Featured
-
-
Article
| Open Access A molecular atlas reveals the tri-sectional spinning mechanism of spider dragline silk
A molecular atlas reveals the tri-sectional spinning mechanism of spider dragline silk
The genetic basis of spider major ampullate (Ma) gland silk production remains unknown. Hu et al. unveil a molecular atlas of this gland for the golden orb-weaving spider combining genome assembly and multiomics, revealing the single-cell spatial architecture of silk production in the Ma gland.
- Wenbo Hu
- , Anqiang Jia
- & Yi Wang
-
Article
| Open Access Lipid-induced transcriptomic changes in blood link to lipid metabolism and allergic response
Lipid-induced transcriptomic changes in blood link to lipid metabolism and allergic response
Circulating lipids can influence immune cell function, which could have implications for inflammatory diseases such as rheumatoid arthritis. Here, the authors use Mendelian randomization to identify genes whose expression is influenced by triglyceride levels in blood, implicating genes involved in lipid metabolism and allergic response.
- Koen F. Dekkers
- , Roderick C. Slieker
- & Bastiaan T. Heijmans
-
Article
| Open Access Correlated evolution of social organization and lifespan in mammals
Correlated evolution of social organization and lifespan in mammals
To elucidate the relationship between sociality and longevity, the authors perform phylogenetic and transcriptomic comparative analysis of mammals. They find that group-living species lived longer than solitary species and identify 31 genes, hormones, and immunity-related pathways involved in this connection.
- Pingfen Zhu
- , Weiqiang Liu
- & Xuming Zhou
-
Article
| Open Access Single-cell analysis reveals prognostic fibroblast subpopulations linked to molecular and immunological subtypes of lung cancer
Single-cell analysis reveals prognostic fibroblast subpopulations linked to molecular and immunological subtypes of lung cancer
Fibroblast heterogeneity is a prominent but poorly understood feature of solid tumours. Here three major fibroblast subpopulations in non-small cell lung cancer are identified and characterised through single cell RNA-sequencing, multiplexed immunohistochemistry and digital cytometry.
- Christopher J. Hanley
- , Sara Waise
- & Gareth J. Thomas
-
Article
| Open Access Spatially resolved transcriptomic profiling of degraded and challenging fresh frozen samples
Spatially resolved transcriptomic profiling of degraded and challenging fresh frozen samples
Spatial transcriptomics relies on RNA quality, which is variable and dependent on sample handling, storage, and/or intrinsic factors. Here, authors present a genome-wide spatial gene expression profiling method called RNA Rescue Spatial Transcriptomics (RRST), designed for the analysis of moderate to low quality fresh frozen tissue samples and demonstrate its robustness on 7 different tissue types.
- Reza Mirzazadeh
- , Zaneta Andrusivova
- & Joakim Lundeberg
-
Article
| Open Access scm6A-seq reveals single-cell landscapes of the dynamic m6A during oocyte maturation and early embryonic development
scm6A-seq reveals single-cell landscapes of the dynamic m6A during oocyte maturation and early embryonic development
Modification of RNA with N6-methyladenosine can regulate RNA metabolism. Here they developed scm6A-seq to profile the methylome and transcriptome in single cells, and reveal the functions of m6A modification during oocyte maturation and early embryo development.
- Huan Yao
- , Chun-Chun Gao
- & Yun-Gui Yang
-
Article
| Open Access Molecular characterization of Richter syndrome identifies de novo diffuse large B-cell lymphomas with poor prognosis
Molecular characterization of Richter syndrome identifies de novo diffuse large B-cell lymphomas with poor prognosis
Richter syndrome (RS) is the transformation of chronic lymphocytic leukaemia (CLL) into aggressive lymphoma, in most cases diffuse large B-cell lymphoma (DLBCL). Here, the authors characterize the DNA methylation and transcriptomic profiles of RS samples, find a clonally-related CLL epigenetic imprint, and develop classifiers for “RS-type” de novo DLBCLs.
