Article
|
Open Access
Featured
-
-
Article
| Open Access Graph-based autoencoder integrates spatial transcriptomics with chromatin images and identifies joint biomarkers for Alzheimer’s disease
Graph-based autoencoder integrates spatial transcriptomics with chromatin images and identifies joint biomarkers for Alzheimer’s disease
Methods for jointly analysing the different spatial data modalities in 3D are lacking. Here the authors report the computational framework STACI (Spatial Transcriptomic data using over-parameterized graph-based Autoencoders with Chromatin Imaging data) which they apply to an Alzheimer’s disease mouse model.
- Xinyi Zhang
- , Xiao Wang
- & Caroline Uhler
-
Article
| Open Access Reference panel guided topological structure annotation of Hi-C data
Reference panel guided topological structure annotation of Hi-C data
Predicting topological structures from Hi-C data provides insight into comprehending gene expression and regulation. Here, the authors present RefHiC, an attention-based deep learning framework that leverages a reference panel of Hi-C datasets to assist topological structure annotation from a given study sample.
- Yanlin Zhang
- & Mathieu Blanchette
-
Article
| Open Access A unified computational framework for single-cell data integration with optimal transport
A unified computational framework for single-cell data integration with optimal transport
Integrating heterogeneous single-cell multi-omics as well as spatially resolved transcriptomic data remains a major challenge. Here the authors report a unified single-cell data integration framework using an unbalanced optimal transport-based deep network.
- Kai Cao
- , Qiyu Gong
- & Lin Wan
-
Article
| Open Access Analysis of the first genetic engineering attribution challenge
Analysis of the first genetic engineering attribution challenge
Identifying the designers of engineered biological sequences would help promote biotechnological innovation while holding designers accountable. Here the authors present the winners of a 2020 data-science competition which improved on previous attempts to attribute plasmid sequences.
- Oliver M. Crook
- , Kelsey Lane Warmbrod
- & William J. Bradshaw
-
Article
| Open Access DNA methylation-based classification of sinonasal tumors
DNA methylation-based classification of sinonasal tumors
Sinonasal tumour diagnosis can be complicated by the heterogeneity of disease and classification systems. Here, the authors use machine learning to classify sinonasal undifferentiated carcinomas into 4 molecular classe with differences in differentiation state and clinical outcome.
- Philipp Jurmeister
- , Stefanie Glöß
- & David Capper
-
Article
| Open Access A single-cell analysis reveals tumor heterogeneity and immune environment of acral melanoma
A single-cell analysis reveals tumor heterogeneity and immune environment of acral melanoma
Studying the cell composition of acral melanoma at the single-cell level could provide some clues about its poor response to immunotherapy. Here, the authors analyse acral and cutaneous melanoma patient samples using single-cell RNA-sequencing, and reveal a severe immunosuppressive state in acral melanomas
- Chao Zhang
- , Hongru Shen
- & Jilong Yang
-
Article
| Open Access DeepPROTACs is a deep learning-based targeted degradation predictor for PROTACs
DeepPROTACs is a deep learning-based targeted degradation predictor for PROTACs
The rational design of PROTACs is difficult due to their obscure structure-activity relationship. Here the authors present a deep neural network model - DeepPROTACs - for predicting the degradation capacity of a proposed PROTAC molecule.
- Fenglei Li
- , Qiaoyu Hu
- & Fang Bai
-
Article
| Open Access Leveraging data-driven self-consistency for high-fidelity gene expression recovery
Leveraging data-driven self-consistency for high-fidelity gene expression recovery
Recovering dropout-affected gene expression values is a challenging problem in bioinformatics. Here, the authors propose a data-driven framework, that first learns the underlying data distribution and then recovers the expression values by imposing a self-consistency on the expression matrix.
- Md Tauhidul Islam
- , Jen-Yeu Wang
- & Lei Xing
-
Article
| Open Access Deep learning to decompose macromolecules into independent Markovian domains
Deep learning to decompose macromolecules into independent Markovian domains
Modeling the dynamics of large proteins reveals a fundamental scaling problem. Here, the authors tackle this challenge by decomposing a large system into smaller independent subsystems, simultaneously modeling each subsystem’s kinetics and ensuring their mutual independence.
- Andreas Mardt
- , Tim Hempel
- & Frank Noé
-
Comment
| Open Access Developing medical imaging AI for emerging infectious diseases
Developing medical imaging AI for emerging infectious diseases
Very few of the COVID-19 ML models were fit for deployment in real-world settings. In this Comment, Huang et al. discuss the main steps required to develop clinically useful models in the context of an emerging infectious disease.
- Shih-Cheng Huang
- , Akshay S. Chaudhari
- & Matthew P. Lungren
-
Article
| Open Access Prediction of inter-chain distance maps of protein complexes with 2D attention-based deep neural networks
Prediction of inter-chain distance maps of protein complexes with 2D attention-based deep neural networks
Predicting inter-chain residue-residue distances of protein complexes is useful for constructing and evaluating quaternary structures of the protein complexes. Here, the authors develop a deep attention-based residual network method (CDPred) to predict inter-chain residue-residue distances of protein dimers.
