Article
|
Open Access
Featured
-
-
Article
| Open Access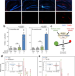 Death Induced by Survival gene Elimination (DISE) correlates with neurotoxicity in Alzheimer’s disease and aging
Death Induced by Survival gene Elimination (DISE) correlates with neurotoxicity in Alzheimer’s disease and aging
Events that cause neurons to die in Alzheimer’s disease (AD) are poorly understood. Here, the authors provide evidence for a role of RNA interference in AD. Short RNAs causing neurotoxicity and DNA damage are seen in AD and aged brains, and are counteracted by nontoxic RNAs.
- Bidur Paudel
- , Si-Yeon Jeong
- & Marcus E. Peter
-
Article
| Open Access Structural basis of antiphage immunity generated by a prokaryotic Argonaute-associated SPARSA system
Structural basis of antiphage immunity generated by a prokaryotic Argonaute-associated SPARSA system
Short prokaryotic Argonaute and Sir2 proteins function as an antivirus system. Here the authors describe structures of SPARSA (a heterodimer of Sir2-APAZ and prokaryotic Argonaute) with and without template DNA and guide RNA, providing structural basis of its assembly and activation by the recognition of the invading virus.
- Xiangkai Zhen
- , Xiaolong Xu
- & Songying Ouyang
-
Article
| Open Access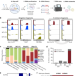 In vivo RNA interactome profiling reveals 3’UTR-processed small RNA targeting a central regulatory hub
In vivo RNA interactome profiling reveals 3’UTR-processed small RNA targeting a central regulatory hub
Here the authors report a new approach to profile RNA-RNA interactions in live bacterial cells. The charted RNA interaction networks unveil a key mRNA regulatory hub targeted by twelve small RNAs, including a novel RNA involved in fatty acid metabolism.
- Fang Liu
- , Ziying Chen
- & Yanjie Chao
-
Article
| Open Access C. elegans germ granules sculpt both germline and somatic RNAome
C. elegans germ granules sculpt both germline and somatic RNAome
Germ granules are membraneless organelles that act as organizing centers for small RNA biogenesis during germline development. Here they show the LOTUS domain protein eggd-1 organizes germ granules, and reveal a germ line-to-soma communication pathway activated upon perturbation of germ granules.
- Ian F. Price
- , Jillian A. Wagner
- & Wen Tang
-
Article
| Open Access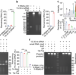 Discovery of the major 15–30 nt mammalian small RNAs, their biogenesis and function
Discovery of the major 15–30 nt mammalian small RNAs, their biogenesis and function
The authors uncover the major 15-30 nt mouse and human small RNAs (sRNAs), which mainly end with 2’,3’-cyclic phosphate, and show that many of these sRNAs can be generated by Angiogenin or RNase 4 and function in Ago2 complex as miRNAs.
- Hejin Lai
- , Ning Feng
- & Qiwei Zhai
-
Article
| Open Access A linear and circular dual-conformation noncoding RNA involved in oxidative stress tolerance in Bacillus altitudinis
A linear and circular dual-conformation noncoding RNA involved in oxidative stress tolerance in Bacillus altitudinis
The presence and/or biological functionality of circular RNAs in bacteria are unclear. Here, the authors identify a dual-conformation (linear and circular) noncoding RNA that promotes tolerance to oxidative stress in Bacillus altitutidinis, and provide evidence for the existence of other circular RNAs in diverse bacterial species.
- Ting-Ting He
- , Yun-Fan Xu
- & Hai-Yan Wang
-
Article
| Open Access Mechanism of U6 snRNA oligouridylation by human TUT1
Mechanism of U6 snRNA oligouridylation by human TUT1
U6 snRNA plays a catalytic role in pre-mRNA splicing. Here the authors report the structure of human terminal uridyltransferase, TUT1, complexed with U6 snRNA revealing the mechanism of specific uridylation of U6 snRNA by TUT1.
- Seisuke Yamashita
- & Kozo Tomita
-
Article
| Open Access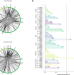 An RNA sponge controls quorum sensing dynamics and biofilm formation in Vibrio cholerae
An RNA sponge controls quorum sensing dynamics and biofilm formation in Vibrio cholerae
Small regulatory RNAs (sRNAs) often act in concert with the RNA-chaperone Hfq to regulate the expression of multiple target transcripts in bacteria. Here, the authors identify Hfq-interacting sRNAs and their targets in the pathogen Vibrio cholerae, including an RNA sponge that binds and inactivates four sRNAs that modulate the quorum sensing pathway.
