Featured
-
-
Article
| Open Access Single-cell transcriptome atlas of the human corpus cavernosum
Single-cell transcriptome atlas of the human corpus cavernosum
The corpus cavernosum is the most important structure for penile erection, and its dysfunction causes physiological and psychological problems. Here the authors perform single-cell RNA-sequencing on corpus cavernosum samples from males with normal erection and erectile dysfunction patients, providing insights into this pathology.
- LiangYu Zhao
- , Sha Han
- & Zheng Li
-
Article
| Open Access Construction of the axolotl cell landscape using combinatorial hybridization sequencing at single-cell resolution
Construction of the axolotl cell landscape using combinatorial hybridization sequencing at single-cell resolution
The Mexican axolotl is a well-established tetrapod model for regeneration and development. Here the authors report a scRNA-seq method to profile neotenic, metamorphic and limb development stages, highlighting unique perturbation patterns of cell type-related gene expression throughout metamorphosis.
- Fang Ye
- , Guodong Zhang
- & Guoji Guo
-
Article
| Open Access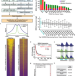 A multi-omic dissection of super-enhancer driven oncogenic gene expression programs in ovarian cancer
A multi-omic dissection of super-enhancer driven oncogenic gene expression programs in ovarian cancer
Super-enhancers and their associated transcription factor networks have been shown to influence ovarian cancer biology. Here, based on an integrated set of genomic and epigenomic datasets, the authors identify clinically relevant super-enhancers amplified in ovarian cancer patients and functionally validate their activity.
- Michael R. Kelly
- , Kamila Wisniewska
- & Hector L. Franco
-
Article
| Open Access Differential analysis of RNA structure probing experiments at nucleotide resolution: uncovering regulatory functions of RNA structure
Differential analysis of RNA structure probing experiments at nucleotide resolution: uncovering regulatory functions of RNA structure
The authors present DiffScan, an advanced tool for normalization and differential analysis of RNA structure probing experiments, combining their power in deciphering the dynamic RNA structurome and facilitating the discovery of RNA regulatory functions.
- Bo Yu
- , Pan Li
- & Lin Hou
-
Article
| Open Access Reactivity-dependent profiling of RNA 5-methylcytidine dioxygenases
Reactivity-dependent profiling of RNA 5-methylcytidine dioxygenases
Kleiner and co-workers profile RNA 5-methylcytidine (m5C) dioxygenase enzymes using an activity-based metabolic probing strategy. They reveal ALKBH1 as the major 5-formylcytidine (f5C) writer and characterize modification sites across mRNA and tRNA.
- A. Emilia Arguello
- , Ang Li
- & Ralph E. Kleiner
-
Article
| Open Access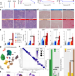 Single-cell analysis highlights differences in druggable pathways underlying adaptive or fibrotic kidney regeneration
Single-cell analysis highlights differences in druggable pathways underlying adaptive or fibrotic kidney regeneration
After acute injury, kidneys either successfully repair/regenerate or become fibrotic. Here the authors use scRNA-seq to study adaptive/maladaptive kidney regeneration and identify proinflammatory/fibrotic proximal tubule cells with pharmacologically targetable pyroptosis/ferroptosis signatures.
- Michael S. Balzer
- , Tomohito Doke
- & Katalin Susztak
-
Article
| Open Access Lipolysis regulates major transcriptional programs in brown adipocytes
Lipolysis regulates major transcriptional programs in brown adipocytes
β-Adrenergic signaling is a core regulator of brown adipocyte function. Here, the authors provide unbiased insight into the transcriptional network controlled by lipolysis in brown adipocytes, showing that lipolysis is required for much of the thermogenic gene program activated by β-adrenergic signals.
- Lasse K. Markussen
- , Elizabeth A. Rondini
- & Susanne Mandrup
-
Article
| Open Access Intestine-enriched apolipoprotein b orthologs are required for stem cell progeny differentiation and regeneration in planarians
Intestine-enriched apolipoprotein b orthologs are required for stem cell progeny differentiation and regeneration in planarians
Lipid metabolism regulates stem cell states and differentiation. Here, the authors demonstrate a requirement in planarians for Apolipoprotein B-mediated neutral lipid transport from intestinal stores to stem cells and their progeny during differentiation and whole-body regeneration.