- Zhiye Guo
- , Jian Liu
- & Jianlin Cheng
-
Article
| Open Access Causal deep learning reveals the comparative effectiveness of antihyperglycemic treatments in poorly controlled diabetes
Causal deep learning reveals the comparative effectiveness of antihyperglycemic treatments in poorly controlled diabetes
Current treatment guidelines for Type-2 diabetes endorse a massive number of potential anti-hyper-glycemic treatment options in various permutations and combinations. Here, the authors present a causal deep learning approach for more personalized recommendations of treatment selection.
- Chinmay Belthangady
- , Stefanos Giampanis
- & Beau Norgeot
-
Article
| Open Access A formal validation of a deep learning-based automated workflow for the interpretation of the echocardiogram
A formal validation of a deep learning-based automated workflow for the interpretation of the echocardiogram
Deep learning can automate the interpretation of medical imaging tests. Here, the authors formally assess the interchangeability of deep learning algorithms with expert human measurements for interpreting echocardiographic studies.
- Jasper Tromp
- , David Bauer
- & Scott D. Solomon
-
Article
| Open Access Systematic tissue annotations of genomics samples by modeling unstructured metadata
Systematic tissue annotations of genomics samples by modeling unstructured metadata
The 1+ million publicly-available human –omics samples currently remain acutely underused. Here the authors present an approach combining natural language processing and machine learning to infer the source tissue of public genomics samples based on their plain text descriptions, making these samples easy to discover and reuse.
- Nathaniel T. Hawkins
- , Marc Maldaver
- & Arjun Krishnan
-
Article
| Open Access Deep learning-based image analysis predicts PD-L1 status from H&E-stained histopathology images in breast cancer
Deep learning-based image analysis predicts PD-L1 status from H&E-stained histopathology images in breast cancer
Programmed death ligand-1 (PD-L1) has been recently adopted for breast cancer as a predictive biomarker for immunotherapies. Here, the authors show that PD-L1 expression can be predicted from H&E-stained images using deep learning.
- Gil Shamai
- , Amir Livne
- & Ron Kimmel
-
Article
| Open Access Learning the histone codes with large genomic windows and three-dimensional chromatin interactions using transformer
Learning the histone codes with large genomic windows and three-dimensional chromatin interactions using transformer
Existing deep learning-based approaches for the prediction of gene expression by histone modifications (HMs) can only focus on narrow and linear genomic regions around promoters. Here, the authors address these problems by developing a transformer-based deep learning architecture named Chromoformer.
- Dohoon Lee
- , Jeewon Yang
- & Sun Kim
-
Article
| Open Access Tempo: an unsupervised Bayesian algorithm for circadian phase inference in single-cell transcriptomics
Tempo: an unsupervised Bayesian algorithm for circadian phase inference in single-cell transcriptomics
Previous efforts to study the circadian clock using scRNA-seq have relied on time course designs that treat cell collection time as a proxy for circadian time. Here, the authors introduce a statistical method to infer circadian timing directly from expression, enabling researchers to study circadian phase heterogeneity.
- Benjamin J. Auerbach
- , Garret A. FitzGerald
- & Mingyao Li
-
Article
| Open Access Uncertainty-informed deep learning models enable high-confidence predictions for digital histopathology
Uncertainty-informed deep learning models enable high-confidence predictions for digital histopathology
Safe clinical deployment of deep learning models for digital pathology requires reliable estimates of predictive uncertainty. Here the authors describe an algorithm for quantifying whole-slide image uncertainty, demonstrating their approach with models trained to distinguish lung cancer subtypes.
- James M. Dolezal
- , Andrew Srisuwananukorn
- & Alexander T. Pearson
-
Article
| Open Access Deep learning empowered volume delineation of whole-body organs-at-risk for accelerated radiotherapy
Deep learning empowered volume delineation of whole-body organs-at-risk for accelerated radiotherapy
Volume delineation of organs-at risk (OARs) and target tumors is an indispensable process for creating radiotherapy treatment planning. Herein, the authors propose a lightweight deep learning framework to empower the rapid and precise volume delineation of whole-body OARs and target tumors.
- Feng Shi
- , Weigang Hu
- & Dinggang Shen
-
Article
| Open Access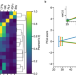 Unsupervised learning of aging principles from longitudinal data
Unsupervised learning of aging principles from longitudinal data
Biomarkers of age and frailty may aid in understanding the aging process, predicting lifespan or health span and in assessing the effects of anti-aging interventions. Here, the authors show that combining physics-based models and deep learning may enhance understanding of aging from big biomedical data, observe effects of anti-aging interventions in laboratory animals, and discover signatures of longevity.
