis and machine learning to predict RNA editing efficiency of specific substrates.
- Xin Liu
- , Tao Sun
- & Jin Billy Li
-
Article
| Open Access Skeletal muscle transcriptome in healthy aging
Skeletal muscle transcriptome in healthy aging
As human skeletal muscle ages, gene expression programs change and reflect damage accumulation and homeostatic resilience mechanisms. Here, the authors present a detailed framework of the global transcriptome that characterizes skeletal muscle during aging in healthy individuals.
- Robert A. Tumasian III
- , Abhinav Harish
- & Luigi Ferrucci
-
Article
| Open Access Aptardi predicts polyadenylation sites in sample-specific transcriptomes using high-throughput RNA sequencing and DNA sequence
Aptardi predicts polyadenylation sites in sample-specific transcriptomes using high-throughput RNA sequencing and DNA sequence
Short read RNA sequencing and DNA sequence contain useful information for profiling polyadenylation sites, but each also possesses inherent limitations when examined independently. Aptardi combines these data and significantly improves annotation of polyadenylation sites in the expressed transcriptome.
- Ryan Lusk
- , Evan Stene
- & Laura M. Saba
-
Article
| Open Access Joint analysis of expression levels and histological images identifies genes associated with tissue morphology
Joint analysis of expression levels and histological images identifies genes associated with tissue morphology
Image features from histological slides can be used as informative endophenotypes in association studies for tissue-localized pathologies. Here, the authors develop ImageCCA, a framework for joint analysis of paired gene expression and histology data derived from automatically extracted image features.
- Jordan T. Ash
- , Gregory Darnell
- & Barbara E. Engelhardt
-
Article
| Open Access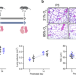 Single cell transcriptomic analysis of murine lung development on hyperoxia-induced damage
Single cell transcriptomic analysis of murine lung development on hyperoxia-induced damage
It is unclear how changes in gene expression are induced by changes in oxygen levels during late lung development. Here, the authors provide data from MULTI-seq scRNAseq in mice showing exposure to higher oxygen levels affects cell fates, especially for alveolarisation, and define gene/cell signatures of impaired lung development under hyperoxia.
- Maria Hurskainen
- , Ivana Mižíková
- & Bernard Thébaud
-
Article
| Open Access Loss of Quaking RNA binding protein disrupts the expression of genes associated with astrocyte maturation in mouse brain
Loss of Quaking RNA binding protein disrupts the expression of genes associated with astrocyte maturation in mouse brain
Quaking RNA binding protein (QKI) is known for its role in oligodendrocyte maturation. Here, the authors define the QKI targets in the mouse brain and show that loss of QKI disrupts the expression of cell maturation-associated genes in astrocytes in vivo.
- Kristina Sakers
- , Yating Liu
- & Joseph D. Dougherty
-
Article
| Open Access Single cell transcriptomics of primate sensory neurons identifies cell types associated with chronic pain
Single cell transcriptomics of primate sensory neurons identifies cell types associated with chronic pain
The contribution of distinct types of dorsal root ganglion neurons to chronic pain is unclear. Here, the authors molecularly profile non-human primate sensory neurons and show that genome-wide associations converge on two neuronal types with different genetic susceptibilities for chronic pain.
- Jussi Kupari
- , Dmitry Usoskin
- & Patrik Ernfors
-
Article
| Open Access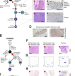 Somatic mutations and single-cell transcriptomes reveal the root of malignant rhabdoid tumours
Somatic mutations and single-cell transcriptomes reveal the root of malignant rhabdoid tumours
Malignant rhabdoid tumours (MRT) have been suggested to originate in the ectoderm-derived neural crest. Here, the authors analyse MRTs using phylogenetics, scRNA-seq, and patient-derived organoids; they find evidence for an MRT origin in the neural crest lineage and suggest differentiation treatment with HDAC/mTOR inhibitors.