- Lily L. Wong
- , Christina G. Bruxvoort
- & David J. Forsthoefel
-
Article
| Open Access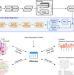 GenomicSuperSignature facilitates interpretation of RNA-seq experiments through robust, efficient comparison to public databases
GenomicSuperSignature facilitates interpretation of RNA-seq experiments through robust, efficient comparison to public databases
Many transcriptomic profiles have been deposited in public archives but are underused for the interpretation of experiments. Here the authors report GenomicSuperSignature for interpreting new transcriptomic datasets through comparison to public archives, without high-performance computing requirements.
- Sehyun Oh
- , Ludwig Geistlinger
- & Sean Davis
-
Article
| Open Access Complex regulation of Gephyrin splicing is a determinant of inhibitory postsynaptic diversity
Complex regulation of Gephyrin splicing is a determinant of inhibitory postsynaptic diversity
The protein gephyrin is involved in organizing synapses. Here, the authors show how different transcripts of gephyrin form and regulate inhibitory synapses.
- Raphaël Dos Reis
- , Etienne Kornobis
- & Eric Allemand
-
Article
| Open Access RIViT-seq enables systematic identification of regulons of transcriptional machineries
RIViT-seq enables systematic identification of regulons of transcriptional machineries
Here the authors present their method ‘regulon identification by in vitro transcription-sequencing’ (RIViT-seq), which enables systematic identification of target genes of transcription factors of interest. They applied RIViT-seq to 13 sigma factors from Streptomyces coelicolor A3(2) and successfully identified target genes of 11 of these, expanding the regulatory characterisation in this organism.
- Hiroshi Otani
- & Nigel J. Mouncey
-
Article
| Open Access Gene expression signatures of individual ductal carcinoma in situ lesions identify processes and biomarkers associated with progression towards invasive ductal carcinoma
Gene expression signatures of individual ductal carcinoma in situ lesions identify processes and biomarkers associated with progression towards invasive ductal carcinoma
Progression from ductal carcinoma in situ (DCIS) to invasive ductal carcinoma (IDC) remains poorly understood. Here, the authors analyse over 2700 micro-dissected samples using transcriptomics to identify genes that characterise different stages of DCIS to IDC progression, and identify IDC-associated markers within early-stage lesions.
- Clare A. Rebbeck
- , Jian Xian
- & Gregory J. Hannon
-
Article
| Open Access A single-cell transcriptomic atlas characterizes the silk-producing organ in the silkworm
A single-cell transcriptomic atlas characterizes the silk-producing organ in the silkworm
The molecular underpinning of silk-producing organs is not well characterized. Here the authors use single-cell RNA sequencing to build an atlas of the silkworm silk gland and reveal the heterogeneity of silk gland cells.
- Yan Ma
- , Wenhui Zeng
- & Hanfu Xu
-
Article
| Open Access Integrating 3D genomic and epigenomic data to enhance target gene discovery and drug repurposing in transcriptome-wide association studies
Integrating 3D genomic and epigenomic data to enhance target gene discovery and drug repurposing in transcriptome-wide association studies
Transcriptome-wide association studies can be used to test the effects of predicted gene expression in a cohort of individuals based on genetic data. Here, the authors developed a transcriptome-wide association method that integrates 3D genomic and epigenomic data with expression quantitative trait loci to improve gene expression predictions.
- Chachrit Khunsriraksakul
- , Daniel McGuire
- & Dajiang J. Liu
-
Article
| Open Access Mapping the cardiac vascular niche in heart failure
Mapping the cardiac vascular niche in heart failure
The cardiac vascular niche is of major importance in homeostasis and disease, but knowledge of its complexity in response to injury remains limited. Here we combine lineage tracing with single cell RNA sequencing to show alterations in fibroblasts, endothelial and mural cells in hypertrophic remodeling.
- Fabian Peisker
- , Maurice Halder
- & Rafael Kramann
-
Article
| Open Access Cellular and genetic drivers of RNA editing variation in the human brain
Cellular and genetic drivers of RNA editing variation in the human brain
Here the authors provide a deep catalogue of cell-specific A-to-I editing sites in the human cortex. Thousands of sites are enriched and elevated in neurons relative to glial cells, and are genetically regulated across multiple brain regions.
- Winston H. Cuddleston
- , Junhao Li
- & Michael S. Breen
-
Article
| Open Access Stochastic expression of invasion genes in Plasmodium falciparum schizonts
Stochastic expression of invasion genes in Plasmodium falciparum schizonts
Genetically identical cells can be phenotypically diverse to allow adaptive flexibility in a given environment. This phenotypic diversity is driven by epigenetic and transcriptional variability. Here, Tripathi et al. perform scRNA-seq of isogenic and non-isogenic Plasmodium falciparum schizont populations to explore transcriptional heterogeneity and stochastic gene expression during the course of development.