- Julien Broséus
- , Sébastien Hergalant
- & Stephan Stilgenbauer
-
Article
| Open Access Semi-quantitative detection of pseudouridine modifications and type I/II hypermodifications in human mRNAs using direct long-read sequencing
Semi-quantitative detection of pseudouridine modifications and type I/II hypermodifications in human mRNAs using direct long-read sequencing
Pseudouridine (psi) is an RNA modification that can affect its physiology, including increased half-life. Here the authors identify sites of psi modification in the human transcriptome using direct RNA sequencing and provide a “ground truth” list of psi sites, sites of high psi occupancy, and transcripts that may be modified at multiple sites.
- Sepideh Tavakoli
- , Mohammad Nabizadeh
- & Sara H. Rouhanifard
-
Article
| Open Access Transcriptional vulnerabilities of striatal neurons in human and rodent models of Huntington’s disease
Transcriptional vulnerabilities of striatal neurons in human and rodent models of Huntington’s disease
In human and mouse models of Huntington’s disease, Matsushima, Pineda et al. show, using snRNAsequencing, the two axes defining identities of striatal projection neurons are multiplexed and differentially compromised, calling for distinct therapies.
- Ayano Matsushima
- , Sergio Sebastian Pineda
- & Ann M. Graybiel
-
Article
| Open Access Molecular subtypes of ALS are associated with differences in patient prognosis
Molecular subtypes of ALS are associated with differences in patient prognosis
Variability in ALS disease onset and progression are poorly understood. Our work identifies three distinct molecular states in post-mortem tissue that capture some of the observed differences in patient age of onset and survival.
- Jarrett Eshima
- , Samantha A. O’Connor
- & Barbara S. Smith
-
Article
| Open Access A comprehensive single-cell map of T cell exhaustion-associated immune environments in human breast cancer
A comprehensive single-cell map of T cell exhaustion-associated immune environments in human breast cancer
T cell exhaustion in breast tumours remains to be fully characterised. Here, single cell transcriptomics and imaging mass cytometry analysis of luminal breast tumours with or without exhausted T cells suggests distinct patterns of PD-1 and CXCL13 expression in T cells, and of MHC-I, but not PD-L1, expression in tumour cells.
- Sandra Tietscher
- , Johanna Wagner
- & Bernd Bodenmiller
-
Article
| Open Access Single-cell transcriptomic analysis reveals diversity within mammalian spinal motor neurons
Single-cell transcriptomic analysis reveals diversity within mammalian spinal motor neurons
How molecular diversity in neurons links to versatile functions is elusive. Here the authors profiled embryonic spinal motor neurons with single-cell RNAseq and identified molecular subtypes targeting distinct muscle groups in different species.
- Ee Shan Liau
- , Suoqin Jin
- & Jun-An Chen
-
Article
| Open Access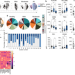 Transcriptional reprogramming from innate immune functions to a pro-thrombotic signature by monocytes in COVID-19
Transcriptional reprogramming from innate immune functions to a pro-thrombotic signature by monocytes in COVID-19
Although myeloid cell dysfunction has been observed in COVID-19, the underlying mechanisms remain incompletely understood. Here, the authors demonstrate that monocytes from patients with mild to moderate COVID-19 show a blunted innate immune response and a pro-thrombotic signature following secondary SARS-CoV-2 challenge.
- Allison K. Maher
- , Katie L. Burnham
- & Margarita Dominguez-Villar
-
Article
| Open Access Exon junction complex shapes the m6A epitranscriptome
Exon junction complex shapes the m6A epitranscriptome
Here the authors show the exon junction complex (EJC) component, EIF4A3, locally restricts METTL3- mediated mRNA methylation at exon junctions to explain the observed widespread enrichment of m6A modification in 3’ untranslated regions.
