Research Highlight |
Featured
-
-
News & Views |
 The promise and peril of deep learning in microscopy
The promise and peril of deep learning in microscopy
- David P. Hoffman
- , Isaac Slavitt
- & Casey A. Fitzpatrick
-
Article |
 Evaluation and development of deep neural networks for image super-resolution in optical microscopy
Evaluation and development of deep neural networks for image super-resolution in optical microscopy
This study explores the performance of deep-learning models for super-resolution imaging and introduces models that utilize frequency content information in the Fourier domain to improve SIM reconstruction under low-SNR conditions.
- Chang Qiao
- , Di Li
- & Dong Li
-
Article |
 Click-ExM enables expansion microscopy for all biomolecules
Click-ExM enables expansion microscopy for all biomolecules
Click-ExM uses click-chemistry-based labeling to increase the versatility of expansion microscopy. Click-ExM enables imaging of numerous classes of biomolecules including lipids, glycans, proteins, DNA, RNA and small molecules.
- De-en Sun
- , Xinqi Fan
- & Xing Chen
-
Article |
 Live-cell super-resolved PAINT imaging of piconewton cellular traction forces
Live-cell super-resolved PAINT imaging of piconewton cellular traction forces
Tension-PAINT integrates molecular tension probes with DNA-PAINT to enable ~25-nm-resolution mapping of piconewton mechanical events. Tension-PAINT can be used to study dynamic forces, and an irreversible variant integrates force history over time.
- Joshua M. Brockman
- , Hanquan Su
- & Khalid Salaita
-
Correspondence |
 SMAP: a modular super-resolution microscopy analysis platform for SMLM data
SMAP: a modular super-resolution microscopy analysis platform for SMLM data
- Jonas Ries
-
Brief Communication |
 Optimizing imaging speed and excitation intensity for single-molecule localization microscopy
Optimizing imaging speed and excitation intensity for single-molecule localization microscopy
A systematic evaluation shows that excitation intensity has a dramatic impact on image quality in localization microscopy and reveals the benefits of lower excitation intensity for improved labeling efficiency and localization precision.
- Robin Diekmann
- , Maurice Kahnwald
- & Jonas Ries
-
-
Brief Communication |
 Up to 100-fold speed-up and multiplexing in optimized DNA-PAINT
Up to 100-fold speed-up and multiplexing in optimized DNA-PAINT
Hundred-fold-faster DNA-PAINT imaging is enabled by the introduction of concatenated, periodic DNA sequence motifs in the docking strand. Six orthogonal sequences are described for speed-optimized and highly multiplexed cellular imaging.
- Sebastian Strauss
- & Ralf Jungmann
-
Article |
 DeepSTORM3D: dense 3D localization microscopy and PSF design by deep learning
DeepSTORM3D: dense 3D localization microscopy and PSF design by deep learning
DeepSTORM3D uses deep learning for accurate localization of point emitters in densely labeled samples in three dimensions for volumetric localization microscopy with high temporal resolution, as well as for optimal point-spread function design.
- Elias Nehme
- , Daniel Freedman
- & Yoav Shechtman
-
Article |
 Three-dimensional nanoscopy of whole cells and tissues with in situ point spread function retrieval
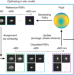
Three-dimensional nanoscopy of whole cells and tissues with in situ point spread function retrieval
In situ point spread function (PSF) retrieval (INSPR) enables precise single-molecule localization in 3D single-molecule localization microscopy of whole cells and tissues. It directly determines PSF from a single-molecule blinking dataset, removing errors associated with sample-induced aberrations.
- Fan Xu
- , Donghan Ma
- & Fang Huang
-
Article |
 Single-molecule displacement mapping unveils nanoscale heterogeneities in intracellular diffusivity
Single-molecule displacement mapping unveils nanoscale heterogeneities in intracellular diffusivity
Single-molecule displacement/diffusivity mapping (SMdM) enables nanoscale mapping of freely diffusing molecules in mammalian cells and reveals the structural basis of variations in local diffusivity in both the cytoplasm and nucleus.
- Limin Xiang
- , Kun Chen
- & Ke Xu
-
Research Highlight |
 CLEM takes a polar plunge
CLEM takes a polar plunge
Combining cryogenic super-resolution microscopy and focused ion beam scanning electron microscopy allows nanoscale views of protein architecture in the context of cellular ultrastructure in whole cells.
- Rita Strack
-
Perspective |
 Visualizing the genome in high resolution challenges our textbook understanding
Visualizing the genome in high resolution challenges our textbook understanding
This Perspective highlights how high-resolution imaging has informed our view and helped overturn the textbook understanding of 4D genome organization.