- Jaishree Tripathi
- , Lei Zhu
- & Zbynek Bozdech
-
Article
| Open Access SpotClean adjusts for spot swapping in spatial transcriptomics data
SpotClean adjusts for spot swapping in spatial transcriptomics data
Spatial transcriptomics experiments profile genome-wide gene expression at localized spots across a tissue. Here, the authors identify spot swapping, an artifact where RNA expressed at one tissue spot binds probes at another, and they propose SpotClean to adjust for it.
- Zijian Ni
- , Aman Prasad
- & Christina Kendziorski
-
Article
| Open Access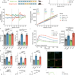 Multiomics reveals persistence of obesity-associated immune cell phenotypes in adipose tissue during weight loss and weight regain in mice
Multiomics reveals persistence of obesity-associated immune cell phenotypes in adipose tissue during weight loss and weight regain in mice
Adipose immune cells contribute to obesity-related disease, but less is known about weight cycling. Here, authors show that weight loss reduces diabetes risk, but inflammatory adipose immune cell populations persist and may contribute to worsened diabetes risk upon weight regain.
- Matthew A. Cottam
- , Heather L. Caslin
- & Alyssa H. Hasty
-
Article
| Open Access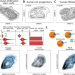 Cell cycle gene regulation dynamics revealed by RNA velocity and deep-learning
Cell cycle gene regulation dynamics revealed by RNA velocity and deep-learning
Single-cell RNA-sequencing technology gives access to cell cycle dynamics without externally perturbing the cell. Here the authors present DeepCycle,a robust deep learning method to infer the cell cycle state in single cells from scRNA-seq data.
- Andrea Riba
- , Attila Oravecz
- & Nacho Molina
-
Article
| Open Access Proteome allocations change linearly with the specific growth rate of Saccharomyces cerevisiae under glucose limitation
Proteome allocations change linearly with the specific growth rate of Saccharomyces cerevisiae under glucose limitation
Understanding how yeast organizes its functional proteome is a fundamental task in systems biology. Here, the authors conduct a multiomics analysis on yeast cells cultured with different growth rates, identifying a linear dependence of the functional proteome on the growth rate.
- Jianye Xia
- , Benjamin J. Sánchez
- & Jens Nielsen
-
Article
| Open Access A bio-functional polymer that prevents retinal scarring through modulation of NRF2 signalling pathway
A bio-functional polymer that prevents retinal scarring through modulation of NRF2 signalling pathway
One common cause of vision loss after retinal detachment surgery is retinal scarring. Here the authors demonstrate that a bio-functional polymer can prevent retinal scarring by suppression of epithelial-mesenchymal transition and hyper-proliferation of retinal pigment epithelial cells.
- Bhav Harshad Parikh
- , Zengping Liu
- & Xinyi Su
-
Article
| Open Access Extended intergenic DNA contributes to neuron-specific expression of neighboring genes in the mammalian nervous system
Extended intergenic DNA contributes to neuron-specific expression of neighboring genes in the mammalian nervous system
A large part of noncoding DNA is intergenic regions, yet how the size of intergenic regions affects gene expression in a tissue-specific manner is unclear. Here the authors present long intergenic DNA length-dependent neural gene expression, reflecting the complexity in the mammalian nervous system.
- Ravneet Jaura
- , Ssu-Yu Yeh
- & Ho Sung Rhee
-
Article
| Open Access Nuclear oligo hashing improves differential analysis of single-cell RNA-seq
Nuclear oligo hashing improves differential analysis of single-cell RNA-seq
Using spike-in controls with current single cell RNA-seq platforms remains a challenge. Here, the authors use a mixture of short, unmodified DNA oligos as a normalization standard for sci-RNAseq to improve the detection of global transcriptome changes.
- Hyeon-Jin Kim
- , Greg Booth
- & Cole Trapnell
-
Article
| Open Access Alternative splicing modulation by G-quadruplexes
Alternative splicing modulation by G-quadruplexes
Here the authors shows that G-quadruplexes, non-canonical DNA/RNA structures, can have a direct impact on alternative splicing and that binding of splicing regulators is affected by their presence.