- Xin Yang
- , Robinson Triboulet
- & Richard I. Gregory
-
Article
| Open Access Single-cell profiling of healthy human kidney reveals features of sex-based transcriptional programs and tissue-specific immunity
Single-cell profiling of healthy human kidney reveals features of sex-based transcriptional programs and tissue-specific immunity
Knowledge of the transcriptional programs of human kidney cell populations at homeostasis is limited. Here, the authors show sex-based differences in gene expression of kidney parenchymal cells and examine the complexity of kidney-resident immune cells using single cell RNA sequencing of healthy living kidney donors.
- Caitriona M. McEvoy
- , Julia M. Murphy
- & Sarah Q. Crome
-
Article
| Open Access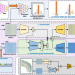 Spatial-ID: a cell typing method for spatially resolved transcriptomics via transfer learning and spatial embedding
Spatial-ID: a cell typing method for spatially resolved transcriptomics via transfer learning and spatial embedding
Comprehensive annotating of cell types in spatially resolved transcriptomics to understand biological processes at the single cell level remains challenging. Here the authors introduce Spatial-ID, a supervision-based cell typing method, that combines the existing knowledge of reference single-cell RNA-seq data and the spatial information of spatially resolved transcriptomics data.
- Rongbo Shen
- , Lin Liu
- & Jianhua Yao
-
Article
| Open Access The transcription factor DDIT3 is a potential driver of dyserythropoiesis in myelodysplastic syndromes
The transcription factor DDIT3 is a potential driver of dyserythropoiesis in myelodysplastic syndromes
Myelodysplastic syndromes (MDS) are age-related pathologies in which alterations of hematopoietic stem cells lead to abnormal formation of blood cells. Here, the authors study the lesions that these cells undergo in aging and disease, characterizing a factor whose alteration in MDS leads to abnormal blood cell production.
- Nerea Berastegui
- , Marina Ainciburu
- & Felipe Prosper
-
Article
| Open Access A unified computational framework for single-cell data integration with optimal transport
A unified computational framework for single-cell data integration with optimal transport
Integrating heterogeneous single-cell multi-omics as well as spatially resolved transcriptomic data remains a major challenge. Here the authors report a unified single-cell data integration framework using an unbalanced optimal transport-based deep network.
- Kai Cao
- , Qiyu Gong
- & Lin Wan
-
Article
| Open Access Altered tRNA processing is linked to a distinct and unusual La protein in Tetrahymena thermophila
Altered tRNA processing is linked to a distinct and unusual La protein in Tetrahymena thermophila
La proteins are conserved factors critical for the maturation of RNA polymerase III transcripts. In the ciliate T. thermophila and related alveolates, La proteins have a novel domain arrangement and are linked to a distinct pre-tRNA processing pathway.
- Kyra Kerkhofs
- , Jyoti Garg
- & Mark A. Bayfield
-
Article
| Open Access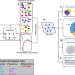 SOTIP is a versatile method for microenvironment modeling with spatial omics data
SOTIP is a versatile method for microenvironment modeling with spatial omics data
Methods that analyse heterogeneity and compare tissue microenvironments using spatial omics data are challenging to develop. Here, the authors present SOTIP, a method that can perform spatial heterogeneity, spatial domain, and differential microenvironment analyses across multiple spatial omics modalities.
- Zhiyuan Yuan
- , Yisi Li
- & Michael Q. Zhang
-
Article
| Open Access A single-cell analysis reveals tumor heterogeneity and immune environment of acral melanoma
A single-cell analysis reveals tumor heterogeneity and immune environment of acral melanoma
Studying the cell composition of acral melanoma at the single-cell level could provide some clues about its poor response to immunotherapy. Here, the authors analyse acral and cutaneous melanoma patient samples using single-cell RNA-sequencing, and reveal a severe immunosuppressive state in acral melanomas
- Chao Zhang
- , Hongru Shen
- & Jilong Yang
-
Article
| Open Access Postnatal expansion of mesenteric lymph node stromal cells towards reticular and CD34+ stromal cell subsets
Postnatal expansion of mesenteric lymph node stromal cells towards reticular and CD34+ stromal cell subsets
Lymph nodes in various locations of the body differ in their cell composition and gene expression signatures. Here authors show that the rapid postnatal expansion of lymph nodes is governed by CD34 + stromal cells and fibroblastic reticular stromal cell progenitors, distinguished by intrinsic, microbiome-independent core epigenetic blueprints.
