Featured
-
-
Article
| Open Access NSD2 overexpression drives clustered chromatin and transcriptional changes in a subset of insulated domains
NSD2 overexpression drives clustered chromatin and transcriptional changes in a subset of insulated domains
In multiple myeloma, a 4;14 translocation induces overexpression of histone methyltransferase NSD2, resulting in expansion of H3K36me2 and shrinkage of H3K27me3 domains. Here the authors find that CTCF, H3K27ac and gene expression changes cluster within a subset of insulated domains implicating 3D chromosome organization as a key factor in the NSD2-mediated phenotype.
- Priscillia Lhoumaud
- , Sana Badri
- & Jane A. Skok
-
Article
| Open Access Transcriptomic profiling of the myeloma bone-lining niche reveals BMP signalling inhibition to improve bone disease
Transcriptomic profiling of the myeloma bone-lining niche reveals BMP signalling inhibition to improve bone disease
Multiple myeloma is a cancer of the bone marrow that can induce bone disease. Here, the authors profile the transcriptome of bone-lining cells and find a targetable role of bone morphogenetic protein (BMP) signalling in myeloma-induced bone-disease
- Sarah Gooding
- , Sam W. Z. Olechnowicz
- & Claire M. Edwards
-
Article
| Open Access Genomic landscape and chronological reconstruction of driver events in multiple myeloma
Genomic landscape and chronological reconstruction of driver events in multiple myeloma
Multiple myeloma evolves continuously. Here the authors chronologically reconstruct driver events in multiple myeloma, noting a limited repertoire of initiating driver events that shape the evolutionary trajectory of the disease.
- Francesco Maura
- , Niccoló Bolli
- & Peter J. Campbell
-
Article
| Open Access A practical guide for mutational signature analysis in hematological malignancies
A practical guide for mutational signature analysis in hematological malignancies
Mutational signature analysis provides important information about the mutational processes underpinning different stages of tumorigenesis. Here, the authors compare publicly available signature extraction tools and suggest a framework for the generation of accurate and reproducible signature data.
- Francesco Maura
- , Andrea Degasperi
- & Niccolò Bolli
-
Article
| Open Access Multiple myeloma immunoglobulin lambda translocations portend poor prognosis
Multiple myeloma immunoglobulin lambda translocations portend poor prognosis
Multiple myeloma is frequently characterised by translocation of genes next to the immunoglobulin heavy chain locus. In this study, the authors sequence a large cohort of high risk myeloma samples and find translocations of cMyc to the immunoglobulin heavy chain locus and this is associated with poor prognosis.
- Benjamin G. Barwick
- , Paola Neri
- & Lawrence H. Boise
-
Article
| Open Access Discovery of Mcl-1-specific inhibitor AZD5991 and preclinical activity in multiple myeloma and acute myeloid leukemia
Discovery of Mcl-1-specific inhibitor AZD5991 and preclinical activity in multiple myeloma and acute myeloid leukemia
High expression of Mcl-1 promotes tumorigenesis and resistance to anticancer therapies. Here they report a macrocyclic molecule with high selectivity and affinity for Mcl-1 that exhibits potent anti-tumor effects as single agent and in combination with bortezomib or venetoclax in preclinical models of multiple myeloma and acute myeloid leukemia.
- Adriana E. Tron
- , Matthew A. Belmonte
- & Alexander W. Hird
-
Article
| Open Access Microbiota-driven interleukin-17-producing cells and eosinophils synergize to accelerate multiple myeloma progression
Microbiota-driven interleukin-17-producing cells and eosinophils synergize to accelerate multiple myeloma progression
The mechanisms through which gut microbiota affect extramucosal tumors are poorly understood. Here the authors show that the gut microbiota promotes multiple myeloma by inducing differentiation and migration of Th17 cells in the bone marrow resulting also in increased recruitment of pro-tumorigenic eosinophils.
