Featured
-
-
Article
| Open Access A qPCR technology for direct quantification of methylation in untreated DNA
A qPCR technology for direct quantification of methylation in untreated DNA
Analysis of DNA methylation usually requires a chemical or an enzymatic pretreatment step. Here, the authors report a PCR-based technology for the detection of DNA methylation in untreated DNA, and present analytical and clinical results from methylation analysis of the MGMT promoter.
- Kamilla Kolding Bendixen
- , Maria Mindegaard
- & Rasmus Koefoed Petersen
-
Article
| Open Access Droplet-based bisulfite sequencing for high-throughput profiling of single-cell DNA methylomes
Droplet-based bisulfite sequencing for high-throughput profiling of single-cell DNA methylomes
Single-cell DNA methylomic studies offer high resolution to differentiate cell subsets based on their epigenomic features. Here, the authors demonstrate Drop-BS, a droplet-based single-cell bisulfite sequencing library preparation method, for DNA methylome profiling.
- Qiang Zhang
- , Sai Ma
- & Chang Lu
-
Article
| Open Access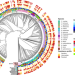 Roving methyltransferases generate a mosaic epigenetic landscape and influence evolution in Bacteroides fragilis group
Roving methyltransferases generate a mosaic epigenetic landscape and influence evolution in Bacteroides fragilis group
Here, Tisza, Dekker, and colleagues perform large scale analysis of genome methylation in the gut commensal and pathogen, Bacteroides fragilis group, revealing immense methyl motif diversity and evidence of widespread methyltransferase exchange among phages.
- Michael J. Tisza
- , Derek D. N. Smith
- & John P. Dekker
-
Article
| Open Access High-throughput robust single-cell DNA methylation profiling with sciMETv2
High-throughput robust single-cell DNA methylation profiling with sciMETv2
Despite the importance of DNA methylation, accessible and high-throughput methods to profile methylation at the single-cell level are lacking. Here, the authors present sciMETv2, a high-throughput workflow that provides high-quality single-cell methylomes in a robust and simple workflow.
- Ruth V. Nichols
- , Brendan L. O’Connell
- & Andrew C. Adey
-
Article
| Open Access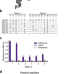 High-affinity chromodomains engineered for improved detection of histone methylation and enhanced CRISPR-based gene repression
High-affinity chromodomains engineered for improved detection of histone methylation and enhanced CRISPR-based gene repression
Engineered chromodomains are promising probes to analyze histone methylation. Here the authors designed high-affinity chromodomains for enhanced genome-wide binding analysis, live-cell imaging and ultra-potent CRISPR-based gene repression.
- G. Veggiani
- , R. Villaseñor
- & S. S. Sidhu
-
Article
| Open Access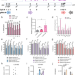 Unraveling the functional role of DNA demethylation at specific promoters by targeted steric blockage of DNA methyltransferase with CRISPR/dCas9
Unraveling the functional role of DNA demethylation at specific promoters by targeted steric blockage of DNA methyltransferase with CRISPR/dCas9
The causal relationship between DNA demethylation and gene expression regulation has not yet been fully resolved. Here the authors develop a nuclease-dead Cas9 (dCas9) and gRNA site-specific targeting approach to physically block DNA methylation at specific promoters to cause DNA demethylation in cells and tackle this question.
- Daniel M. Sapozhnikov
- & Moshe Szyf
-
Article
| Open Access EpiScanpy: integrated single-cell epigenomic analysis
EpiScanpy: integrated single-cell epigenomic analysis
The authors present epiScanpy: a computational framework for the analysis of single-cell epigenomic data, both ATAC-seq and DNA methylation data, with examples for clustering, cell type identification, trajectory learning and atlas integration - and show its performance in distinguishing cell types.
- Anna Danese
- , Maria L. Richter
- & Maria Colomé-Tatché
-
Article
| Open Access DNA methylation predicts age and provides insight into exceptional longevity of bats
DNA methylation predicts age and provides insight into exceptional longevity of bats
DNA methylation profiles from 26 bat species accurately predicts chronological age, while longevity-related methylation patterns across the genome suggest that bat longevity results from augmented immune response and cancer suppression.
- Gerald S. Wilkinson
- , Danielle M. Adams
- & Steve Horvath
-
Article
| Open Access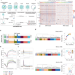 Single-cell multiomics sequencing reveals the functional regulatory landscape of early embryos
Single-cell multiomics sequencing reveals the functional regulatory landscape of early embryos
Extensive epigenetic reprogramming occurs during preimplantation embryo development. Here the authors develop a single cell multiomics sequencing technology that enables profiling of genome-wide chromatin accessibility, DNA methylation and RNA expression in the same individual cell and apply this method to study mouse preimplantation embryos.
