Featured
-
-
Article
| Open Access Site-specific identification and quantitation of endogenous SUMO modifications under native conditions
Site-specific identification and quantitation of endogenous SUMO modifications under native conditions
SUMOylation is post-translational modification implicated in several biological pathways. Here the authors describe an approach for the global profiling of SUMO attachment sites under native conditions that also allows the parallel determination of SUMO and Ub attachments.
- Ryan J. Lumpkin
- , Hongbo Gu
- & Elizabeth A. Komives
-
Article
| Open Access Evolution of AF6-RAS association and its implications in mixed-lineage leukemia
Evolution of AF6-RAS association and its implications in mixed-lineage leukemia
Several rearrangements of the MLL gene are associated with acute leukemia, including the fusion of MLL with a RAS effector protein, AF6. Here the authors show that the truncated AF6 can induce AF6-MLL dimerization and drive its oncogenic activity.
- Matthew J. Smith
- , Elizabeth Ottoni
- & Mitsuhiko Ikura
-
Article
| Open Access An HDAC3-PROX1 corepressor module acts on HNF4α to control hepatic triglycerides
An HDAC3-PROX1 corepressor module acts on HNF4α to control hepatic triglycerides
HDAC3 is a critical mediator of hepatic lipid metabolism and its loss leads to fatty liver. Here, the authors characterize the liver HDAC3 interactome in vivo, provide evidence that HDAC3 interacts with PROX1, and show that HDAC3 and PROX1 control expression of genes regulating lipid homeostasis.
- Sean M. Armour
- , Jarrett R. Remsberg
- & Mitchell A. Lazar
-
Article
| Open Access Split-BioID a conditional proteomics approach to monitor the composition of spatiotemporally defined protein complexes
Split-BioID a conditional proteomics approach to monitor the composition of spatiotemporally defined protein complexes
The BioID approaches takes advantage of the promiscuous biotinylation enzyme (BirA*) to identify proteins that closely interact. Here the authors improve the resolution of BioID using a protein fragment complementation approach that allows the assignment of protein-protein interactions to specific complexes within a common interactome.
- Isabel Myriam Schopp
- , Cinthia Claudia Amaya Ramirez
- & Julien Béthune
-
Article
| Open Access R2TP/Prefoldin-like component RUVBL1/RUVBL2 directly interacts with ZNHIT2 to regulate assembly of U5 small nuclear ribonucleoprotein
R2TP/Prefoldin-like component RUVBL1/RUVBL2 directly interacts with ZNHIT2 to regulate assembly of U5 small nuclear ribonucleoprotein
The R2TP/Prefoldin-like cochaperone complex is involved in the assembly of a number of protein complexes. Here the authors provide evidence that RUVBL1/RUVBL2, subunits of that cochaperone complex, directly interact with ZNHIT2 to regulate assembly of U5 small ribonucleoprotein.
- Philippe Cloutier
- , Christian Poitras
- & Benoit Coulombe
-
Article
| Open Access WIPI3 and WIPI4 β-propellers are scaffolds for LKB1-AMPK-TSC signalling circuits in the control of autophagy
WIPI3 and WIPI4 β-propellers are scaffolds for LKB1-AMPK-TSC signalling circuits in the control of autophagy
During autophagy, AMPK and mTOR associate with ULK1 and regulate phosphatidylinositol 3-phosphate (PtdIns3P) production that mediates autophagosome formation via WIPI proteins. Here the authors show WIPI3 and WIPI4 have a scaffolding function upstream of PtdIns3P production and have a role in the PtdIns3P effector function of WIPI1-WIPI2 at nascent autophagosomes.
- Daniela Bakula
- , Amelie J. Müller
- & Tassula Proikas-Cezanne
-
Article
| Open Access Optimized fragmentation schemes and data analysis strategies for proteome-wide cross-link identification
Optimized fragmentation schemes and data analysis strategies for proteome-wide cross-link identification
Chemical cross-linking combined with mass spectrometry (XL-MS) can provide information on protein conformations and interactions in highly complex samples. Here the authors describe an improved XL-MS workflow to increase the depth and fidelity of cross-link identification using whole proteome databases.