- Melike Lakadamyali
- & Maria Pia Cosma
-
Article |
 MINFLUX nanoscopy delivers 3D multicolor nanometer resolution in cells
MINFLUX nanoscopy delivers 3D multicolor nanometer resolution in cells
Advances in MINFLUX nanoscopy enable multicolor imaging over large fields of view, bringing true nanometer-scale fluorescence imaging to labeled structures in fixed and living cells.
- Klaus C. Gwosch
- , Jasmin K. Pape
- & Stefan W. Hell
-
Article |
 Nanoscale subcellular architecture revealed by multicolor three-dimensional salvaged fluorescence imaging
Nanoscale subcellular architecture revealed by multicolor three-dimensional salvaged fluorescence imaging
4Pi single-molecule switching microscopy combined with ‘salvaged fluorescence’ enables improved ratiometric imaging that bypasses chromatic aberrations and allows for multicolor whole-cell imaging with sub-10-nm localization precision.
- Yongdeng Zhang
- , Lena K. Schroeder
- & Joerg Bewersdorf
-
Brief Communication |
 Localization microscopy at doubled precision with patterned illumination
Localization microscopy at doubled precision with patterned illumination
SIMFLUX combines elements of MINFLUX with structured illumination to double localization precision and improve resolution in localization microscopy. The approach was demonstrated on DNA origami and on cellular microtubules.
- Jelmer Cnossen
- , Taylor Hinsdale
- & Sjoerd Stallinga
-
-
-
Brief Communication |
 An order of magnitude faster DNA-PAINT imaging by optimized sequence design and buffer conditions
An order of magnitude faster DNA-PAINT imaging by optimized sequence design and buffer conditions
DNA-PAINT is sped up by an order of magnitude by optimizing sequences and buffer conditions, enabling faster imaging with no compromise to image quality or resolution, improved single-molecule counting and enhanced cellular imaging.
- Florian Schueder
- , Johannes Stein
- & Ralf Jungmann
-
Research Highlight |
Lipid biology in super-resolution
BODIPY dimerization is exploited for super-resolution imaging and single-particle tracking of lipids in living cells.
- Rita Strack
-
-
Article |
 Nuclear pores as versatile reference standards for quantitative superresolution microscopy
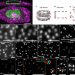
Nuclear pores as versatile reference standards for quantitative superresolution microscopy
Cell lines in which Nup96 is endogenously tagged with mEGFP, SNAP-tag, HaloTag or mMaple serve as versatile reference samples, enabling 3D resolution calibration, assessment of labeling efficiency and precise molecular counting.
- Jervis Vermal Thevathasan
- , Maurice Kahnwald
- & Jonas Ries
-
Brief Communication |
 Molecular resolution imaging by repetitive optical selective exposure
Molecular resolution imaging by repetitive optical selective exposure
Repetitive optical selective exposure (ROSE) is an interferometric single-molecule localization microscopy method offering twofold improvement in lateral resolution with the same photon budget compared with conventional approaches.
- Lusheng Gu
- , Yuanyuan Li
- & Wei Ji
-
Brief Communication |
 Oblique-plane single-molecule localization microscopy for tissues and small intact animals
Oblique-plane single-molecule localization microscopy for tissues and small intact animals
Single-molecule oblique-plane microscopy (obSTORM) enables deep volumetric super-resolution imaging in a light-sheet microscopy platform that is convenient for standard tissues and small intact animals.
- Jeongmin Kim
- , Michal Wojcik
- & Xiang Zhang
-
-
Brief Communication |
 Mechanistic investigation of mEos4b reveals a strategy to reduce track interruptions in sptPALM
Mechanistic investigation of mEos4b reveals a strategy to reduce track interruptions in sptPALM
The red form of the photoconvertible fluorescent protein mEos4b has a long-lived dark state with specific chromophore conformation. Weak 488-nm light depopulates this state, improving track lengths in single-particle tracking experiments.
- Elke De Zitter
- , Daniel Thédié
- & Dominique Bourgeois
-
Brief Communication |
 Epi-illumination SPIM for volumetric imaging with high spatial-temporal resolution
Epi-illumination SPIM for volumetric imaging with high spatial-temporal resolution
Epi-illumination SPIM enables fast, volumetric, high-resolution, subcellular imaging of any sample compatible with a standard inverted fluorescence microscope.
- Bin Yang
- , Xingye Chen
- & Bo Huang
-
Research Highlight |
 Bypassing bleaching with fluxional fluorophores
Bypassing bleaching with fluxional fluorophores
Combining photoactivation and spontaneous blinking enables long time-lapse localization microscopy.
- Rita Strack
-
Analysis |
 Super-resolution fight club: assessment of 2D and 3D single-molecule localization microscopy software
Super-resolution fight club: assessment of 2D and 3D single-molecule localization microscopy software
This study reports results from the second community-wide single-molecule localization microscopy software challenge, which tested over 30 software packages on realistic simulated data for multiple popular 3D image acquisition modes, as well as 2D localization microscopy.