- Ilias Georgakopoulos-Soares
- , Guillermo E. Parada
- & Martin Hemberg
-
Article
| Open Access Reference-free cell type deconvolution of multi-cellular pixel-resolution spatially resolved transcriptomics data
Reference-free cell type deconvolution of multi-cellular pixel-resolution spatially resolved transcriptomics data
Identifying cell-type-specific spatial patterns in ST data is critical for understanding tissue organization but current methods rely on external references. Here the authors develop a reference-free method to effectively recover cell-type transcriptional profiles and proportions.
- Brendan F. Miller
- , Feiyang Huang
- & Jean Fan
-
Article
| Open Access Single cell transcriptomic analysis reveals cellular diversity of murine esophageal epithelium
Single cell transcriptomic analysis reveals cellular diversity of murine esophageal epithelium
The level of cellular diversity in the esophageal epithelium has yet to be classified at the single cell level. Here the authors analyze the transcriptome of 44,679 murine esophageal keratinocytes to identify an unexpected level of cellular heterogeneity.
- Mohammad Faujul Kabir
- , Adam L. Karami
- & Kelly A. Whelan
-
Article
| Open Access Regulated interaction of ID2 with the anaphase-promoting complex links progression through mitosis with reactivation of cell-type-specific transcription
Regulated interaction of ID2 with the anaphase-promoting complex links progression through mitosis with reactivation of cell-type-specific transcription
Tissue-specific transcriptional activity is silenced in mitotic cells. Here the authors reveal a general phosphorylation-dependent mechanism of recognition for the anaphase-promoting complex (APC) substrates, and show that the APC targets ID2 during the establishment of post-mitotic transcription.
- Sang Bae Lee
- , Luciano Garofano
- & Anna Lasorella
-
Article
| Open Access Generation of human islet cell type-specific identity genesets
Generation of human islet cell type-specific identity genesets
Cell therapy to replace β-cells is a potential therapeutic avenue to treat diabetes, but the production of insulin-secreting replacement cells requires reliable tools to assess islet cellular identity. Here the authors use single-cell transcriptomics meta-analysis to construct gene sets that describe the identity of human α-, β-, γ- and δ-cells.
- Léon van Gurp
- , Leon Fodoulian
- & Pedro L. Herrera
-
Article
| Open Access Cooperative interaction between ERα and the EMT-inducer ZEB1 reprograms breast cancer cells for bone metastasis
Cooperative interaction between ERα and the EMT-inducer ZEB1 reprograms breast cancer cells for bone metastasis
The epithelial mesenchymal transition (EMT) is important in the metastatic spread of cancer cells. Here, the authors show that the EMT transcription factor, ZEB1, can modify estrogen receptor α during EMT and facilitate the migration of breast cancer cells to the bone
- Nastaran Mohammadi Ghahhari
- , Magdalena K. Sznurkowska
- & Didier Picard
-
Article
| Open Access Single-cell RNA sequencing coupled to TCR profiling of large granular lymphocyte leukemia T cells
Single-cell RNA sequencing coupled to TCR profiling of large granular lymphocyte leukemia T cells
T cell large granular lymphocyte leukemia (T-LGLL) and the cellular phenotype underlying response to therapy is not well understood. Here the authors use single cell sequencing to better understand changes in T cell clonal frequency and gene expression before and after therapy in T-LGLL.
- Shouguo Gao
- , Zhijie Wu
- & Neal S. Young
-
Article
| Open Access Cell-intrinsic Aryl Hydrocarbon Receptor signalling is required for the resolution of injury-induced colonic stem cells
Cell-intrinsic Aryl Hydrocarbon Receptor signalling is required for the resolution of injury-induced colonic stem cells
Rapid intestinal regeneration after injury is critical to maintain barrier integrity and homeostasis, but must be tightly controlled to prevent tumorigenesis. Here they show that the aryl hydrocarbon receptor is required to terminate the regenerative response after wound healing.
- Kathleen Shah
- , Muralidhara Rao Maradana
- & Brigitta Stockinger
-
Article
| Open Access Single-cell transcriptomics identifies potential cells of origin of MYC rhabdoid tumors
Single-cell transcriptomics identifies potential cells of origin of MYC rhabdoid tumors
Rhabdoid tumors (RT) are aggressive paediatric cancers with yet unknown cells of origin. Here, the authors establish genetically engineered mouse models of RT and, using single-cell RNA-seq and epigenomics, identify potential cells of origin for the SHH and MYC subtypes.
- Monika Graf
- , Marta Interlandi
- & Kornelius Kerl
-
Article
| Open Access Comprehensive evaluation of deconvolution methods for human brain gene expression
Comprehensive evaluation of deconvolution methods for human brain gene expression
Transcriptome deconvolution aims to estimate cellular composition based on gene expression data. Here the authors evaluate deconvolution methods for human brain transcriptome and conclude that partial deconvolution algorithms work best, but that appropriate cell-type signatures are also important.