- Fan Liu
- , Philip Lössl
- & Albert J. R. Heck
-
Article
| Open Access Defective Gpsm2/Gαi3 signalling disrupts stereocilia development and growth cone actin dynamics in Chudley-McCullough syndrome
Defective Gpsm2/Gαi3 signalling disrupts stereocilia development and growth cone actin dynamics in Chudley-McCullough syndrome
Mutations inGPSM2cause a rare disease characterized by deafness and brain abnormalities. Here the authors show that Gpsm2 forms a molecular complex with a heterotrimeric G-protein subunit, whirlin and a myosin motor to regulate actin dynamics in neurons and auditory hair cell stereocilia.
- Stephanie A. Mauriac
- , Yeri E. Hien
- & Mireille Montcouquiol
-
Article
| Open Access Comparative influenza protein interactomes identify the role of plakophilin 2 in virus restriction
Comparative influenza protein interactomes identify the role of plakophilin 2 in virus restriction
Protein interaction networks can identify host proteins that affect virus replication. Here, the authors compare the protein interactomes of several influenza A virus strains and identify plakophilin 2 as a restriction factor that inhibits formation of the viral polymerase complex.
- Lingyan Wang
- , Bishi Fu
- & Shitao Li
-
Article
| Open Access In vivo protein interaction network analysis reveals porin-localized antibiotic inactivation in Acinetobacter baumannii strain AB5075
In vivo protein interaction network analysis reveals porin-localized antibiotic inactivation in Acinetobacter baumannii strain AB5075
The bacterial pathogen Acinetobacter baumannii has evolved resistance to many antibiotics, including carbapenems. Here, Wu et al. show that the carbapenemase Oxa-23 interacts with the outer membrane porin CarO in an A. baumanniiisolate, indicative of porin-localised antibiotic inactivation.
- Xia Wu
- , Juan D. Chavez
- & James E. Bruce
-
Article
| Open Access Genetically encoded protein photocrosslinker with a transferable mass spectrometry-identifiable label
Genetically encoded protein photocrosslinker with a transferable mass spectrometry-identifiable label
Mapping protein-protein interaction using crosslinking and mass spectroscopy strategies is hampered by a high rate of false-positive results. Here, the authors develop a genetically encoded photo-affinity probe for accurate identification of protein interaction partners and crosslinking sites.
- Yi Yang
- , Haiping Song
- & Peng R. Chen
-
Article
| Open Access Magneto-nanosensor platform for probing low-affinity protein–protein interactions and identification of a low-affinity PD-L1/PD-L2 interaction
Magneto-nanosensor platform for probing low-affinity protein–protein interactions and identification of a low-affinity PD-L1/PD-L2 interaction
Measurement of low-affinity protein–protein interactions is challenging; surface plasmon resonance requires high concentrations of reagents. Here the authors combine magneto-nanosensors with microfluidic chips and protein-conjugated magnetic nanoparticles to discover a low-affinity interaction between T-cell inhibitory receptors PD-L1 and PD-L2.
- Jung-Rok Lee
- , Daniel J. B. Bechstein
- & Shan X. Wang
-
Article
| Open Access 14-3-3 proteins regulate Tctp–Rheb interaction for organ growth in Drosophila
14-3-3 proteins regulate Tctp–Rheb interaction for organ growth in Drosophila
14-3-3 proteins regulate several signalling pathways but often act redundantly; however, the molecular mechanisms behind such redundancy are unclear. Here, the authors show that 14-3-3 proteins regulate two interacting components of Tor signalling in Drosophila, Tctp and Rheb, disrupting organ development.
- Thao Phuong Le
- , Linh Thuong Vuong
- & Kwang-Wook Choi
-
Article
| Open Access The Ku-binding motif is a conserved module for recruitment and stimulation of non-homologous end-joining proteins
The Ku-binding motif is a conserved module for recruitment and stimulation of non-homologous end-joining proteins
Werner syndrome is a progeroid disease characterised by genetic instability due to mutations to the WRN helicase/exonuclease. Here the authors define a novel Ku binding motif (KBM) and show that two such motifs facilitate the involvement of WRN in DNA double-strand break repair.