- Daniel Sage
- , Thanh-An Pham
- & Seamus Holden
-
Research Highlight |
Bridging the nanoscale measurement gap
Single-molecule imaging enables accurate distance measurements from two to hundreds of nanometers.
- Rita Strack
-
-
-
Research Highlight |
 Quantum imaging in biological samples
Quantum imaging in biological samples
The combination of image scanning microscopy and quantum imaging improves resolution up to fourfold compared with the classical diffraction barrier.
- Christian Schnell
-
Brief Communication |
 A robust and versatile platform for image scanning microscopy enabling super-resolution FLIM
A robust and versatile platform for image scanning microscopy enabling super-resolution FLIM
A single-photon detector array enables robust and versatile image scanning microscopy (ISM) on any confocal microscope. This implementation makes super-resolution FLIM possible and eases a transition from confocal microscopy to ISM.
- Marco Castello
- , Giorgio Tortarolo
- & Giuseppe Vicidomini
-
-
Perspective |
 Expansion microscopy: principles and uses in biological research
Expansion microscopy: principles and uses in biological research
Expansion microscopy allows super-resolution images of diverse samples to be acquired on conventional microscopes, thus democratizing super-resolution imaging. This Perspective reviews available methods and provides practical guidance for users.
- Asmamaw T. Wassie
- , Yongxin Zhao
- & Edward S. Boyden
-
Brief Communication |
 Imaging cellular ultrastructures using expansion microscopy (U-ExM)
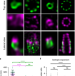
Imaging cellular ultrastructures using expansion microscopy (U-ExM)
U-ExM enables near-native expansion microscopy of samples in vitro and in cells. The combination of U-ExM with confocal microscopy and HyVolution revealed details of centriole chirality that were previously accessible only by electron microscopy.
- Davide Gambarotto
- , Fabian U. Zwettler
- & Paul Guichard
-
Article |
 Deep learning enables cross-modality super-resolution in fluorescence microscopy
Deep learning enables cross-modality super-resolution in fluorescence microscopy
Deep learning enables cross-modality super-resolution imaging, including confocal-to-STED and TIRF-to-TIRF-SIM image transformation. Imaging of a larger FOV and greater depth of field is possible with higher resolution and SNR at lower light doses.
- Hongda Wang
- , Yair Rivenson
- & Aydogan Ozcan
-
Technology Feature |
 Specialty probes give super-res imaging that special blink
Specialty probes give super-res imaging that special blink
Users of super-resolution imaging describe how they match probes to imaging modalities
- Vivien Marx
-
Correspondence |
 Impact of optical aberrations on axial position determination by photometry
Impact of optical aberrations on axial position determination by photometry
- Rasmus Ø. Thorsen
- , Christiaan N. Hulleman
- & Bernd Rieger
-
Perspective |
 Faster, sharper, and deeper: structured illumination microscopy for biological imaging
Faster, sharper, and deeper: structured illumination microscopy for biological imaging
A Perspective on super-resolution structured illumination microscopy reviews advances in these methods and focuses on matching user needs in terms of imaging speed, sample depth, and desired resolution with the appropriate instrumentation.
- Yicong Wu
- & Hari Shroff
-
-
Brief Communication |
 Analyzing complex single-molecule emission patterns with deep learning
Analyzing complex single-molecule emission patterns with deep learning
The deep neural network smNet extracts multiplexed parameters such as 3D position, orientation and wavefront distortion from emission patterns of single molecules.
- Peiyi Zhang
- , Sheng Liu
- & Fang Huang
-
-
Correspondence |
 Assessing photodamage in live-cell STED microscopy
Assessing photodamage in live-cell STED microscopy
- Nicole Kilian
- , Alexander Goryaynov
- & Joerg Bewersdorf
-
News & Views |
 Single-particle analysis for fluorescence nanoscopy
Single-particle analysis for fluorescence nanoscopy
Building on methods from electron microscopy, researchers are applying nanoscale fluorescence imaging to address questions in structural biology.
- Mark Bates
-
News & Views |
 Improving probes for super-resolution
Improving probes for super-resolution
Chemically modified DNA aptamers enable quantitative super-resolution imaging.
- Regan P. Moore
- & Wesley R. Legant
-
Brief Communication |
 Modified aptamers enable quantitative sub-10-nm cellular DNA-PAINT imaging
Modified aptamers enable quantitative sub-10-nm cellular DNA-PAINT imaging
Slow off-rate modified aptamer (SOMAmer) reagents are small and versatile probes for DNA-PAINT super-resolution microscopy that enable multiplexed, quantitative, and high-resolution imaging in fixed and live cells.
- Sebastian Strauss
- , Philipp C. Nickels
- & Ralf Jungmann
-