- Michaela Huber
- , Anne Lippegaus
- & Kai Papenfort
-
Article
| Open Access Altered tRNA processing is linked to a distinct and unusual La protein in Tetrahymena thermophila
Altered tRNA processing is linked to a distinct and unusual La protein in Tetrahymena thermophila
La proteins are conserved factors critical for the maturation of RNA polymerase III transcripts. In the ciliate T. thermophila and related alveolates, La proteins have a novel domain arrangement and are linked to a distinct pre-tRNA processing pathway.
- Kyra Kerkhofs
- , Jyoti Garg
- & Mark A. Bayfield
-
Article
| Open Access A cancer-associated RNA polymerase III identity drives robust transcription and expression of snaR-A noncoding RNA
A cancer-associated RNA polymerase III identity drives robust transcription and expression of snaR-A noncoding RNA
RNA polymerase III changes its subunit composition during mammalian development. Here the authors report that loss of subunit POLR3G, which re-emerges in cancer, promotes expression of small NF90-associated RNA (snaR-A), a noncoding RNA implicated in cell proliferation and metastasis.
- Kevin Van Bortle
- , David P. Marciano
- & Michael P. Snyder
-
Article
| Open Access Diversity of bacterial small RNAs drives competitive strategies for a mutual chaperone
Diversity of bacterial small RNAs drives competitive strategies for a mutual chaperone
Bacterial sRNAs compete for limited copies of an RNA chaperone. Here, real-time single-molecule imaging shows that pairwise interactions of sRNAs with Hfq determine if the sRNAs are actively exchanged or stably coexist on the shared protein.
- Jorjethe Roca
- , Andrew Santiago-Frangos
- & Sarah A. Woodson
-
Article
| Open Access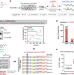 TRMT6/61A-dependent base methylation of tRNA-derived fragments regulates gene-silencing activity and the unfolded protein response in bladder cancer
TRMT6/61A-dependent base methylation of tRNA-derived fragments regulates gene-silencing activity and the unfolded protein response in bladder cancer
RNA modifications are important regulators of RNA biology. Here we report N1-methyladenosine (m1A) enrichment on 22-nucleotide tRNA fragments and its effect on gene-silencing. Higher level of m1A in bladder cancer is accompanied by gene dysregulation in unfolded protein response.
- Zhangli Su
- , Ida Monshaugen
- & Anindya Dutta
-
Article
| Open Access Secondary structure RNA elements control the cleavage activity of DICER
Secondary structure RNA elements control the cleavage activity of DICER
MicroRNA precursors are cleaved by DICER to generate mature microRNAs in the cytoplasm. Here the authors employ high-throughput analysis of DICER cleavage activity and identify RNA secondary elements in precursor miRNAs and shRNAs, including a single nucleotide bulge, which govern its cleavage efficiency and accuracy.
- Trung Duc Nguyen
- , Tam Anh Trinh
- & Tuan Anh Nguyen
-
Article
| Open Access The core and accessory Hfq interactomes across Pseudomonas aeruginosa lineages
The core and accessory Hfq interactomes across Pseudomonas aeruginosa lineages
Protein Hfq regulates bacterial mRNA translation and stability by interacting with mRNAs and small noncoding RNAs. Here, the authors identify Hfq-interacting RNAs in three strains representing major phylogenetic lineages of Pseudomonas aeruginosa, highlighting intra-species diversity in post-transcriptional regulatory networks.
- Julian Trouillon
- , Kook Han
- & Stephen Lory
-
Article
| Open Access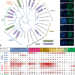 Small RNA pathways in the nematode Ascaris in the absence of piRNAs
Small RNA pathways in the nematode Ascaris in the absence of piRNAs
The parasitic nematode Ascaris lacks piRNAs. Here the authors compare Argonaute proteins and small RNAs from C. elegans and Ascaris, expanding our understanding of the conservation, divergence, and flexibility of Argonautes and small RNA pathways in nematodes.
- Maxim V. Zagoskin
- , Jianbin Wang
- & Richard E. Davis
-
Article
| Open Access An anionic ligand snap-locks a long-range interaction in a magnesium-folded riboswitch
An anionic ligand snap-locks a long-range interaction in a magnesium-folded riboswitch
A single molecule fluorescence study revealed three dynamically interconverting conformations of the fluoride riboswitch from Bacillus cereus, where an anionic ligand snap-locks a docked conformation through a long-range interaction necessary for downstream gene regulation.