- Yang Wang
- , Peng Yuan
- & Liying Yan
-
Article
| Open Access The genome-wide impact of trisomy 21 on DNA methylation and its implications for hematopoiesis
The genome-wide impact of trisomy 21 on DNA methylation and its implications for hematopoiesis
Down syndrome has a high co-morbidity with immune and hematopoietic disorders. Here, the authors perform an epigenome-wide association study in newborns with and without Down syndrome to find differential methylation across the genome, including in hematopoietic regulators RUNX1 and FLI1.
- Ivo S. Muskens
- , Shaobo Li
- & Adam J. de Smith
-
Article
| Open Access Comprehensive characterization of claudin-low breast tumors reflects the impact of the cell-of-origin on cancer evolution
Comprehensive characterization of claudin-low breast tumors reflects the impact of the cell-of-origin on cancer evolution
Claudin-low tumors are a rare aggressive subtype of breast cancers. In this study, the authors use a multiomics approach to demonstrate that these tumors are heterogeneous and comprise three main subgroups that emerge from different evolutionary processes.
- Roxane M. Pommier
- , Amélien Sanlaville
- & Alain Puisieux
-
Article
| Open Access Atlas of quantitative single-base-resolution N6-methyl-adenine methylomes
Atlas of quantitative single-base-resolution N6-methyl-adenine methylomes
N6-methyladenosine (m6A) and N6,2′-O-dimethyladenosine (m6Am) are eukaryotic mRNA modifications. Here the authors develop m6A-Crosslinking-Exonuclease-sequencing to map quantitative methylome changes at single-base-resolution after individually knocking out each known methyltransferase or demethylase.
- Casslynn W. Q. Koh
- , Yeek Teck Goh
- & W. S. Sho Goh
-
Article
| Open Access N6-methyladenosine mRNA marking promotes selective translation of regulons required for human erythropoiesis
N6-methyladenosine mRNA marking promotes selective translation of regulons required for human erythropoiesis
Erythropoiesis can be regulated by transcriptional, epigenetic, and post-transcriptional mechanisms. Here the authors report that N6-methyladenosine mRNA methyltransferase complex stimulates erythropoiesis by promoting translation of specific mRNAs.
- Daniel A. Kuppers
- , Sonali Arora
- & Patrick J. Paddison
-
Article
| Open Access Cell-type-specific resolution epigenetics without the need for cell sorting or single-cell biology
Cell-type-specific resolution epigenetics without the need for cell sorting or single-cell biology
Compared to bulk data, cell-type-specific DNA methylation data provide higher resolution of epigenetic variation. Here, the authors introduce Tensor Composition Analysis, a novel computational approach for learning cell-type-specific DNA methylation from tissue-level bulk data, and show its application in epigenome-wide association studies.
- Elior Rahmani
- , Regev Schweiger
- & Eran Halperin
-
Article
| Open Access Metaepigenomic analysis reveals the unexplored diversity of DNA methylation in an environmental prokaryotic community
Metaepigenomic analysis reveals the unexplored diversity of DNA methylation in an environmental prokaryotic community
Our knowledge of DNA methylation systems in prokaryotes is mostly limited to those of culturable microbes. Here, Hiraoka et al. analyse DNA methylation patterns in metagenomic data from a microbial community, revealing new methylated motifs and experimentally validating the methyltransferases’ specificities.
- Satoshi Hiraoka
- , Yusuke Okazaki
- & Wataru Iwasaki
-
Article
| Open Access Non-invasive detection of human cardiomyocyte death using methylation patterns of circulating DNA
Non-invasive detection of human cardiomyocyte death using methylation patterns of circulating DNA
The detection of cardiomyocyte death is a critical aspect in the diagnosis and monitoring of heart diseases. Here the authors show that cardiomyocyte-specific methylation patterns of circulating cell-free DNA may serve as a biomarker of cardiac cell death in infarcted and septic patients.
- Hai Zemmour
- , David Planer
- & Yuval Dor
-
Article
| Open Access scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells
scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells
Relationships between DNA methylation and transcription, and methylation and DNA accessibility can be probed but interrogating all three in the same single cells has not been possible. Here, the authors report the first single-cell method for parallel chromatin accessibility, DNA methylation and transcriptome profiling.
- Stephen J. Clark
- , Ricard Argelaguet
- & Wolf Reik

 Epigenome-wide association analysis of infant bronchiolitis severity: a multicenter prospective cohort study
Epigenome-wide association analysis of infant bronchiolitis severity: a multicenter prospective cohort study