Featured
-
-
Article
| Open Access Evaluating the impact of curfews and other measures on SARS-CoV-2 transmission in French Guiana
Evaluating the impact of curfews and other measures on SARS-CoV-2 transmission in French Guiana
Identifying effective combinations of control measures in different populations is important for SARS-CoV-2 control. Here, the authors show that in French Guiana, which has a relatively young population, curfews and localised lockdowns appeared to contribute to reducing transmission.
- Alessio Andronico
- , Cécile Tran Kiem
- & Simon Cauchemez
-
Article
| Open Access A deep learning method for HLA imputation and trans-ethnic MHC fine-mapping of type 1 diabetes
A deep learning method for HLA imputation and trans-ethnic MHC fine-mapping of type 1 diabetes
Human leukocyte antigen (HLA) genes contribute to risk of many complex traits, yet understanding inter-ethnic heterogeneity is computationally challenging. Here, the authors develop DEEP*HLA for imputation of HLA genotypes and show its ability to disentangle HLA variant risk effects in diverse populations.
- Tatsuhiko Naito
- , Ken Suzuki
- & Yukinori Okada
-
Article
| Open Access Real-time tracking and prediction of COVID-19 infection using digital proxies of population mobility and mixing
Real-time tracking and prediction of COVID-19 infection using digital proxies of population mobility and mixing
Digital proxies of human mobility can be used to monitor social distancing, and therefore have potential to infer COVID-19 dynamics. Here, the authors integrate travel card data from Hong Kong into a transmission model and show that it can be used to track transmissibility in near real-time.
- Kathy Leung
- , Joseph T. Wu
- & Gabriel M. Leung
-
Article
| Open Access Prioritizing non-coding regions based on human genomic constraint and sequence context with deep learning
Prioritizing non-coding regions based on human genomic constraint and sequence context with deep learning
Intolerance to variation is a strong indicator of disease relevance for coding regions of the human genome. Here, the authors present JARVIS, a deep learning method integrating intolerance to variation in non-coding regions and sequence-specific annotations to infer non-coding variant pathogenicity.
- Dimitrios Vitsios
- , Ryan S. Dhindsa
- & Slavé Petrovski
-
Article
| Open Access Accurate imputation of human leukocyte antigens with CookHLA
Accurate imputation of human leukocyte antigens with CookHLA
Human leukocyte antigen (HLA) genes influence many immune phenotypes, however methods to impute HLA type have been limited in accuracy. Here, the authors present an HLA imputation method, CookHLA, which uses locally embedded prediction markers to adaptively impute HLA genes across a range of scenarios.
- Seungho Cook
- , Wanson Choi
- & Buhm Han
-
Article
| Open Access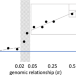 Phenotypic covariance across the entire spectrum of relatedness for 86 billion pairs of individuals
Phenotypic covariance across the entire spectrum of relatedness for 86 billion pairs of individuals
Assigning inter-individual similarities to genetic and non-genetic factors is central to quantitative genetics. Here, the authors look at phenotypic covariance among pairs of individuals for 32 traits across the UK Biobank, from nominally unrelated pairs through to monozygotic twins.
- Kathryn E. Kemper
- , Loic Yengo
- & Peter M. Visscher
-
Article
| Open Access A practical solution to pseudoreplication bias in single-cell studies
A practical solution to pseudoreplication bias in single-cell studies
Single cell genomics uses cells from the same individual, or pseudoreplicates, that can introduce biases and inflate type I error rates. Here the authors apply generalized linear mixed models with a random effect for individual, to properly account for both zero inflation and the correlation structure among cells within an individual.
- Kip D. Zimmerman
- , Mark A. Espeland
- & Carl D. Langefeld
-
Article
| Open Access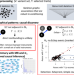 Graphical analysis for phenome-wide causal discovery in genotyped population-scale biobanks
Graphical analysis for phenome-wide causal discovery in genotyped population-scale biobanks
Mendelian randomization is a popular method to detect causal relationships between traits, but can be confounded by instances of horizontal pleiotropy. Here, the authors present a Mendelian randomization workflow which includes causal discovery analysis and filtering of genetic instruments based on their conditional independencies.