- Lars Custers
- , Eleonora Khabirova
- & Jarno Drost
-
Article
| Open Access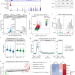 Multi-omics analysis reveals contextual tumor suppressive and oncogenic gene modules within the acute hypoxic response
Multi-omics analysis reveals contextual tumor suppressive and oncogenic gene modules within the acute hypoxic response
The response to hypoxia can significantly impact oncogenic processes. Here, the authors define the early transcriptional response to acute hypoxia and identify HIF1A target genes as part of this acute response, providing a resource for investigating context-dependent roles of HIF1A in the biology of cancer.
- Zdenek Andrysik
- , Heather Bender
- & Joaquin M. Espinosa
-
Article
| Open Access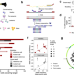 Identification of leukemic and pre-leukemic stem cells by clonal tracking from single-cell transcriptomics
Identification of leukemic and pre-leukemic stem cells by clonal tracking from single-cell transcriptomics
Leukaemic stem cells drive acute myeloid leukaemia (AML) progression and relapse but they are incompletely characterized. Here, the authors combine single-cell transcriptomics and clonal tracking using nuclear and mitochondrial somatic variants to distinguish healthy, pre-leukaemic and leukaemic stem cells in AML.
- Lars Velten
- , Benjamin A. Story
- & Lars M. Steinmetz
-
Article
| Open Access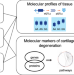 A molecular quantitative trait locus map for osteoarthritis
A molecular quantitative trait locus map for osteoarthritis
Understanding the molecular effects of disease variants in relevant tissues is essential to understanding and treating disease. Here, the authors discover expression and protein quantitative trait loci in cartilage and synovium from 115 osteoarthritis patients to pinpoint genes of action and potential drug treatments.
- Julia Steinberg
- , Lorraine Southam
- & Eleftheria Zeggini
-
Article
| Open Access Distinct subtypes of proprioceptive dorsal root ganglion neurons regulate adaptive proprioception in mice
Distinct subtypes of proprioceptive dorsal root ganglion neurons regulate adaptive proprioception in mice
Molecular diversity of proprioceptive neuron types (Ia, Ib and II PNs) is unclear. Here, the authors characterized the functional organization and development of eight subtypes of PNs in mice. Importantly, Ia subtypes are plastic, suggesting a role in adaptive proprioception during motor behavior.
- Haohao Wu
- , Charles Petitpré
- & François Lallemend
-
Article
| Open Access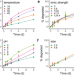 Photodegradation of carbon dots cause cytotoxicity
Photodegradation of carbon dots cause cytotoxicity
Carbon dots have attracted much attention for biomedical applications but potential degradation and associated toxicity are still poorly understood. Here, the authors report on a study into the photo-degradation of carbon dots, the products produced and associated cytotoxicity.
- Yue-Yue Liu
- , Nan-Yang Yu
- & Ai-Jun Miao
-
Article
| Open Access A practical solution to pseudoreplication bias in single-cell studies
A practical solution to pseudoreplication bias in single-cell studies
Single cell genomics uses cells from the same individual, or pseudoreplicates, that can introduce biases and inflate type I error rates. Here the authors apply generalized linear mixed models with a random effect for individual, to properly account for both zero inflation and the correlation structure among cells within an individual.
- Kip D. Zimmerman
- , Mark A. Espeland
- & Carl D. Langefeld
-
Article
| Open Access Identification and analysis of splicing quantitative trait loci across multiple tissues in the human genome
Identification and analysis of splicing quantitative trait loci across multiple tissues in the human genome
The profiling of genetic variants affecting splicing can give insight into disease mechanisms. Here, the authors develop a pipeline for discovery of variants affecting splicing (sQTLs) and with application to the GTEx dataset they generate a catalog of human sQTLs.
- Diego Garrido-Martín
- , Beatrice Borsari
- & Roderic Guigó
-
Article
| Open Access Uncovering de novo gene birth in yeast using deep transcriptomics
Uncovering de novo gene birth in yeast using deep transcriptomics
Genome-wide studies of de novo genes have tended to focus on genomic open reading frames (ORFs). Here, Blevins et al. use deep transcriptomics and synteny information to identify de novo transcripts in the yeast Saccharomyces cerevisiae, many of which are expressed from the alternative DNA strand.