- Rajeev Yadav
- , Julia R. Widom
- & Nils G. Walter
-
Article
| Open Access Angiogenin mediates paternal inflammation-induced metabolic disorders in offspring through sperm tsRNAs
Angiogenin mediates paternal inflammation-induced metabolic disorders in offspring through sperm tsRNAs
Paternal environmental inputs can influence various phenotypes in offspring, with important implications for basic biology and public health. Here the authors show that Ang-mediated biogenesis of 5′-tsRNAs in sperm contributes to paternal inflammation-induced metabolic disorders in offspring.
- Yanwen Zhang
- , Li Ren
- & Bin He
-
Article
| Open Access Association of snR190 snoRNA chaperone with early pre-60S particles is regulated by the RNA helicase Dbp7 in yeast
Association of snR190 snoRNA chaperone with early pre-60S particles is regulated by the RNA helicase Dbp7 in yeast
The molecular events underlying the assembly and maturation of the early pre-60S particles during eukaryotic ribosome synthesis are not well understood. Here, the authors combine yeast genetics and biochemical experiments to characterise the functions of two important players of eukaryotic ribosome biogenesis, the box C/D snoRNP snR190 and the helicase Dbp7, which both interact. They show that the snR190 snoRNA acts as a RNA chaperone that assists the structuring of the 25S rRNA during the maturation of early pre-60S particles and that Dbp7 is important for facilitating remodeling events in the peptidyl transferase center region of the 25S rRNAs during the maturation of early pre-60S particles.
- Mariam Jaafar
- , Julia Contreras
- & Anthony K. Henras
-
Article
| Open Access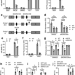 Loss of SNORA73 reprograms cellular metabolism and protects against steatohepatitis
Loss of SNORA73 reprograms cellular metabolism and protects against steatohepatitis
Lipid induced stress contributes to metabolic diseases. Here the authors identify small nucleolar RNA 73 (SNORA73) in a screen for genes that protect against lipotoxicity and show that deficiency of SNORA73 reprograms oxidative metabolism and protects against steatohepatitis in mice.
- Arthur C. Sletten
- , Jessica W. Davidson
- & Jean E. Schaffer
-
Article
| Open Access RNase κ promotes robust piRNA production by generating 2′,3′-cyclic phosphate-containing precursors
RNase κ promotes robust piRNA production by generating 2′,3′-cyclic phosphate-containing precursors
Mature piRNAs are produced from long single-stranded precursor RNAs. Here the authors show that RNase κ generates 2′,3′-cyclic phosphate-containing piRNA precursors and promotes piRNA production.
- Megumi Shigematsu
- , Takuya Kawamura
- & Yohei Kirino
-
Article
| Open Access Small RNA mediated gradual control of lipopolysaccharide biosynthesis affects antibiotic resistance in Helicobacter pylori
Small RNA mediated gradual control of lipopolysaccharide biosynthesis affects antibiotic resistance in Helicobacter pylori
The small RNA RepG modulates expression of chemotaxis receptor TlpB in Helicobacter pylori by targeting a length-variable G-repeat in the tlpB mRNA. Here, Pernitzsch et al. show that RepG also gradually controls lipopolysaccharide biosynthesis, antibiotic susceptibility, and in-vivo colonization of the stomach, by regulating a gene that is co-transcribed with tlpB.
- Sandy R. Pernitzsch
- , Mona Alzheimer
- & Cynthia M. Sharma
-
Article
| Open Access Translation and codon usage regulate Argonaute slicer activity to trigger small RNA biogenesis
Translation and codon usage regulate Argonaute slicer activity to trigger small RNA biogenesis
22G-RNAs are single-stranded antisense small RNAs that are expressed in C. elegans germline. Here the authors show that CSR-1 dependent 22G-RNAs are produced in the cytosol on mRNAs actively engaged in translation and that codon usage of an mRNA regulates the biogenesis of CSR-1 dependent 22G-RNAs.