- David Amar
- , Nasa Sinnott-Armstrong
- & Manuel A. Rivas
-
Article
| Open Access Lossless integration of multiple electronic health records for identifying pleiotropy using summary statistics
Lossless integration of multiple electronic health records for identifying pleiotropy using summary statistics
Thus far, pleiotropy analysis using individual-level Electronic Health Records data has been limited to data from one site. Here, the authors introduce Sum-Share, a method designed to efficiently and losslessly integrate EHR and genetic data from multiple sites to perform pleiotropy analysis.
- Ruowang Li
- , Rui Duan
- & Jason H. Moore
-
Article
| Open Access Improving the informativeness of Mendelian disease-derived pathogenicity scores for common disease
Improving the informativeness of Mendelian disease-derived pathogenicity scores for common disease
Pathogenicity scores are instrumental in prioritizing variants for Mendelian disease, yet their application to common disease is largely unexplored. Here, the authors assess the utility of pathogenicity scores for 41 complex traits and develop a framework to improve their informativeness for common disease.
- Samuel S. Kim
- , Kushal K. Dey
- & Alkes L. Price
-
Article
| Open Access muscat detects subpopulation-specific state transitions from multi-sample multi-condition single-cell transcriptomics data
muscat detects subpopulation-specific state transitions from multi-sample multi-condition single-cell transcriptomics data
Single-cell transcriptomics enhanced our ability to profile heterogeneous cell populations. It is not known which statistical frameworks are performant to detect subpopulation-level responses. Here, the authors developed a simulation framework to evaluate various methods across a range of scenarios.
- Helena L. Crowell
- , Charlotte Soneson
- & Mark D. Robinson
-
Article
| Open Access Ensemble dimensionality reduction and feature gene extraction for single-cell RNA-seq data
Ensemble dimensionality reduction and feature gene extraction for single-cell RNA-seq data
Dimensionality reduction is used to make the analysis of single-cell RNA sequencing data more efficient. Here the authors propose a method, EDGE, which simultaneously carries out dimensionality reduction and feature gene extraction.
- Xiaoxiao Sun
- , Yiwen Liu
- & Lingling An
-
Article
| Open Access A computational method for detection of ligand-binding proteins from dose range thermal proteome profiles
A computational method for detection of ligand-binding proteins from dose range thermal proteome profiles
2D-thermal proteome profiling (2D-TPP) is a powerful assay for probing interactions of proteins with small molecules in their native context. Here the authors provide a statistical method for false discovery rate controlled analysis for 2D-TPP applications.
- Nils Kurzawa
- , Isabelle Becher
- & Mikhail M. Savitski
-
Article
| Open Access Collider bias undermines our understanding of COVID-19 disease risk and severity
Collider bias undermines our understanding of COVID-19 disease risk and severity
Many published studies of the current SARS-CoV-2 pandemic have analysed data from non-representative samples from populations. Here, using UK BioBank samples, Gibran Hemani and colleagues discuss the potential for such studies to suffer from collider bias, and provide suggestions for optimising study design to account for this.
- Gareth J. Griffith
- , Tim T. Morris
- & Gibran Hemani
-
Article
| Open Access Heritability of the HIV-1 reservoir size and decay under long-term suppressive ART
Heritability of the HIV-1 reservoir size and decay under long-term suppressive ART
The HIV reservoir is a major hurdle for a cure of HIV, but the factors determining its size and dynamics remain unclear. Here the authors show in a large cohort of 610 HIV-1 infected individuals, who are on suppressive ART for a median of 5.4 years, that viral genetic factors contribute substantially to the HIV-1 reservoir size.
- Chenjie Wan
- , Nadine Bachmann
- & Sabine Yerly
-
Article
| Open Access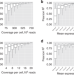 Optimized design of single-cell RNA sequencing experiments for cell-type-specific eQTL analysis
Optimized design of single-cell RNA sequencing experiments for cell-type-specific eQTL analysis
Single cell RNA-sequencing can be a powerful approach to characterizing cell composition in a population of cells but is thought to be too expensive for population-scale analyses. Here, the authors show how lower coverage of more samples can increase the power to detect cell-type-specific eQTL.
- Igor Mandric
- , Tommer Schwarz
- & Eran Halperin
-
Article
| Open Access Pattern recognition based on machine learning identifies oil adulteration and edible oil mixtures
Pattern recognition based on machine learning identifies oil adulteration and edible oil mixtures
Fraudulent adulteration of edible oils is based on the fact that their characteristic fatty acid profile can be mimicked with mixtures of other oil types. Here, the authors use a deep learning method to uncover fatty acid patterns discriminative for ten different plant oil types and to discern composition of mixtures.