- William R. Blevins
- , Jorge Ruiz-Orera
- & M. Mar Albà
-
Article
| Open Access RUNX1/RUNX1T1 mediates alternative splicing and reorganises the transcriptional landscape in leukemia
RUNX1/RUNX1T1 mediates alternative splicing and reorganises the transcriptional landscape in leukemia
The fusion gene RUNX1/RUNX1T1 is oncogenic in acute myeloid leukemia. Here, the authors show that the fusion gene alters the transcriptional landscape of the cells by changing the structure of the 5’UTR, altering isoform expression, and controlling the expression of splicing factors.
- Vasily V. Grinev
- , Farnaz Barneh
- & Olaf Heidenreich
-
Article
| Open Access A spatially resolved brain region- and cell type-specific isoform atlas of the postnatal mouse brain
A spatially resolved brain region- and cell type-specific isoform atlas of the postnatal mouse brain
Alternative RNA splicing varies across the brain. Its mapping at single cell resolution is unclear. Here, the authors provide a spatial and single-cell splicing atlas reporting brain region- and cell type-specific expression of different isoforms in the postnatal mouse brain.
- Anoushka Joglekar
- , Andrey Prjibelski
- & Hagen U. Tilgner
-
Article
| Open Access Core transcription regulatory circuitry orchestrates corneal epithelial homeostasis
Core transcription regulatory circuitry orchestrates corneal epithelial homeostasis
Corneal epithelium shares similar molecular signatures to other stratified epithelia. Here, the authors map super-enhancers and accessible chromatin in corneal epithelium, identifying a transcription regulatory circuit, including RUNX1, PAX6, and SMAD3, required for corneal epithelial identity and homeostasis.
- Mingsen Li
- , Huaxing Huang
- & Hong Ouyang
-
Article
| Open Access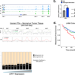 The circadian cryptochrome, CRY1, is a pro-tumorigenic factor that rhythmically modulates DNA repair
The circadian cryptochrome, CRY1, is a pro-tumorigenic factor that rhythmically modulates DNA repair
Cryptochrome 1 (CRY1) is a transcriptional coregulator associated with the circadian clock. Here the authors reveal that CRY1 is hormone-regulated, stabilized by genomic insult, and promotes DNA repair and cell survival through temporal transcriptional regulation.
- Ayesha A. Shafi
- , Chris M. McNair
- & Karen E. Knudsen
-
Article
| Open Access isoCirc catalogs full-length circular RNA isoforms in human transcriptomes
isoCirc catalogs full-length circular RNA isoforms in human transcriptomes
Circular RNAs have been identified using short-read RNA sequencing. Here, the authors report isoCirc, a long-read sequencing method to characterize full-length circRNA isoforms and generate a catalogue of full-length circRNA isoforms in 12 human tissues and one human cell line.
- Ruijiao Xin
- , Yan Gao
- & Yi Xing
-
Article
| Open Access Cellular and molecular landscape of mammalian sinoatrial node revealed by single-cell RNA sequencing
Cellular and molecular landscape of mammalian sinoatrial node revealed by single-cell RNA sequencing
The spontaneous bioelectrical activity of pacemaker cells in sinoatrial node (SAN) triggers the heartbeats. Here, the authors perform single-cell RNA sequencing in the mouse SAN and identify molecular and cellular features of the SAN conserved in rabbit and cynomolgus monkey, identifying a new potential SAN marker.
- Dandan Liang
- , Jinfeng Xue
- & Yi-Han Chen
-
Article
| Open Access Single-cell transcriptome profiling of the vaginal wall in women with severe anterior vaginal prolapse
Single-cell transcriptome profiling of the vaginal wall in women with severe anterior vaginal prolapse
Anterior vaginal prolapse (AVP), the most common form of pelvic organ prolapse, has deleterious effects on women’s health. Here the authors employ single-cell RNA-seq to construct a transcriptomic atlas of vaginal wall cells from AVP patients, and find that extracellular matrix dysregulation and immune reaction are associated with AVP.