- Gabrielle J. Grundy
- , Stuart L. Rulten
- & Keith W. Caldecott
-
Article
| Open Access BTG2 bridges PABPC1 RNA-binding domains and CAF1 deadenylase to control cell proliferation
BTG2 bridges PABPC1 RNA-binding domains and CAF1 deadenylase to control cell proliferation
BTG2 promotes mRNA poly(A) tail shortening and regulates cellular differentiation. Here, Stupfler et al. show that the BTG2 APRO domain interacts with PABPC1 RRM1, allowing the former to recruit and stimulate the poly(A) tail shortening activity of CAF1 deadenylase and to control cell proliferation.
- Benjamin Stupfler
- , Catherine Birck
- & Fabienne Mauxion
-
Article
| Open Access Chemical basis for the recognition of trimethyllysine by epigenetic reader proteins
Chemical basis for the recognition of trimethyllysine by epigenetic reader proteins
A structurally diverse set of epigenetic reader proteins can recognize methylated lysine residues on histones. Here the authors show that recognition of trimethyllysine occurs through a combination of favourable cation–πinteractions and the release of water molecules occupying the aromatic cages of reader proteins.
- Jos J.A.G. Kamps
- , Jiaxin Huang
- & Jasmin Mecinović
-
Article
| Open Access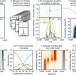 Quantifying the stabilizing effects of protein–ligand interactions in the gas phase
Quantifying the stabilizing effects of protein–ligand interactions in the gas phase
Relatively few techniques can quantitatively measure the effect of ligands on membrane protein stability. Here the authors demonstrate the use of ion-mobility mass spectrometry to accurately measure and quantify ligand-induced protein stabilization in the gas phase.
- Timothy M. Allison
- , Eamonn Reading
- & Carol V. Robinson
-
Article
| Open Access CPI motif interaction is necessary for capping protein function in cells
CPI motif interaction is necessary for capping protein function in cells
Capping protein regulates actin filament dynamics by binding to barbed ends and preventing their growth. Edwards et al. show that capping protein also requires interactions with proteins containing a capping protein interaction motif to promote its proper localization and regulation of actin dynamics.
- Marc Edwards
- , Patrick McConnell
- & John A. Cooper
-
Article |
 Multiple cellular proteins interact with LEDGF/p75 through a conserved unstructured consensus motif
Multiple cellular proteins interact with LEDGF/p75 through a conserved unstructured consensus motif
LEDGF/p75, a protein involved in HIV integration and leukaemia, interacts with various cellular proteins via its integrase binding domain (IBD). Here, the authors show that the interaction is mediated by an intrinsically disordered IBD-binding motif (IBM) on all known cellular partners of LEDGF/p75.
- Petr Tesina
- , Kateřina Čermáková
- & Pavlína Řezáčová
-
Article
| Open Access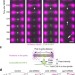 Direct interaction between centralspindlin and PRC1 reinforces mechanical resilience of the central spindle
Direct interaction between centralspindlin and PRC1 reinforces mechanical resilience of the central spindle
The central spindle is an anti parallel bundle of microtubules that forms between segregating chromosomes and links the two halves of the mitotic spindle. Lee et al.reveal that interaction between two microtubule bundling proteins at the central spindle confers robustness to cortical pulling forces.
- Kian-Yong Lee
- , Behrooz Esmaeili
- & Masanori Mishima
-
Article
| Open Access Proximity-dependent initiation of hybridization chain reaction
Proximity-dependent initiation of hybridization chain reaction
Proximity ligation assays are a sensitive method for detecting protein interactions, but require the addition of enzymes. Here the authors introduce proxHCR, an enzyme-free method of detecting interactions in close proximity by inducing a hybribization chain reaction (HCR) of fluorescently labelled oligonucleotides.
- Björn Koos
- , Gaëlle Cane
- & Ola Söderberg
-
Article
| Open Access Extreme multifunctional proteins identified from a human protein interaction network
Extreme multifunctional proteins identified from a human protein interaction network
Proteins are sometimes implicated in separate and seemingly unrelated processes, so called moonlighting functions. Here the authors use bioinformatics tools to identify extreme multifunctional proteins and define a signature of extreme multifunctionality.
- Charles E. Chapple
- , Benoit Robisson
- & Christine Brun
-
Article |
 Engineered pairs of distinct photoswitches for optogenetic control of cellular proteins
Engineered pairs of distinct photoswitches for optogenetic control of cellular proteins
Photoreceptor-based photoswitches have proved to be powerful tools for the specific control of protein activity in live cells. Here the authors describe Magnets, a new set of photoswitches based on the Vivid photoreceptor with enhanced hetero-dimerization specificity and variable activation kinetics.