- Kevin Lim
- , Kun Pan
- & Rong Hui Xiao
-
Article
| Open Access Mendelian randomization while jointly modeling cis genetics identifies causal relationships between gene expression and lipids
Mendelian randomization while jointly modeling cis genetics identifies causal relationships between gene expression and lipids
Mendelian randomization is a useful tool to infer causal relationships between traits, but can be confounded by the presence of pleiotropy. Here, the authors have developed MR-link, a Mendelian randomization method which accounts for unobserved pleiotropy and linkage disequilibrium between instrumental variables.
- Adriaan van der Graaf
- , Annique Claringbould
- & Serena Sanna
-
Article
| Open Access A cell-type deconvolution meta-analysis of whole blood EWAS reveals lineage-specific smoking-associated DNA methylation changes
A cell-type deconvolution meta-analysis of whole blood EWAS reveals lineage-specific smoking-associated DNA methylation changes
Smoking-associated DNA methylation changes in whole blood have been reported by many EWAS. Here, the authors use a cell-type deconvolution algorithm to identify cell-type specific DNA methylation signals in seven EWAS, identifying lineage-specific smoking-associated DNA methylation changes.
- Chenglong You
- , Sijie Wu
- & Andrew E. Teschendorff
-
Article
| Open Access CORE GREML for estimating covariance between random effects in linear mixed models for complex trait analyses
CORE GREML for estimating covariance between random effects in linear mixed models for complex trait analyses
Linear mixed models have bias due to the assumed independence between random effects. Here, the authors describe a genome-based restricted maximum likelihood, CORE GREML, which estimates covariance between random effects. Application to UK Biobank data highlights this as an important parameter for multi-omics analyses of phenotypic variance.
- Xuan Zhou
- , Hae Kyung Im
- & S. Hong Lee
-
Article
| Open Access Efficient variance components analysis across millions of genomes
Efficient variance components analysis across millions of genomes
Variance components analysis may be used for a variety of applications including heritability estimation and association mapping. Here, the authors present a computationally efficient method, scalable to extremely large GWAS datasets, and use it for heritabilty analysis of 22 traits from UK Biobank
- Ali Pazokitoroudi
- , Yue Wu
- & Sriram Sankararaman
-
Article
| Open Access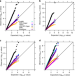 Testing and controlling for horizontal pleiotropy with probabilistic Mendelian randomization in transcriptome-wide association studies
Testing and controlling for horizontal pleiotropy with probabilistic Mendelian randomization in transcriptome-wide association studies
Transcriptome-wide association studies integrate GWAS and transcriptome data to examine the molecular mechanisms underlying disease etiology. Here the authors present PMR-Egger, a powerful TWAS method based on probabilistic Mendelian Randomization.
- Zhongshang Yuan
- , Huanhuan Zhu
- & Xiang Zhou
-
Article
| Open Access Theoretical and empirical quantification of the accuracy of polygenic scores in ancestry divergent populations
Theoretical and empirical quantification of the accuracy of polygenic scores in ancestry divergent populations
Polygenic scores (PGS) are often based on GWAS data from individuals of European ancestry, thus limiting their use in populations of non-European ancestry. Here, the authors predict the relative accuracy of PGS across ancestries and suggest that causal variants are mostly shared across continents.
- Ying Wang
- , Jing Guo
- & Loic Yengo
-
Article
| Open Access Using sigLASSO to optimize cancer mutation signatures jointly with sampling likelihood
Using sigLASSO to optimize cancer mutation signatures jointly with sampling likelihood
The next generation sequencing has provided the opportunity to look for signatures of carcinogenesis on a genome wide scale. Here, the authors develop the algorithm, sigLASSO, that provides confidence in assigning mutational signatures when the mutation count is low and the samples used are variable.
- Shantao Li
- , Forrest W. Crawford
- & Mark B. Gerstein
-
Article
| Open Access A universal and independent synthetic DNA ladder for the quantitative measurement of genomic features
A universal and independent synthetic DNA ladder for the quantitative measurement of genomic features
Standard units of measurement are required for a quantitative description of the genome. Here, the authors present a universal synthetic DNA ladder that can measure genetic abundance in next-generation sequencing libraries.