- Joern Pezoldt
- , Carolin Wiechers
- & Jochen Huehn
-
Article
| Open Access Spatially aware dimension reduction for spatial transcriptomics
Spatially aware dimension reduction for spatial transcriptomics
Spatial transcriptomics analyses can be affected by noise and spatial correlation across tissue locations. Here, the authors develop SpatialPCA, a spatially-aware dimensionality reduction method that explicitly models spatial correlation structures, and show its application to the analysis of healthy and tumour tissues.
- Lulu Shang
- & Xiang Zhou
-
Article
| Open Access Genomic signatures of recent convergent transitions to social life in spiders
Genomic signatures of recent convergent transitions to social life in spiders
Sociality has evolved repeatedly in arthropods. Tong et al. compare the genomes of 22 spider species with a range of social complexity and eight independent origins of sociality, and identify specific genetic changes associated with the evolution of sociality in spiders.
- Chao Tong
- , Leticia Avilés
- & Timothy A. Linksvayer
-
Article
| Open Access A non-coding GWAS variant impacts anthracycline-induced cardiotoxic phenotypes in human iPSC-derived cardiomyocytes
A non-coding GWAS variant impacts anthracycline-induced cardiotoxic phenotypes in human iPSC-derived cardiomyocytes
Germline variants may pre-dispose patients to an increased risk of developing anthracycline-induced cardiotoxicity. This report provides insights into the mechanism by which a common genetic variant, rs28714259, may confer an increased risk of cardiac damage.
- Xi Wu
- , Fei Shen
- & Bryan Paul Schneider
-
Article
| Open Access CLIMB: High-dimensional association detection in large scale genomic data
CLIMB: High-dimensional association detection in large scale genomic data
Comparisons among experimental results with large amounts of data can be more precise and meaningful when done across multiple different conditions simultaneously. Koch et al. introduce a method, called CLIMB, that does this, and captures interpretable and biologically meaningful information.
- Hillary Koch
- , Cheryl A. Keller
- & Qunhua Li
-
Article
| Open Access A flexible cross-platform single-cell data processing pipeline
A flexible cross-platform single-cell data processing pipeline
As the throughput of single-cell RNA-seq studies increases, there is a need for tools that can make the data analysis steps more streamlined and convenient. Here, the authors develop UniverSC, a tool that unifies single-cell RNA-seq analysis workflows and also facilitates their use for non-experts.
- Kai Battenberg
- , S. Thomas Kelly
- & Aki Minoda
-
Article
| Open Access Systematic characterization of cancer transcriptome at transcript resolution
Systematic characterization of cancer transcriptome at transcript resolution
Modification of transcribed mRNAs enables regulation of transcription but its extent in cancer cells is incompletely understood. Here, the authors analyse transcript assembly in over 1000 cancer cell lines and find unannotated transcripts are common, and are associated with drug sensitivity.
- Wei Hu
- , Yangjun Wu
- & Shengli Li
-
Article
| Open Access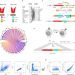 speedingCARs: accelerating the engineering of CAR T cells by signaling domain shuffling and single-cell sequencing
speedingCARs: accelerating the engineering of CAR T cells by signaling domain shuffling and single-cell sequencing
Chimeric antigen receptors (CAR) are a promising option for cell-based immunotherapy for cancer and other immune diseases. Here the authors develop speedingCARs, an integrated CAR design and screening platform based on modular signaling domain shuffling and single cell transcriptomic analyses, and test its potential for identifying and validating novel CAR designs.