- Yaqian Li
- , Qing-Yang Zhang
- & Lan Zhu
-
Article
| Open Access Transcriptional and morphological profiling of parvalbumin interneuron subpopulations in the mouse hippocampus
Transcriptional and morphological profiling of parvalbumin interneuron subpopulations in the mouse hippocampus
The relationship between gene expression and morphology to classify PV interneurons is unclear. Here, the authors show transcriptional continuity of morphologically distinct mouse hippocampal PV interneurons subtypes, combining single-cell RNA sequencing and electrophysiology.
- Lin Que
- , David Lukacsovich
- & Csaba Földy
-
Article
| Open Access Error correction enables use of Oxford Nanopore technology for reference-free transcriptome analysis
Error correction enables use of Oxford Nanopore technology for reference-free transcriptome analysis
Nanopore sequencing technologies applied to transcriptome analysis suffer from high error rates, limiting them largely to reference-based analyses. Here, the authors develop a computational error correction method for transcriptome analysis that reduces the median error rate from ~7% to ~1%.
- Kristoffer Sahlin
- & Paul Medvedev
-
Article
| Open Access RNA structure-wide discovery of functional interactions with multiplexed RNA motif library
RNA structure-wide discovery of functional interactions with multiplexed RNA motif library
Structured RNA motifs can be obtained by structure probing, duplex capture, and motif prediction. Here the authors develop a multiplexed affinity assay system to identify functional protein interactors from an RNA structure library with validated or predicted RNA motifs.
- Kaoru R. Komatsu
- , Toshiki Taya
- & Hirohide Saito
-
Article
| Open Access XBP1 links the 12-hour clock to NAFLD and regulation of membrane fluidity and lipid homeostasis
XBP1 links the 12-hour clock to NAFLD and regulation of membrane fluidity and lipid homeostasis
Hepatocyte 12-hour rhythms have a role in cellular stress and metabolic functions. Here, the authors demonstrate disrupting the 12-hour clock through deletion of XBP1 is associated with the development of NAFLD as well as disruption of phospholipid composition and the maintenance of lipid homeostasis.
- Huan Meng
- , Naomi M. Gonzales
- & Bert W. O’Malley
-
Article
| Open Access muscat detects subpopulation-specific state transitions from multi-sample multi-condition single-cell transcriptomics data
muscat detects subpopulation-specific state transitions from multi-sample multi-condition single-cell transcriptomics data
Single-cell transcriptomics enhanced our ability to profile heterogeneous cell populations. It is not known which statistical frameworks are performant to detect subpopulation-level responses. Here, the authors developed a simulation framework to evaluate various methods across a range of scenarios.
- Helena L. Crowell
- , Charlotte Soneson
- & Mark D. Robinson
-
Article
| Open Access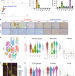 Functional compensation precedes recovery of tissue mass following acute liver injury
Functional compensation precedes recovery of tissue mass following acute liver injury
The liver possesses the ability to regenerate following sudden injury. Here, the authors use single-cell RNA-sequencing and in situ transcriptional analyses to identify a new phase of liver regeneration in mice aimed at maintaining essential functions throughout the regenerative process.
- Chad M. Walesky
- , Kellie E. Kolb
- & Wolfram Goessling
-
Article
| Open Access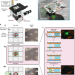 Digital microfluidic isolation of single cells for -Omics
Digital microfluidic isolation of single cells for -Omics
Multi-Omic approaches are a powerful way for obtaining in-depth understanding of a cell’s state. Here the authors present DISCO, combining digital microfluidics, laser cell lysis, and artificial intelligence-driven image processing to analyze single-cell genomes, transcriptomes and proteomes in a mixed population.
- Julian Lamanna
- , Erica Y. Scott
- & Aaron R. Wheeler
-
Article
| Open Access Benchmarking of cell type deconvolution pipelines for transcriptomics data
Benchmarking of cell type deconvolution pipelines for transcriptomics data
Inferring cell type proportions from transcriptomics data is affected by data transformation, normalization, choice of method and the markers used. Here, the authors use single-cell RNAseq datasets to evaluate the impact of these factors and propose guidelines to maximise deconvolution performance.