- Arianna Calcinotto
- , Arianna Brevi
- & Matteo Bellone
-
Article
| Open Access Identification of multiple risk loci and regulatory mechanisms influencing susceptibility to multiple myeloma
Identification of multiple risk loci and regulatory mechanisms influencing susceptibility to multiple myeloma
Multiple myeloma is a cancer of the plasma cells, and the complete aetiology of the disease is still unclear. Here the authors perform an additional GWAS analysis followed by a meta-analysis with existing GWAS and replication genotyping and identify 6 novel risk loci and utilise gene expression, epigenetic profiling and in situ Hi-C data to further our understanding of MM susceptibility.
- Molly Went
- , Amit Sud
- & Stephen N. Thibodeau
-
Article
| Open Access Genomic patterns of progression in smoldering multiple myeloma
Genomic patterns of progression in smoldering multiple myeloma
Smoldering MM (SMM) is a premalignant stage of multiple myeloma (MM). Here the authors perform whole genome sequencing of unique paired samples of SMM progressing to MM, and show that the genomic landscape at the SMM stage is very similar to MM, but trajectories of evolution can vary from patient to patient.
- Niccolò Bolli
- , Francesco Maura
- & Nikhil Munshi
-
Article
| Open Access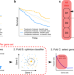 Predicting treatment benefit in multiple myeloma through simulation of alternative treatment effects
Predicting treatment benefit in multiple myeloma through simulation of alternative treatment effects
Selection of the right cancer treatment is still a challenge. Here, the authors introduce a framework to analyze treatment benefits, using the idea that patients with similar genetic tumor profiles receiving different treatments can be used to model their responses to the alternative treatment.
- Joske Ubels
- , Pieter Sonneveld
- & Jeroen de Ridder
-
Article
| Open Access Whole-exome sequencing of cell-free DNA and circulating tumor cells in multiple myeloma
Whole-exome sequencing of cell-free DNA and circulating tumor cells in multiple myeloma
Circulating tumor cells (CTCs) and cell-free DNA (cfDNA) enables characterization of a patient’s cancer. Here, the authors analyse CTCs, cfDNA, and tumor biopsies from multiple myeloma patients to show these approaches are complementary for mutation detection, together enabling a greater fraction of patient tumors to be profiled.
- S. Manier
- , J. Park
- & I. M. Ghobrial
-
Article
| Open Access The multiple myeloma risk allele at 5q15 lowers ELL2 expression and increases ribosomal gene expression
The multiple myeloma risk allele at 5q15 lowers ELL2 expression and increases ribosomal gene expression
ELL2 was recently discovered as a susceptibility gene for multiple myeloma (MM). Here, they show that the MM risk allele lowers ELL2 expression in plasma cells, that it also upregulates gene sets related to ribosome biogenesis, and that one of the linked variants reduces binding of MAFF/G/K family transcription factors.
- Mina Ali
- , Ram Ajore
- & Björn Nilsson
-
Article
| Open Access Ikaros family zinc finger 1 regulates dendritic cell development and function in humans
Ikaros family zinc finger 1 regulates dendritic cell development and function in humans
IKZF1 is a transcription factor known to regulate mammalian B-cell development. Here the authors show that IKZF1 is required for human pDC development and regulation of DC cytokine production in patients with IKZF1 haploinsufficiency, findings which are recapitulated in lenalidomide-induced IKZF1 deficiency.
- Urszula Cytlak
- , Anastasia Resteu
- & Venetia Bigley
-
Article
| Open Access Alu-dependent RNA editing of GLI1 promotes malignant regeneration in multiple myeloma
Alu-dependent RNA editing of GLI1 promotes malignant regeneration in multiple myeloma
The treatment of multiple myeloma is challenging due to high relapse rates. Here the authors show that expression of ADAR1 correlates with poor patient outcomes, and that ADAR1-mediated editing of GLI1 is a mechanism relevant in the context of multiple myeloma progression and drug resistance.