- Rocío Castellanos-Rueda
- , Raphaël B. Di Roberto
- & Sai T. Reddy
-
Article
| Open Access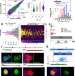 Translation and natural selection of micropeptides from long non-canonical RNAs
Translation and natural selection of micropeptides from long non-canonical RNAs
Translation of 100 to 300 micropeptides from small ORFs within lncRNA was detected by Ribosomal Profiling in Drosophila embryos. These translated small ORFs showed natural selection conserving micropeptide sequence and function.
- Pedro Patraquim
- , Emile G. Magny
- & Juan Pablo Couso
-
Article
| Open Access De novo analysis of bulk RNA-seq data at spatially resolved single-cell resolution
De novo analysis of bulk RNA-seq data at spatially resolved single-cell resolution
Current methods to reanalyze bulk RNA-seq at spatially resolved single-cell resolution have limitations. Here, the authors develop Bulk2Space, a spatial deconvolution algorithm using single-cell and spatial transcriptomics as references, providing new insights into spatial heterogeneity within bulk tissue.
- Jie Liao
- , Jingyang Qian
- & Xiaohui Fan
-
Article
| Open Access SUMMIT: An integrative approach for better transcriptomic data imputation improves causal gene identification
SUMMIT: An integrative approach for better transcriptomic data imputation improves causal gene identification
Genes with moderate-low expression heritability cannot be sufficiently captured with conventional TWAS. This study introduces a new method, Summary-level Unified Method for Modeling Integrated Transcriptome (SUMMIT), to improve the expression prediction of TWAS by using eQTL summary-level data.
- Zichen Zhang
- , Ye Eun Bae
- & Chong Wu
-
Article
| Open Access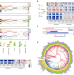 RNA G-quadruplex structure contributes to cold adaptation in plants
RNA G-quadruplex structure contributes to cold adaptation in plants
During evolution, plants have adapted to habitats with distinct temperature ranges. In this study, scientists report that a specific RNA structure motif, RNA G-quadruplex (RG4) is enriched across genomes of plant species growing in colder climates.
- Xiaofei Yang
- , Haopeng Yu
- & Yiliang Ding
-
Article
| Open Access The genome and lifestage-specific transcriptomes of a plant-parasitic nematode and its host reveal susceptibility genes involved in trans-kingdom synthesis of vitamin B5
The genome and lifestage-specific transcriptomes of a plant-parasitic nematode and its host reveal susceptibility genes involved in trans-kingdom synthesis of vitamin B5
Plant-parasitic nematodes are a threat to crop production. Combining bioinformatics, genetic and biochemical approaches, the authors show that the plant pathogen beet cyst nematode possesses an incomplete vitamin B5 synthesis pathway, of potential prokaryotic origin, complemented by its plant host.
- Shahid Siddique
- , Zoran S. Radakovic
- & Sebastian Eves-van den Akker
-
Article
| Open Access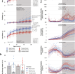 Chronic intake of high dietary sucrose induces sexually dimorphic metabolic adaptations in mouse liver and adipose tissue
Chronic intake of high dietary sucrose induces sexually dimorphic metabolic adaptations in mouse liver and adipose tissue
Dietary sugar intake may contribute to the development non-alcoholic fatty liver disease. Here the authors investigated the effects of chronic dietary sucrose on the liver-adipose-microbiome axis in mice, and report that sex is a moderating factor that influences sucrose-driven lipid storage in the liver and adipose tissue lipolysis.
- Erin J. Stephenson
- , Amanda S. Stayton
- & Joan C. Han
-
Article
| Open Access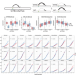 Tissue-specific impacts of aging and genetics on gene expression patterns in humans
Tissue-specific impacts of aging and genetics on gene expression patterns in humans
Age is a risk factor for many diseases, but the impact of aging on molecular phenotypes is not fully understood. Here, the authors quantify the relative contributions of genetics and aging to gene expression patterns across 27 tissues in humans, showing that age and genetics each play distinct roles in shaping expression phenotypes.