- Fuun Kawano
- , Hideyuki Suzuki
- & Moritoshi Sato
-
Article |
 Pre-B cell receptor binding to galectin-1 modifies galectin-1/carbohydrate affinity to modulate specific galectin-1/glycan lattice interactions
Pre-B cell receptor binding to galectin-1 modifies galectin-1/carbohydrate affinity to modulate specific galectin-1/glycan lattice interactions
Galectin-1 (GAL1) is a secreted protein that binds to glycans and to the pre-B-cell receptor (pre-BCR). Here Bonzi et al. show that pre-BCR binding to GAL1 causes a conformational change in the GAL1 carbohydrate-binding site to inhibit binding to selected glycans.
- Jeremy Bonzi
- , Olivier Bornet
- & Latifa Elantak
-
Article
| Open Access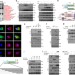 Small heterodimer partner interacts with NLRP3 and negatively regulates activation of the NLRP3 inflammasome
Small heterodimer partner interacts with NLRP3 and negatively regulates activation of the NLRP3 inflammasome
Excessive NLRP3 inflammasome activation underlies inflammatory diseases such as gout. Here the authors show that orphan nuclear receptor small heterodimer partner protein (SHP) negatively regulates NLRP3, and its loss leads to accumulation of damaged mitochondria and gout-like immunopathology.
- Chul-Su Yang
- , Jwa-Jin Kim
- & Eun-Kyeong Jo
-
Article |
 Tumour suppressor TRIM33 targets nuclear β-catenin degradation
Tumour suppressor TRIM33 targets nuclear β-catenin degradation
Aberrant activation of β-catenin in the nucleus has been implicated in several cancers, but the mechanisms regulating nuclear β-catenin are not well understood. Here the authors identify Trim33 as new E3 ligase targeting nuclear β-catenin independently of Wnt signal.
- Jianfei Xue
- , Yaohui Chen
- & Suyun Huang
-
Article
| Open Access Super-resolution imaging and tracking of protein–protein interactions in sub-diffraction cellular space
Super-resolution imaging and tracking of protein–protein interactions in sub-diffraction cellular space
Protein–protein interactions are ubiquitous in cells and these contacts are crucial for a wide number of cellular processes. Here, the authors present a technique for the super-resolution imaging and tracking of protein–protein interactions in cells.
- Zhen Liu
- , Dong Xing
- & Yujie Sun
-
Article
| Open Access Protein interaction network of alternatively spliced isoforms from brain links genetic risk factors for autism
Protein interaction network of alternatively spliced isoforms from brain links genetic risk factors for autism
Autism spectrum disorder (ASD) is a complex genetic trait that encompasses a range of neurodevelopmental disorders. Here, the authors clone brain-expressed alternatively-spliced isoforms of ASD risk factors and construct a network of protein interactions that provides further insight into the disease aetiology.
- Roser Corominas
- , Xinping Yang
- & Lilia M. Iakoucheva
-
Article |
 Cholesterol modulates cell signaling and protein networking by specifically interacting with PDZ domain-containing scaffold proteins
Cholesterol modulates cell signaling and protein networking by specifically interacting with PDZ domain-containing scaffold proteins
Cholesterol indirectly regulates intracellular signalling by modulating the physical properties of lipid membranes. Sheng et al.now show that many PDZ domains contain a functional cholesterol-binding motif, revealing that cholesterol can also control the localization and function of signalling proteins directly.
- Ren Sheng
- , Yong Chen
- & Wonhwa Cho
-
Article |
 Protein-binding assays in biological liquids using microscale thermophoresis
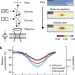
Protein-binding assays in biological liquids using microscale thermophoresis
Protein interactions in biological environments are expected to differ from the situationin vitro. In this study, a thermophoresis-based technique is described that allows the analysis of protein and small-molecule interactions in biological liquids; the work may allow more efficient drug development.
- Christoph J. Wienken
- , Philipp Baaske
- & Stefan Duhr

 Quantitative analysis of protein-protein interactions and post-translational modifications in rare immune populations
Quantitative analysis of protein-protein interactions and post-translational modifications in rare immune populations