- Andre L. M. Reis
- , Ira W. Deveson
- & Tim R. Mercer
-
Article
| Open Access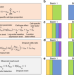 Flexible experimental designs for valid single-cell RNA-sequencing experiments allowing batch effects correction
Flexible experimental designs for valid single-cell RNA-sequencing experiments allowing batch effects correction
It is not clear which designs, other than completely randomized ones, are valid for scRNA-seq experiments so that batch effects can be adjusted. Here the authors show that under flexible reference panel and chain-type designs, biological variability can also be separated from batch effects, at least by BUSseq.
- Fangda Song
- , Ga Ming Angus Chan
- & Yingying Wei
-
Article
| Open Access Single-cell lineage tracing by integrating CRISPR-Cas9 mutations with transcriptomic data
Single-cell lineage tracing by integrating CRISPR-Cas9 mutations with transcriptomic data
Lineage tracing studies combining CRISPR-Cas9 editing and scRNA-seq face several challenges and cannot integrate lineages from multiple individuals. Here the authors show that integration of mutation and expression leads to accurate lineage tree inference and enables the learning of a species-invariant lineage tree.
- Hamim Zafar
- , Chieh Lin
- & Ziv Bar-Joseph
-
Article
| Open Access Bayesian reassessment of the epigenetic architecture of complex traits
Bayesian reassessment of the epigenetic architecture of complex traits
Linking epigenetic marks to clinical outcomes promises insight into the underlying processes. Here, the authors introduce a statistical approach to estimate associations between a phenotype and all epigenetic probes jointly, and to estimate the proportion of variation captured by epigenetic effects.
- Daniel Trejo Banos
- , Daniel L. McCartney
- & Matthew R. Robinson
-
Article
| Open Access Reconstructing Mayaro virus circulation in French Guiana shows frequent spillovers
Reconstructing Mayaro virus circulation in French Guiana shows frequent spillovers
Mayaro virus (MAYV) is an emerging arbovirus, but cross-reactivity with other alphaviruses makes analysis of its epidemiology difficult. Here, the authors develop an analytical framework to assess MAYV epidemiology and find evidence for an important sylvatic cycle and seroprevalences of up to 18% in some areas of French Guiana.
- Nathanaël Hozé
- , Henrik Salje
- & Simon Cauchemez
-
Article
| Open Access Multi-trait analysis of rare-variant association summary statistics using MTAR
Multi-trait analysis of rare-variant association summary statistics using MTAR
Methods to integrate association evidence across multiple traits often focus on individual common variants GWAS. Here the authors present multi-trait analysis of rare-variant associations (MTAR), a framework for joint analysis of association summary statistics between multiple rare variants and different traits.
- Lan Luo
- , Judong Shen
- & Zheng-Zheng Tang
-
Article
| Open Access Model-based analysis of sample index hopping reveals its widespread artifacts in multiplexed single-cell RNA-sequencing
Model-based analysis of sample index hopping reveals its widespread artifacts in multiplexed single-cell RNA-sequencing
Sample index hopping results in various artefacts in multiplexed scRNA-seq experiments. Here, the authors introduce a statistical model to estimate sample index hopping rate in droplet-based scRNA-seq data and show that artifacts can be corrected by purging phantom molecules from the data.
- Rick Farouni
- , Haig Djambazian
- & Hamed S. Najafabadi
-
Article
| Open Access Deep learning enables accurate clustering with batch effect removal in single-cell RNA-seq analysis
Deep learning enables accurate clustering with batch effect removal in single-cell RNA-seq analysis
Increasingly large scRNA-seq datasets demand better and more scalable analysis tools. Here, the authors introduce a scalable unsupervised deep embedding algorithm that clusters scRNA-seq data by iteratively optimizing a clustering objective function and enables removal of batch effects.
- Xiangjie Li
- , Kui Wang
- & Mingyao Li
-
Article
| Open Access Integrative differential expression and gene set enrichment analysis using summary statistics for scRNA-seq studies
Integrative differential expression and gene set enrichment analysis using summary statistics for scRNA-seq studies
Differential expression (DE) and gene set enrichment (GSE) analysis tend to be carried out separately. Here, the authors present iDEA (integrative Differential expression and gene set Enrichment Analysis) for the analysis of scRNAseq data which uses a Baysian approach to jointly model DE and GSE for improved power in both tasks.