- Elisa Lazzari
- , Phoebe K. Mondala
- & Catriona H. M. Jamieson
-
Article
| Open Access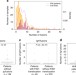 Expressed fusion gene landscape and its impact in multiple myeloma
Expressed fusion gene landscape and its impact in multiple myeloma
Multiple myeloma is a malignancy of plasma cells in the blood. Here, the authors establish the landscape of fusion genes within this disease, identifying novel recurrent fusion genes that impact survival and may drive disease progression.
- A. Cleynen
- , R. Szalat
- & H. Avet-Loiseau
-
Article
| Open Access Determining therapeutic susceptibility in multiple myeloma by single-cell mass accumulation
Determining therapeutic susceptibility in multiple myeloma by single-cell mass accumulation
Multiple myeloma is characterized by high rates of drug resistance and relapse. Here the authors utilize a functional assay to assess the ex vivo drug sensitivity of single multiple myeloma cells based on measuring the mass accumulation rate of individual cells.
- Arif E. Cetin
- , Mark M. Stevens
- & Scott R. Manalis
-
Article
| Open Access The non-canonical poly(A) polymerase FAM46C acts as an onco-suppressor in multiple myeloma
The non-canonical poly(A) polymerase FAM46C acts as an onco-suppressor in multiple myeloma
FAM46C is one of the most frequently mutated genes in multiple myeloma (MM), but its molecular function remains unknown. Here the authors show that FAM46C is a poly(A) polymerase and that loss of function of FAM46C drives multiple myeloma through the destabilisation of ER response transcripts.
- Seweryn Mroczek
- , Justyna Chlebowska
- & Andrzej Dziembowski
-
Article
| Open Access Spatial genomic heterogeneity in multiple myeloma revealed by multi-region sequencing
Spatial genomic heterogeneity in multiple myeloma revealed by multi-region sequencing
In multiple myeloma, malignant cells expand within bone marrow. Here, the authors use multi-region sequencing in patient samples to analyse spatial clonal architecture and heterogeneity, providing novel insight into multiple myeloma progression and evolution.
- L. Rasche
- , S. S. Chavan
- & N. Weinhold
-
Article
| Open Access Multiple myeloma risk variant at 7p15.3 creates an IRF4-binding site and interferes with CDCA7L expression
Multiple myeloma risk variant at 7p15.3 creates an IRF4-binding site and interferes with CDCA7L expression
Genome wide association studies have identified multiple risk loci for multiple myeloma. Here, the authors show that the expression of CDCA7Lis associated with patient survival and expression of the gene is influenced by a risk variant at 7p15.3, which creates a transcription factor binding site for IRF4.
- Ni Li
- , David C. Johnson
- & Richard S. Houlston
-
Article
| Open Access Genome-wide association study identifies multiple susceptibility loci for multiple myeloma
Genome-wide association study identifies multiple susceptibility loci for multiple myeloma
Previous genome-wide association studies have identified loci associated with the risk of multiple myeloma. Here, the authors present a meta-analysis of six genome wide association studies of the disease and identify eight new loci; functional studies identify genes as candidates for the basis of these associations.
- Jonathan S. Mitchell
- , Ni Li
- & Richard S. Houlston
-
Article
| Open Access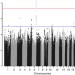 Genome-wide association study identifies variation at 6q25.1 associated with survival in multiple myeloma
Genome-wide association study identifies variation at 6q25.1 associated with survival in multiple myeloma
The prognosis of multiple myeloma patients varies widely. Here, to identify genetic factors associated with differing prognoses, the authors carried out a meta-analysis of four genome-wide association studies and identified a risk variant associated with survival interval.