- Meetali Singh
- , Eric Cornes
- & Germano Cecere
-
Article
| Open Access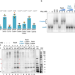 RNA binding of Hfq monomers promotes RelA-mediated hexamerization in a limiting Hfq environment
RNA binding of Hfq monomers promotes RelA-mediated hexamerization in a limiting Hfq environment
RelA stimulates RyhB small RNA–target mRNA interaction by promoting assembly of Hfq monomers into hexamers. Here the authors show that RelA-mediated Hfq hexamerization requires an initial binding of RNA to Hfq monomers.
- Pallabi Basu
- , Maya Elgrably-Weiss
- & Shoshy Altuvia
-
Article
| Open Access A bacterial small RNA regulates the adaptation of Helicobacter pylori to the host environment
A bacterial small RNA regulates the adaptation of Helicobacter pylori to the host environment
Long-term infection of the stomach with Helicobacter pylori can cause gastric cancer. Here, Kinoshita-Daitoku et al. show that a small non-coding RNA of H. pylori regulates bacterial adaptation to the stomach environment and bacterial oncoprotein production.
- Ryo Kinoshita-Daitoku
- , Kotaro Kiga
- & Hitomi Mimuro
-
Article
| Open Access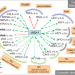 The box C/D snoRNP assembly factor Bcd1 interacts with the histone chaperone Rtt106 and controls its transcription dependent activity
The box C/D snoRNP assembly factor Bcd1 interacts with the histone chaperone Rtt106 and controls its transcription dependent activity
Biogenesis of small nucleolar RNAs ribonucleoproteins (snoRNPs) requires dedicated assembly machinery. Here, the authors show that a subset of snoRNP assembly factors interacts, genetically or directly, with factors modulating chromatin architecture, suggesting a link between ribosome formation and chromatin functions.
- Benoît Bragantini
- , Christophe Charron
- & Bruno Charpentier
-
Article
| Open Access Effects of individual base-pairs on in vivo target search and destruction kinetics of bacterial small RNA
Effects of individual base-pairs on in vivo target search and destruction kinetics of bacterial small RNA
Bacterial small RNA SgrS binds and regulates its primary target, ptsG mRNA. Here the authors employ Sort-Seq and super resolution imaging to investigate in vivo target recognition and rejection kinetics of SgrS.
- Anustup Poddar
- , Muhammad S. Azam
- & Taekjip Ha
-
Article
| Open Access Structure of the TFIIIC subcomplex τA provides insights into RNA polymerase III pre-initiation complex formation
Structure of the TFIIIC subcomplex τA provides insights into RNA polymerase III pre-initiation complex formation
Transcription factor TFIIIC plays roles in Pol III transcription and in chromatin organization. CryoEM structure of the yeast TFIIIC subcomplex τA, a negative stain reconstruction of τA bound to the TFIIIB subunits Brf1 and TBP and accompanying biochemistry suggest how τA achieves positioning of TFIIIB upstream of the TSS and remodeling of the TFIIIC complex during assembly of TFIIIB.
- Matthias K. Vorländer
- , Anna Jungblut
- & Christoph W. Müller
-
Article
| Open Access Jouvence a small nucleolar RNA required in the gut extends lifespan in Drosophila
Jouvence a small nucleolar RNA required in the gut extends lifespan in Drosophila
Small non-coding RNAs contribute to the regulation of aging. Here the authors identify a small nucleolar RNA, the snoRNA jouvence, which extends the lifespan of fruit flies through its function in the gut, and is conserved in humans.
- Stéphanie Soulé
- , Lucille Mellottée
- & Jean-René Martin
-
Article
| Open Access Argonaute bypasses cellular obstacles without hindrance during target search
Argonaute bypasses cellular obstacles without hindrance during target search
Argonaute protein is loaded with small RNA and scans long stretch of sequences to find complementary target sites. Here, using single-molecule FRET and kinetic modelling, the authors showed that prokaryotic Argonaute protein binds target DNA loosely and slides along the DNA during target search.
- Tao Ju Cui
- , Misha Klein
- & Chirlmin Joo
-
Article
| Open Access Modification of messenger RNA by 2′-O-methylation regulates gene expression in vivo
Modification of messenger RNA by 2′-O-methylation regulates gene expression in vivo
The post-transcriptional modification of mRNAs provides an additional layer to gene expression regulation. Here the authors show that 2′-O-methylation mediated by box C/D snoRNAs and fibrillarin can inhibit the translation of target mRNAs.