- Gavin J. Sutton
- , Daniel Poppe
- & Irina Voineagu
-
Article
| Open Access Landscape of adenosine-to-inosine RNA recoding across human tissues
Landscape of adenosine-to-inosine RNA recoding across human tissues
Gabay et al. provide a highly-accurate atlas of recoding by A-to-I RNA editing in human, profiled across tissues and cell subpopulations. Most highly edited sites are evolutionary conserved in non-primate mammals, attesting for adaptation.
- Orshay Gabay
- , Yoav Shoshan
- & Eli Eisenberg
-
Article
| Open Access Modelling Chlamydia and HPV co-infection in patient-derived ectocervix organoids reveals distinct cellular reprogramming
Modelling Chlamydia and HPV co-infection in patient-derived ectocervix organoids reveals distinct cellular reprogramming
Here, Koster et al., model human papillomavirus and Chlamydia coinfection dynamics in patient-derived ectocervical organoids, and characterize the effects of multiple infections in the cellular microenvironment, potentially contributing to neoplasia.
- Stefanie Koster
- , Rajendra Kumar Gurumurthy
- & Cindrilla Chumduri
-
Article
| Open Access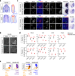 The histone demethylase Kdm6b regulates subtype diversification of mouse spinal motor neurons during development
The histone demethylase Kdm6b regulates subtype diversification of mouse spinal motor neurons during development
Neural cell type diversification during development is a complex and highly regulated process. Here, the authors show that the histone H3-lysine 27 demethylase Kdm6b promotes and inhibits the generation of specific motor neuron subtypes during the development of the mouse spinal cord.
- Wenxian Wang
- , Hyeyoung Cho
- & Soo-Kyung Lee
-
Article
| Open Access SM-Omics is an automated platform for high-throughput spatial multi-omics
SM-Omics is an automated platform for high-throughput spatial multi-omics
The spatial organisation of cells and molecules plays a key role in tissue function. Here the authors report Spatial MultiOmics (SM-Omics) as a fully automated, high-throughput all-sequencing based platform for combined and spatially resolved transcriptomics and antibody-based protein measurements.
- S. Vickovic
- , B. Lötstedt
- & A. Regev
-
Article
| Open Access Lymphocyte infiltration and thyrocyte destruction are driven by stromal and immune cell components in Hashimoto’s thyroiditis
Lymphocyte infiltration and thyrocyte destruction are driven by stromal and immune cell components in Hashimoto’s thyroiditis
Hashimoto’s Thyroiditis is an autoimmune disease with a complex pathomechanism. Authors here show by single cell RNA sequencing that the thyroidal microenvironment in the disease is characterised by three stromal cell subtypes that are potentially responsible for the recruitment of infiltrating inflammatory immune cells, such as macrophages and dendritic cells.
- Qian-Yue Zhang
- , Xiao-Ping Ye
- & Huai-Dong Song
-
Article
| Open Access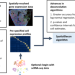 Advances in mixed cell deconvolution enable quantification of cell types in spatial transcriptomic data
Advances in mixed cell deconvolution enable quantification of cell types in spatial transcriptomic data
The deconvolution of cell types is challenging in spatially-resolved transcriptomics. Here, the authors present SpatialDecon, a method for the deconvolution and quantification of cell types in spatial transcriptomics data, and show how it can be used to analyse immune response heterogeneity in cancer.
- Patrick Danaher
- , Youngmi Kim
- & Joseph M. Beechem
-
Article
| Open Access Transcriptional programs regulating neuronal differentiation are disrupted in DLG2 knockout human embryonic stem cells and enriched for schizophrenia and related disorders risk variants
Transcriptional programs regulating neuronal differentiation are disrupted in DLG2 knockout human embryonic stem cells and enriched for schizophrenia and related disorders risk variants
Coordinated programs of gene expression drive brain development. Here, the authors use human embryonic stem cells and foetal cortical tissue as well as available GWAS statistics and analysis of genetic variants associated with neuropsychiatric disorders and cognition revealing a convergence on transcriptional programs regulating excitatory cortical neurogenesis.