- Ying Ma
- , Shiquan Sun
- & Xiang Zhou
-
Article
| Open Access Quantification of the overall contribution of gene-environment interaction for obesity-related traits
Quantification of the overall contribution of gene-environment interaction for obesity-related traits
Most gene-by-environment interaction methods rely on the availability of the interacting environment. Here, the authors propose a robust maximum likelihood method for estimating the overall statistical interaction between a genetic risk score for a continuous outcome and all environmental variables.
- Jonathan Sulc
- , Ninon Mounier
- & Zoltán Kutalik
-
Article
| Open Access Trajectory-based differential expression analysis for single-cell sequencing data
Trajectory-based differential expression analysis for single-cell sequencing data
Downstream of trajectory inference for cell lineages based on scRNA-seq data, differential expression analysis yields insight into biological processes. Here, Van den Berge et al. develop tradeSeq, a framework for the inference of within and between-lineage differential expression, based on negative binomial generalized additive models.
- Koen Van den Berge
- , Hector Roux de Bézieux
- & Lieven Clement
-
Article
| Open Access Multi-resolution localization of causal variants across the genome
Multi-resolution localization of causal variants across the genome
GWAS analysis currently relies mostly on linear mixed models, which do not account for linkage disequilibrium (LD) between tested variants. Here, Sesia et al. propose KnockoffZoom, a non-parametric statistical method for the simultaneous discovery and fine-mapping of causal variants, assuming only that LD is described by hidden Markov models (HMMs).
- Matteo Sesia
- , Eugene Katsevich
- & Chiara Sabatti
-
Article
| Open Access Exploiting horizontal pleiotropy to search for causal pathways within a Mendelian randomization framework
Exploiting horizontal pleiotropy to search for causal pathways within a Mendelian randomization framework
In Mendelian randomization (MR) studies, one typically selects SNPs as instrumental variables that do not directly affect the outcome to avoid violation of MR assumptions. Here, Cho et al. present a framework, MR-TRYX, that leverages knowledge of such outliers of horizontal pleiotropy to identify putative causal relationships between exposure and outcome.
- Yoonsu Cho
- , Philip C. Haycock
- & Gibran Hemani
-
Article
| Open Access Discovering the genes mediating the interactions between chronic respiratory diseases in the human interactome
Discovering the genes mediating the interactions between chronic respiratory diseases in the human interactome
Complex diseases often share genetic determinants and symptoms, but the mechanistic basis of disease interactions remains elusive. Here, the authors propose a network topological measure to identify proteins linking complex diseases in the interactome, and identify mediators between COPD and asthma.
- Enrico Maiorino
- , Seung Han Baek
- & Amitabh Sharma
-
Article
| Open Access Determining sequencing depth in a single-cell RNA-seq experiment
Determining sequencing depth in a single-cell RNA-seq experiment
For single-cell RNA-seq experiments the sequencing budget is limited, and how it should be optimally allocated to maximize information is not clear. Here the authors develop a mathematical framework to show that, for estimating many gene properties, the optimal allocation is to sequence at the depth of one read per cell per gene.
- Martin Jinye Zhang
- , Vasilis Ntranos
- & David Tse
-
Article
| Open Access Characterizing chromatin landscape from aggregate and single-cell genomic assays using flexible duration modeling
Characterizing chromatin landscape from aggregate and single-cell genomic assays using flexible duration modeling
Most currently available statistical tools for the analysis of ATAC-seq data were repurposed from tools developed for other functional genomics data (e.g. ChIP-seq). Here, Gabitto et al develop ChromA, a Bayesian statistical approach for the analysis of both bulk and single-cell ATAC-seq data.
- Mariano I. Gabitto
- , Anders Rasmussen
- & Richard Bonneau
-
Article
| Open Access Mechanistic insights into transcription factor cooperativity and its impact on protein-phenotype interactions
Mechanistic insights into transcription factor cooperativity and its impact on protein-phenotype interactions
Although transcription factor (TF) cooperativity is widespread, a global mechanistic understanding of the role of TF cooperativity is still lacking. Here the authors introduce a statistical learning framework that provides structural insight into TF cooperativity and its functional consequences based on next generation sequencing data and provide mechanistic insights into TF cooperativity and its impact on protein-phenotype interactions.