- Brittany A. Elliott
- , Hsiang-Ting Ho
- & Christopher L. Holley
-
Article
| Open Access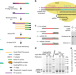 Single-cell CAS-seq reveals a class of short PIWI-interacting RNAs in human oocytes
Single-cell CAS-seq reveals a class of short PIWI-interacting RNAs in human oocytes
PIWI-interacting RNAs (piRNAs) are ~25–33 nt small RNAs expressed in animal germ cells. Here, the authors develop a single-cell small RNA sequencing method and report that a class of ~20-nt piRNAs lacking 3′ end 2′-O-methylation are associated with PIWIL3 protein and predominantly expressed in human and monkey oocytes.
- Qiyuan Yang
- , Ronghong Li
- & Ligang Wu
-
Article
| Open Access Modular one-pot assembly of CRISPR arrays enables library generation and reveals factors influencing crRNA biogenesis
Modular one-pot assembly of CRISPR arrays enables library generation and reveals factors influencing crRNA biogenesis
CRISPR array generation is difficult due to reoccurring repeat sequences. Here the authors present CRATES—a modular, one-pot assembly method—and demonstrate the creation of arrays for Cas9, Cas12a and Cas13a for cell-free, bacterial, yeast and mammalian systems.
- Chunyu Liao
- , Fani Ttofali
- & Chase L. Beisel
-
Article
| Open Access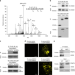 Dual regulation of Arabidopsis AGO2 by arginine methylation
Dual regulation of Arabidopsis AGO2 by arginine methylation
AGO2 is a core component of the RNAi machinery and contributes to plant immunity during bacterial infection. Here the authors show that AGO2 activity is suppressed by arginine methylation which not only promotes AGO2 degradation but also recruits TSN proteins to degrade AGO2-associated small RNAs.
- Po Hu
- , Hongwei Zhao
- & Hailing Jin
-
Article
| Open Access Distinct adaptive mechanisms drive recovery from aneuploidy caused by loss of the Ulp2 SUMO protease
Distinct adaptive mechanisms drive recovery from aneuploidy caused by loss of the Ulp2 SUMO protease
Transient aneuploidy enables cells to survive sudden environmental changes before longterm cellular adaptations are established. Here, the authors show that yeast cells respond to the acute loss of Ulp2 SUMO protease by rapid induction of aneuploidy, and reveal predictable long-term adaptation mechanisms that restore euploidy.
- Hong-Yeoul Ryu
- , Francesc López-Giráldez
- & Mark Hochstrasser
-
Article
| Open Access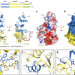 Structural and biochemical insights into small RNA 3′ end trimming by Arabidopsis SDN1
Structural and biochemical insights into small RNA 3′ end trimming by Arabidopsis SDN1
Small RNA degrading nucleases (SDNs) can degrade short RNAs. Here the authors report the crystal structure of Arabidopsis SDN1 in complex with a single-stranded RNA, and provide new insight into 3′ end trimming mechanism of 3′ to 5′ riboexonucleases in the metabolism of various species of small RNAs.
- Jiayi Chen
- , Li Liu
- & Jinbiao Ma
-
Article
| Open Access Gating of miRNA movement at defined cell-cell interfaces governs their impact as positional signals
Gating of miRNA movement at defined cell-cell interfaces governs their impact as positional signals
Movement of small RNA between cells is critical to plant development and stress responses. Here the authors uncover a gate-keeping mechanism that can restrict small RNA movement at cell-cell interfaces, providing selectivity in long-distance signalling and limiting the scope of local mobility.
- Damianos S. Skopelitis
- , Kristine Hill
- & Marja C. P. Timmermans
-
Article
| Open Access nc886 is induced by TGF-β and suppresses the microRNA pathway in ovarian cancer
nc886 is induced by TGF-β and suppresses the microRNA pathway in ovarian cancer
Ovarian cancers often display elevated TGF-β signaling but repressed miRNA expression. Here the authors identify that non-coding RNA nc886 expression is induced by TGF-β, which then binds to DICER and impairs miRNA maturation.
- Ji-Hye Ahn
- , Hyun-Sung Lee
- & Yong Sun Lee
-
Article
| Open Access Coding and noncoding landscape of extracellular RNA released by human glioma stem cells
Coding and noncoding landscape of extracellular RNA released by human glioma stem cells
While circulating DNA has been extensively explored as a potential cancer biomarker, RNA potential has been overlooked so far. Here the authors present a comprehensive analysis of extracellular RNA secreted by glioblastoma cells that could prove a valuable resource for biomarker discovery and a means of intercellular communication.