- Francisco Avila Cobos
- , José Alquicira-Hernandez
- & Katleen De Preter
-
Article
| Open Access Nuclear gene proximity and protein interactions shape transcript covariations in mammalian single cells
Nuclear gene proximity and protein interactions shape transcript covariations in mammalian single cells
Gene expression covariation can be studied by single-cell RNA sequencing. Here the authors analyze intrinsically covarying gene pairs by eliminating the confounding effects in single-cell experiments and observe covariation of proximal genes and miRNA-induced covariation of target mRNAs.
- Marcel Tarbier
- , Sebastian D. Mackowiak
- & Marc R. Friedländer
-
Article
| Open Access A multiresolution framework to characterize single-cell state landscapes
A multiresolution framework to characterize single-cell state landscapes
Dissecting the cellular heterogeneity embedded in single-cell transcriptomic data is challenging. Here, the authors introduce the concept of multiresolution cell-state decomposition as a practical approach to simultaneously capture both fine- and coarse-grain patterns of variability.
- Shahin Mohammadi
- , Jose Davila-Velderrain
- & Manolis Kellis
-
Article
| Open Access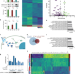 Gene expression and functional deficits underlie TREM2-knockout microglia responses in human models of Alzheimer’s disease
Gene expression and functional deficits underlie TREM2-knockout microglia responses in human models of Alzheimer’s disease
Mutations in TREM2 alter risk for Alzheimer’s disease, though the mechanisms underlying risk in human cells are unclear. Here, the authors use iPS-microglia and chimeric mice to highlight altered survival, phagocytosis, migration, and transcriptional programs in microglia lacking TREM2.
- Amanda McQuade
- , You Jung Kang
- & Mathew Blurton-Jones
-
Article
| Open Access Genome-wide translational profiling of amygdala Crh-expressing neurons reveals role for CREB in fear extinction learning
Genome-wide translational profiling of amygdala Crh-expressing neurons reveals role for CREB in fear extinction learning
Fear and fear extinction learning are dynamic. These dynamic changes are underlined by transcriptional changes. Here, the authors translationally profiled Crh neurons in the amygdala and and identified relevant gene networks.
- Kenneth M. McCullough
- , Chris Chatzinakos
- & Kerry J. Ressler
-
Article
| Open Access The structural variation landscape in 492 Atlantic salmon genomes
The structural variation landscape in 492 Atlantic salmon genomes
This study presents and validates a novel approach to reliably identify structural variations (SVs) in non-model genomes using whole genome sequencing, which was used to detect 15,483 SVs in 492 Atlantic salmon, shedding light on their roles in genome evolution and the genetic architecture of domestication.
- Alicia C. Bertolotti
- , Ryan M. Layer
- & Daniel J. Macqueen
-
Article
| Open Access Single-cell RNA cap and tail sequencing (scRCAT-seq) reveals subtype-specific isoforms differing in transcript demarcation
Single-cell RNA cap and tail sequencing (scRCAT-seq) reveals subtype-specific isoforms differing in transcript demarcation
Most single-cell RNA sequencing methods have limited ability to profile the transcriptome at isoform resolution. Here the authors develop scRCAT-seq to characterize the 5′- and 3′-ends of transcripts in single cells, and identify cell-type specific alternative transcript isoforms.
- Youjin Hu
- , Jiawei Zhong
- & Yizhi Liu
-
Article
| Open Access Type 2 and interferon inflammation regulate SARS-CoV-2 entry factor expression in the airway epithelium
Type 2 and interferon inflammation regulate SARS-CoV-2 entry factor expression in the airway epithelium
ACE2 and TMPRSS2 have received recent attention as entry factors for SARS-CoV-2. Here the authors analyze nasal airway transcriptome data from 695 children determining ACE2 and TMPRSS2 expression is induced by viral and type2 inflammation, respectively, and both exhibit eQTLs that vary across world populations.
- Satria P. Sajuthi
- , Peter DeFord
- & Max A. Seibold
-
Article
| Open Access Reconstructing the maize leaf regulatory network using ChIP-seq data of 104 transcription factors
Reconstructing the maize leaf regulatory network using ChIP-seq data of 104 transcription factors
Transcriptional factors (TFs) bind in a combinatorial fashion to specify the on-and-off states of genes in a complex and redundant regulatory network. Here, the authors construct the transcription regulatory network in maize leaf using 104 TFs ChIP-seq data and train machine learning models to predict TF binding and colocalization.