- Ryo Yamamoto
- , Ryan Chung
- & Peter H. Sudmant
-
Article
| Open Access Adar-mediated A-to-I editing is required for embryonic patterning and innate immune response regulation in zebrafish
Adar-mediated A-to-I editing is required for embryonic patterning and innate immune response regulation in zebrafish
Additional roles for A-to-I editing of RNA continue to be uncovered. Niescierowicz et al. report prevalent A-to-I editing in the zebrafish transcriptome, and the distinct maternal and zygotic functions of the editing enzyme Adar in embryonic patterning and in the regulation of innate immune response, respectively.
- Katarzyna Niescierowicz
- , Leszek Pryszcz
- & Cecilia Winata
-
Article
| Open Access Spatio-temporal analysis of prostate tumors in situ suggests pre-existence of treatment-resistant clones
Spatio-temporal analysis of prostate tumors in situ suggests pre-existence of treatment-resistant clones
Spatial heterogeneity in prostate cancer can contribute to its resistance to androgen deprivation therapy (ADT). Here, the authors analyse prostate cancer samples before and after ADT using Spatial Transcriptomics, and find heterogeneous pre-treatment tumour cell populations and stromal cells that are associated with resistance.
- Maja Marklund
- , Niklas Schultz
- & Joakim Lundeberg
-
Article
| Open Access Mettl3-dependent m6A modification attenuates the brain stress response in Drosophila
Mettl3-dependent m6A modification attenuates the brain stress response in Drosophila
The brain is vulnerable to stress and disease, with much work focused on defining mechanisms that impact the brain’s resilience. Here the author’s reveal in Drosophila that m6A epitranscriptomic modification of RNA dampens the brain’s capacity to mitigate stress by regulating RNA stability and translation.
- Alexandra E. Perlegos
- , Emily J. Shields
- & Nancy M. Bonini
-
Article
| Open Access Splicing QTL analysis focusing on coding sequences reveals mechanisms for disease susceptibility loci
Splicing QTL analysis focusing on coding sequences reveals mechanisms for disease susceptibility loci
Splicing QTL (sQTL), genetic variants regulating alternative splicing, can be biologically important, but complex to detect and interpret. Here, the authors identify sQTL by focusing on protein coding sequences, as an alternative to junction-based approaches.
- Kensuke Yamaguchi
- , Kazuyoshi Ishigaki
- & Yuta Kochi
-
Article
| Open Access The whole blood transcriptional regulation landscape in 465 COVID-19 infected samples from Japan COVID-19 Task Force
The whole blood transcriptional regulation landscape in 465 COVID-19 infected samples from Japan COVID-19 Task Force
Genetic mechanisms influencing COVID-19 susceptibility are not well understood. Here, the authors analyzed whole blood RNA-seq data of 465 Japanese individuals with COVID-19, highlighting thousands of fine-mapped variants affecting expression and splicing of genes, as well as the presence of COVID-19 severity-interaction eQTLs.
- Qingbo S. Wang
- , Ryuya Edahiro
- & Yukinori Okada
-
Article
| Open Access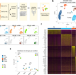 Single cell transcriptomic analysis of the immune cell compartment in the human small intestine and in Celiac disease
Single cell transcriptomic analysis of the immune cell compartment in the human small intestine and in Celiac disease
Celiac disease is linked to responsiveness to dietary gluten, which manifests itself as immune cell activation and the immunopathology including destruction of the epithelium of the small intestine. Here the authors apply single cell transcriptomics to characterise the immune cell compartment of the human small intestine during active Celiac disease.