- Zhiyun Wei
- , Arsen O. Batagov
- & Anna M. Krichevsky
-
Article
| Open Access Identification of functional tetramolecular RNA G-quadruplexes derived from transfer RNAs
Identification of functional tetramolecular RNA G-quadruplexes derived from transfer RNAs
RNA G-quadruplexes (RG4) occur in vivo and have regulatory roles in mRNA metabolism. Here the authors show that the guanine residue stretches at the 5’ end of tRNA-derived stress-induced RNAs mediate the formation of tetramolecular RG4 structures, which play a role in the post-transcriptional regulation of gene expression.
- Shawn M. Lyons
- , Dorota Gudanis
- & Pavel Ivanov
-
Article
| Open Access Computational design of small transcription activating RNAs for versatile and dynamic gene regulation
Computational design of small transcription activating RNAs for versatile and dynamic gene regulation
The structural basis of RNA-based gene control offers the possibility of de novo design. Here the authors present a computational design approach for Small Transcription Activating RNAs a bacterial RNA regulator that allows for versatile and dynamic control of genes, pathways and genetic circuits.
- James Chappell
- , Alexandra Westbrook
- & Julius B. Lucks
-
Article
| Open Access Mutant p53 shapes the enhancer landscape of cancer cells in response to chronic immune signaling
Mutant p53 shapes the enhancer landscape of cancer cells in response to chronic immune signaling
Inflammation is known to affect cancer development, yet the mechanisms by which immune signaling drives transformation remain unclear. Here, the authors provide evidence that chronic TNF-α signaling promotes the enhancer binding and transcriptional interplay between mutant p53 and NFκB.
- Homa Rahnamoun
- , Hanbin Lu
- & Shannon M. Lauberth
-
Article
| Open Access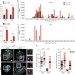 DNA damage response inhibition at dysfunctional telomeres by modulation of telomeric DNA damage response RNAs
DNA damage response inhibition at dysfunctional telomeres by modulation of telomeric DNA damage response RNAs
The DNA damage response (DDR) involves site-specific small non-coding RNAs. Here the authors show that telomere dysfunction induces transcription of telomeric DNA damage response RNAs that are necessary for DDR activation, which can be specifically muted by antisense inhibitory oligonucleotides.
- Francesca Rossiello
- , Julio Aguado
- & Fabrizio d’Adda di Fagagna
-
Article
| Open Access Diverse human extracellular RNAs are widely detected in human plasma
Diverse human extracellular RNAs are widely detected in human plasma
Extracellular miRNAs are present in a variety of bodily fluids. Here, Freedman et al. analysed plasma-derived RNA by RNA-seq from 40 people followed by targeted RT-qPCR in an additional 2,763 people, and report over 1,000 extracellular RNAs including microRNAs, piwi-interacting RNA and small nucleolar RNAs.
- Jane E. Freedman
- , Mark Gerstein
- & Kahraman Tanriverdi
-
Article |
 DSIF and NELF interact with Integrator to specify the correct post-transcriptional fate of snRNA genes
DSIF and NELF interact with Integrator to specify the correct post-transcriptional fate of snRNA genes
The elongation factors DSIF and NELF have established roles in polymerase pausing, elongation and 3'-end processing of replication-dependent histone mRNAs. Here the authors demonstrate that DSIF and NELF form a complex with Integrator and allow proper 3'-processing of snRNA transcripts by preventing the recruitment of CstF.
- Junichi Yamamoto
- , Yuri Hagiwara
- & Yuki Yamaguchi
-
Article |
 Sample sequencing of vascular plants demonstrates widespread conservation and divergence of microRNAs
Sample sequencing of vascular plants demonstrates widespread conservation and divergence of microRNAs
Small RNAs and microRNAs are important regulators of gene expression. In this study, Chávez Montes et al.examine these molecules in 34 plant species, and explore the correlations between abundance, conservation and variability of microRNA sequences across all of the species studied.
- Ricardo A. Chávez Montes
- , de Fátima Flor Rosas-Cárdenas
- & Stefan de Folter

 RNA compaction and iterative scanning for small RNA targets by the Hfq chaperone
RNA compaction and iterative scanning for small RNA targets by the Hfq chaperone