- Xiaoyu Tu
- , María Katherine Mejía-Guerra
- & Silin Zhong
-
Article
| Open Access Genetic circuit characterization by inferring RNA polymerase movement and ribosome usage
Genetic circuit characterization by inferring RNA polymerase movement and ribosome usage
Debugging a genetic circuit is frustrated by the inability to characterize parts in the context of the circuit. Here the authors use RNA-seq and ribosome profiling to take ‘snapshots’ of a large circuit in different states.
- Amin Espah Borujeni
- , Jing Zhang
- & Christopher A. Voigt
-
Article
| Open Access Cardiac dopamine D1 receptor triggers ventricular arrhythmia in chronic heart failure
Cardiac dopamine D1 receptor triggers ventricular arrhythmia in chronic heart failure
The pathophysiological role of dopamine D1 receptor (D1R) in chronic heart failure remains elusive. Here the authors show that D1R-expressing cardiomyocytes appear in chronic heart failure and play a pivotal role in triggering lethal ventricular arrhythmias.
- Toshihiro Yamaguchi
- , Tomokazu S. Sumida
- & Issei Komuro
-
Article
| Open Access Antibody-secreting cell destiny emerges during the initial stages of B-cell activation
Antibody-secreting cell destiny emerges during the initial stages of B-cell activation
The development of activated B cells into antibody-secreting cells (ASC) is a critical step for humoral immunity. Here the authors show, using adoptive transfers and single cell RNA sequencing, that commitment to ASC occurs soon following B cell activation, and is coordinated by specific transcriptome programs and proliferation kinetics.
- Christopher D. Scharer
- , Dillon G. Patterson
- & Jeremy M. Boss
-
Article
| Open Access Single-cell analysis uncovers fibroblast heterogeneity and criteria for fibroblast and mural cell identification and discrimination
Single-cell analysis uncovers fibroblast heterogeneity and criteria for fibroblast and mural cell identification and discrimination
To define and distinguish fibroblasts from vascular mural cells have remained challenging. Here, using single-cell RNA sequencing and tissue imaging, the authors provide a molecular basis for cell type classification and reveal inter- and intra-organ diversity of these cell types.
- Lars Muhl
- , Guillem Genové
- & Christer Betsholtz
-
Article
| Open Access Neonatal genetics of gene expression reveal potential origins of autoimmune and allergic disease risk
Neonatal genetics of gene expression reveal potential origins of autoimmune and allergic disease risk
Some immune-mediated diseases may originate in early childhood. The authors mapped eQTLs and response eQTLs to various stimuli in neonatal myeloid cells and T cells, and revealed their potential role in immune-mediated diseases using colocalisation and Mendelian randomisation.
- Qin Qin Huang
- , Howard H. F. Tang
- & Michael Inouye
-
Article
| Open Access 3D genome organization contributes to genome instability at fragile sites
3D genome organization contributes to genome instability at fragile sites
Common fragile sites are regions susceptible to replication stress and are prone to chromosomal instability. Here, the authors, by analyzing the contribution of 3D chromatin organization, identify and characterize a fragility signature and precisely map these fragility regions.
- Dan Sarni
- , Takayo Sasaki
- & Batsheva Kerem
-
Article
| Open Access PRDM15 is a key regulator of metabolism critical to sustain B-cell lymphomagenesis
PRDM15 is a key regulator of metabolism critical to sustain B-cell lymphomagenesis
The transcriptional regulator PRDM15 is expressed at low levels in normal tissues but overexpressed in B-cell lymphomas. Here, the authors show that PRDM15 depletion does not affect adult somatic cell homeostasis but leads to a metabolic crisis which impairs B-cell lymphomagenesis.
- Slim Mzoughi
- , Jia Yi Fong
- & Ernesto Guccione

 Neuronal genes deregulated in Cornelia de Lange Syndrome respond to removal and re-expression of cohesin
Neuronal genes deregulated in Cornelia de Lange Syndrome respond to removal and re-expression of cohesin