- Bret Sanders
- , Daniel D’Andrea
- & Eunju Shin
-
Article
| Open Access Single cell transcriptomic landscape of diabetic foot ulcers
Single cell transcriptomic landscape of diabetic foot ulcers
Diabetic foot ulcers (DFUs) remain a complication of diabetes that are difficult to heal and lead to disability. Here the authors use single-cell RNA-sequencing and spatial transcriptomics to characterize the DFU cellular landscape and identify a population of fibroblasts that is associated with successful wound closure.
- Georgios Theocharidis
- , Beena E. Thomas
- & Manoj Bhasin
-
Article
| Open Access G-quadruplex DNA structures in human stem cells and differentiation
G-quadruplex DNA structures in human stem cells and differentiation
Whether G-quadruplexes (G4s) regulate stem cell self-renewal and fate determination during embryonic development is not well understood. Here, the authors reveal that the embryonic stem cell state is defined by very high G4 abundance. G4s are progressively lost during differentiation as cells transit to lower lineage potential while artificial G4 stabilisation leads to delayed differentiation.
- Katherine G. Zyner
- , Angela Simeone
- & Shankar Balasubramanian
-
Article
| Open Access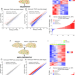 Reactive astrocytes acquire neuroprotective as well as deleterious signatures in response to Tau and Aß pathology
Reactive astrocytes acquire neuroprotective as well as deleterious signatures in response to Tau and Aß pathology
Alzheimer’s disease is associated with changes in astrocytes. Here the authors investigated the astrocyte translatome associated with amyloid-ß and tau pathology.
- Zoeb Jiwaji
- , Sachin S. Tiwari
- & Giles E. Hardingham
-
Article
| Open Access Integrated single-cell transcriptome analysis reveals heterogeneity of esophageal squamous cell carcinoma microenvironment
Integrated single-cell transcriptome analysis reveals heterogeneity of esophageal squamous cell carcinoma microenvironment
The microenvironment of oesophageal squamous cell carcinomas (ESCC) is heterogeneous and can strongly impact response to treatment. Here, the authors characterize the ESCC tumour microenvironment with single-cell RNA-seq, finding CST1 + myofibroblasts with potential biological and prognostic significance as well as immunosuppression signatures.
- Huy Q. Dinh
- , Feng Pan
- & En-Min Li
-
Article
| Open Access RNA modifications detection by comparative Nanopore direct RNA sequencing
RNA modifications detection by comparative Nanopore direct RNA sequencing
Nanopore direct RNA Sequencing data contain information about the presence of RNA modifications, but their detection poses substantial challenges. Here the authors introduce Nanocompore, a new methodology for modification detection from Nanopore data.
- Adrien Leger
- , Paulo P. Amaral
- & Tony Kouzarides
-
Article
| Open Access Single cell atlas for 11 non-model mammals, reptiles and birds
Single cell atlas for 11 non-model mammals, reptiles and birds
Here the authors report single-nucleus RNA sequencing for several anatomical locations in 11 species, including cat, dog, hamster, lizard, goat, rabbit, duck, pigeon, pangolin, tiger, and deer, highlighting coexpression of SARS-CoV-2 entry factors ACE2 and TMPRSS2.
- Dongsheng Chen
- , Jian Sun
- & Xun Xu
-
Article
| Open Access High-intensity training induces non-stoichiometric changes in the mitochondrial proteome of human skeletal muscle without reorganisation of respiratory chain content
High-intensity training induces non-stoichiometric changes in the mitochondrial proteome of human skeletal muscle without reorganisation of respiratory chain content
Exercise training can be therapeutic but how mitochondria respond remains unclear. Here, the authors use multiple omics techniques to reveal a complex network of non-stoichiometric mitochondrial adaptations that are prioritized or deprioritised during different phases of exercise training.
- Cesare Granata
- , Nikeisha J. Caruana
- & David J. Bishop
-
Article
| Open Access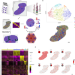 Spatial Transcriptomics to define transcriptional patterns of zonation and structural components in the mouse liver
Spatial Transcriptomics to define transcriptional patterns of zonation and structural components in the mouse liver
Global transcriptional differences across lobular units in the liver remain unknown. Here the authors perform spatial transcriptomics of liver tissue to delineate transcriptional differences in physical space, confirm lobular zonation along transcriptional gradients and suggest the presence of previously uncharacterized structures within liver tissue.
- Franziska Hildebrandt
- , Alma Andersson
- & Johan Ankarklev

 Combi-seq for multiplexed transcriptome-based profiling of drug combinations using deterministic barcoding in single-cell droplets
Combi-seq for multiplexed transcriptome-based profiling of drug combinations using deterministic barcoding in single-cell droplets