- David C. Johnson
- , Niels Weinhold
- & Gareth J. Morgan
-
Article
| Open Access The KDM3A–KLF2–IRF4 axis maintains myeloma cell survival
The KDM3A–KLF2–IRF4 axis maintains myeloma cell survival
Several histone modifiers have been implicated in the survival of multiple myeloma cells. Here, the authors reveal a role for the histone demethylase KDM3A in the survival of this haematologic cancer, and show that mechanistically KDM3A removes H3K9 methylation from the promoters of KLF2 and IRF4, genes essential for myeloma cell survival.
- Hiroto Ohguchi
- , Teru Hideshima
- & Kenneth C. Anderson
-
Article
| Open Access Osteoclasts control reactivation of dormant myeloma cells by remodelling the endosteal niche
Osteoclasts control reactivation of dormant myeloma cells by remodelling the endosteal niche
Therapy resistant dormant myeloma cells contribute to disease relapse. Here, the authors use intravital microscopy to track the location of these cells and demonstrate that they hone to the endosteal niche within the bone.
- Michelle A. Lawson
- , Michelle M. McDonald
- & Peter I. Croucher
-
Article |
 Transcriptional repression by the HDAC4–RelB–p52 complex regulates multiple myeloma survival and growth
Transcriptional repression by the HDAC4–RelB–p52 complex regulates multiple myeloma survival and growth
NF-κB has largely been known as a transcriptional activator. Here the authors show that a transcriptionally repressive NF-κB complex, HDAC4–RelB–p52, maintains repressive chromatin at proapoptotic genes Bim and BMF, and regulates multiple myeloma survival and growth in an ERK1 dependent manner.
- Subrahmanya D. Vallabhapurapu
- , Sunil K. Noothi
- & Sivakumar Vallabhapurapu
-
Article |
 Genome-wide association study identifies variants at 16p13 associated with survival in multiple myeloma patients
Genome-wide association study identifies variants at 16p13 associated with survival in multiple myeloma patients
Multiple myeloma is an incurable blood cancer with family history being a strong contributing risk factor. Here Ziv et al.perform a genome-wide association study for genetic variation associated with myeloma survival, identifying FOPNL variants associated with worse clinical outcomes.
- Elad Ziv
- , Eric Dean
- & Celine Vachon
-
Article
| Open Access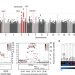 Variants in ELL2 influencing immunoglobulin levels associate with multiple myeloma
Variants in ELL2 influencing immunoglobulin levels associate with multiple myeloma
Multiple myeloma is an incurable and fatal disease characterized by uninhibited growth of plasma cells in the bone marrow. Here, Swaminathan et al. conduct a genome-wide association study and identify a novel risk locus at ELL2, which encodes a key component of the super-elongation complex.
- Bhairavi Swaminathan
- , Guðmar Thorleifsson
- & Björn Nilsson
-
Article |
 APOBEC family mutational signatures are associated with poor prognosis translocations in multiple myeloma
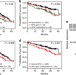
APOBEC family mutational signatures are associated with poor prognosis translocations in multiple myeloma
Rearrangements of the Ig loci are essential for generating antibody diversity but abnormal translocations can be a driving event for myeloma. Here Walker et al. perform whole exome sequencing on myeloma patients to capture the diversity of mutational changes.
- Brian A. Walker
- , Christopher P. Wardell
- & Gareth J. Morgan
-
Article
| Open Access Heterogeneity of genomic evolution and mutational profiles in multiple myeloma
Heterogeneity of genomic evolution and mutational profiles in multiple myeloma
Multiple myeloma is a malignant plasma cell disorder with a complex molecular pathogenesis. Here, the authors perform whole-exome sequencing, copy-number profiling and cytogenetic analysis in 84 myeloma samples and highlight the diversity and evolution of the mutational profile underlying the disease.
- Niccolo Bolli
- , Hervé Avet-Loiseau
- & Nikhil C. Munshi
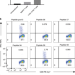
 Anti-BCMA chimeric antigen receptors with fully human heavy-chain-only antigen recognition domains
Anti-BCMA chimeric antigen receptors with fully human heavy-chain-only antigen recognition domains