- Ignacio L. Ibarra
- , Nele M. Hollmann
- & Judith B. Zaugg
-
Article
| Open Access Selecting likely causal risk factors from high-throughput experiments using multivariable Mendelian randomization
Selecting likely causal risk factors from high-throughput experiments using multivariable Mendelian randomization
Multivariable Mendelian randomization (MR) extends the standard MR framework to consider multiple risk factors in a single model. Here, Zuber et al. propose MR-BMA, a Bayesian variable selection approach to identify the likely causal determinants of a disease from many candidate risk factors as for example high-throughput data sets.
- Verena Zuber
- , Johanna Maria Colijn
- & Stephen Burgess
-
Article
| Open Access Using somatic variant richness to mine signals from rare variants in the cancer genome
Using somatic variant richness to mine signals from rare variants in the cancer genome
Sequencing cancer genomes reveals low frequency novel somatic variants without known function. Here, the authors leverage statistical methodology from the fields of computational linguistics and ecology to highlight the potentially important signals harboured by these novel variants that are often dismissed.
- Saptarshi Chakraborty
- , Arshi Arora
- & Ronglai Shen
-
Article
| Open Access Estimating heritability and genetic correlations from large health datasets in the absence of genetic data
Estimating heritability and genetic correlations from large health datasets in the absence of genetic data
Disease heritability and genetic correlations between traits depend on genetics, the environment and their interaction. Here, Jia et al. compute disease prevalence curves and disease embeddings from electronic health records and impute heritability for hundreds of diseases and genetic correlations for thousands of disease pairs.
- Gengjie Jia
- , Yu Li
- & Andrey Rzhetsky
-
Article
| Open Access A Bayesian mixture model for the analysis of allelic expression in single cells
A Bayesian mixture model for the analysis of allelic expression in single cells
Allele-specific expression at single-cell resolution can reveal stochastic and dynamic features of gene expression in greater detail. The authors propose scBASE, a soft zero-and-one inflated model that improves estimation of cellular allelic proportions by pooling information across cells.
- Kwangbom Choi
- , Narayanan Raghupathy
- & Gary A. Churchill
-
Article
| Open Access Improved polygenic prediction by Bayesian multiple regression on summary statistics
Improved polygenic prediction by Bayesian multiple regression on summary statistics
Various approaches are being used for polygenic prediction including Bayesian multiple regression methods that require access to individual-level genotype data. Here, the authors extend BayesR to utilise GWAS summary statistics (SBayesR) and show that it outperforms other summary statistic-based methods.
- Luke R. Lloyd-Jones
- , Jian Zeng
- & Peter M. Visscher
-
Article
| Open Access Species abundance information improves sequence taxonomy classification accuracy
Species abundance information improves sequence taxonomy classification accuracy
Taxonomy classification of amplicon sequences is an important step in investigating microbial communities in microbiome analysis. Here, the authors show incorporating environment-specific taxonomic abundance information can lead to improved species-level classification accuracy across common sample types.
- Benjamin D. Kaehler
- , Nicholas A. Bokulich
- & Gavin A. Huttley
-
Article
| Open Access Thermodynamic control of −1 programmed ribosomal frameshifting
Thermodynamic control of −1 programmed ribosomal frameshifting
Programmed ribosomal frameshifting (PRF) is an alternative translation strategy that causes controlled slippage of the ribosome along the mRNA, changing the sequence of the synthesized protein. Here the authors provide a thermodynamic framework that explains how mRNA sequence determines the efficiency of frameshifting.
- Lars V. Bock
- , Neva Caliskan
- & Helmut Grubmüller
-
Article
| Open Access Identification of significant chromatin contacts from HiChIP data by FitHiChIP
Identification of significant chromatin contacts from HiChIP data by FitHiChIP
HiChIP/PLAC-seq assay is popular for profiling 3D genome interactions among regulatory elements at kilobase resolution. Here the authors describe FitHiChIP an empirical null-based, flexible computational method for statistical significance estimation and loop calling from HiChIP data.
- Sourya Bhattacharyya
- , Vivek Chandra
- & Ferhat Ay

 Model-based deep embedding for constrained clustering analysis of single cell RNA-seq data
Model-based deep embedding for constrained clustering analysis of single cell RNA-seq data