- Nader Atlasy
- , Anna Bujko
- & Hendrik G. Stunnenberg
-
Article
| Open Access Spatiotemporal dynamics of macrophage heterogeneity and a potential function of Trem2hi macrophages in infarcted hearts
Spatiotemporal dynamics of macrophage heterogeneity and a potential function of Trem2hi macrophages in infarcted hearts
Cellular composition and function are not clearly defined in heart failure after myocardial infarction. Here, using single cell and spatial transcriptomics in a MI-HF mouse model, the authors show that macrophages expressing Trem2 are found within the infarcts and this could be a useful biomarker.
- Seung-Hyun Jung
- , Byung-Hee Hwang
- & Yeun-Jun Chung
-
Article
| Open Access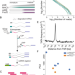 Transcriptomic diversity in human medullary thymic epithelial cells
Transcriptomic diversity in human medullary thymic epithelial cells
The thymus generates all T cells, including those that underly autoimmune diseases. Here, using deep sequencing, the authors profile human medullary thymic epithelial cells and establish a web portal to query their transcriptome, which may serve as a tool to help identify the drivers of autoimmunity.
- Jason A. Carter
- , Léonie Strömich
- & Hannah V. Meyer
-
Article
| Open Access Combi-seq for multiplexed transcriptome-based profiling of drug combinations using deterministic barcoding in single-cell droplets
Combi-seq for multiplexed transcriptome-based profiling of drug combinations using deterministic barcoding in single-cell droplets
Current screens to assess tumour drug resistance require a large amount of material, normally not available from patients. Here the authors report CombiSeq, a scalable microfluidic workflow to screen hundreds of drug combinations in picoliter-size droplets using transcriptome changes as a readout.
- L. Mathur
- , B. Szalai
- & C. A. Merten
-
Article
| Open Access Single-cell transcriptome atlas of the human corpus cavernosum
Single-cell transcriptome atlas of the human corpus cavernosum
The corpus cavernosum is the most important structure for penile erection, and its dysfunction causes physiological and psychological problems. Here the authors perform single-cell RNA-sequencing on corpus cavernosum samples from males with normal erection and erectile dysfunction patients, providing insights into this pathology.
- LiangYu Zhao
- , Sha Han
- & Zheng Li
-
Article
| Open Access Construction of the axolotl cell landscape using combinatorial hybridization sequencing at single-cell resolution
Construction of the axolotl cell landscape using combinatorial hybridization sequencing at single-cell resolution
The Mexican axolotl is a well-established tetrapod model for regeneration and development. Here the authors report a scRNA-seq method to profile neotenic, metamorphic and limb development stages, highlighting unique perturbation patterns of cell type-related gene expression throughout metamorphosis.
- Fang Ye
- , Guodong Zhang
- & Guoji Guo
-
Article
| Open Access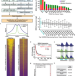 A multi-omic dissection of super-enhancer driven oncogenic gene expression programs in ovarian cancer
A multi-omic dissection of super-enhancer driven oncogenic gene expression programs in ovarian cancer
Super-enhancers and their associated transcription factor networks have been shown to influence ovarian cancer biology. Here, based on an integrated set of genomic and epigenomic datasets, the authors identify clinically relevant super-enhancers amplified in ovarian cancer patients and functionally validate their activity.
- Michael R. Kelly
- , Kamila Wisniewska
- & Hector L. Franco
-
Article
| Open Access Differential analysis of RNA structure probing experiments at nucleotide resolution: uncovering regulatory functions of RNA structure
Differential analysis of RNA structure probing experiments at nucleotide resolution: uncovering regulatory functions of RNA structure
The authors present DiffScan, an advanced tool for normalization and differential analysis of RNA structure probing experiments, combining their power in deciphering the dynamic RNA structurome and facilitating the discovery of RNA regulatory functions.
- Bo Yu
- , Pan Li
- & Lin Hou

 Differential regulation of mRNA stability modulates transcriptional memory and facilitates environmental adaptation
Differential regulation of mRNA stability modulates transcriptional memory and facilitates environmental adaptation