- Konstantin Avchaciov
- , Marina P. Antoch
- & Peter O. Fedichev
-
Article
| Open Access De novo analysis of bulk RNA-seq data at spatially resolved single-cell resolution
De novo analysis of bulk RNA-seq data at spatially resolved single-cell resolution
Current methods to reanalyze bulk RNA-seq at spatially resolved single-cell resolution have limitations. Here, the authors develop Bulk2Space, a spatial deconvolution algorithm using single-cell and spatial transcriptomics as references, providing new insights into spatial heterogeneity within bulk tissue.
- Jie Liao
- , Jingyang Qian
- & Xiaohui Fan
-
Article
| Open Access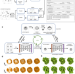 Isotropic reconstruction for electron tomography with deep learning
Isotropic reconstruction for electron tomography with deep learning
Cryogenic electron tomography suffers from anisotropic resolution due to the missing-wedge problem. Here, the authors present IsoNet, a neural network that learn the feature representation from similar structures in the tomogram and recover the missing information for isotropic tomogram reconstruction.
- Yun-Tao Liu
- , Heng Zhang
- & Z. Hong Zhou
-
Article
| Open Access Protein language models trained on multiple sequence alignments learn phylogenetic relationships
Protein language models trained on multiple sequence alignments learn phylogenetic relationships
Protein language models taking multiple sequence alignments as inputs capture protein structure and mutational effects. Here, the authors show that these models also encode phylogenetic relationships, and can disentangle correlations due to structural constraints from those due to phylogeny.
- Umberto Lupo
- , Damiano Sgarbossa
- & Anne-Florence Bitbol
-
Article
| Open Access Combining mass spectrometry and machine learning to discover bioactive peptides
Combining mass spectrometry and machine learning to discover bioactive peptides
Bioactive peptides regulate many physiological functions but progress in discovering them has been slow. Here, the authors use a machine learning framework to predict mammalian peptide candidates from the global and local structure of large-scale tissue-specific mass spectrometry data.
- Christian T. Madsen
- , Jan C. Refsgaard
- & Ulrik de Lichtenberg
-
Article
| Open Access Rapid protein assignments and structures from raw NMR spectra with the deep learning technique ARTINA
Rapid protein assignments and structures from raw NMR spectra with the deep learning technique ARTINA
The analysis of protein NMR spectra is time-consuming and can occupy a human expert for weeks or months. The researchers in this work present a deep learning-based method that delivers signal positions, chemical shift assignments, and structures of proteins within hours after completion of the NMR measurements.
- Piotr Klukowski
- , Roland Riek
- & Peter Güntert
-
Article
| Open Access Comprehensive and clinically accurate head and neck cancer organs-at-risk delineation on a multi-institutional study
Comprehensive and clinically accurate head and neck cancer organs-at-risk delineation on a multi-institutional study
Accurate organ at risk (OAR) segmentation is critical to reduce the radiotherapy post-treatment complications. Here, the authors develop an automated OAR segmentation system to delineate a comprehensive set of 42 H&N OARs.
- Xianghua Ye
- , Dazhou Guo
- & Tsung-Ying Ho
-
Article
| Open Access Using domain knowledge for robust and generalizable deep learning-based CT-free PET attenuation and scatter correction
Using domain knowledge for robust and generalizable deep learning-based CT-free PET attenuation and scatter correction
Deep learning-based methods have been proposed to substitute CT-based PET attenuation and scatter correction to achieve CT-free PET imaging. Here, the authors present a simple way to integrate domain knowledge in deep learning for CT-free PET imaging.
- Rui Guo
- , Song Xue
- & Kuangyu Shi
-
Article
| Open Access Adversarial attacks and adversarial robustness in computational pathology
Adversarial attacks and adversarial robustness in computational pathology
Artificial Intelligence can support diagnostic workflows in oncology, but they are vulnerable to adversarial attacks. Here, the authors show that convolutional neural networks are highly susceptible to white- and black-box adversarial attacks in clinically relevant classification tasks.
- Narmin Ghaffari Laleh
- , Daniel Truhn
- & Jakob Nikolas Kather
-
Article
| Open Access Deciphering microbial gene function using natural language processing
Deciphering microbial gene function using natural language processing
The function of many microbial genes is yet unknown. Here the authors repurposed natural language processing algorithms to explore “gene semantics” and infer function for thousands of genes with defense and secretion systems found to have the most discovery potential.
- Danielle Miller
- , Adi Stern
- & David Burstein
-
Article
| Open Access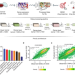 Gene expression based inference of cancer drug sensitivity
Gene expression based inference of cancer drug sensitivity
Predicting treatment response in cancer remains a highly complex task. Here, the authors develop Precily, a deep neural network framework to predict treatment response in cancer by considering gene expression, pathway activity estimates and drug features, and test this method in multiple datasets and preclinical models.
- Smriti Chawla
- , Anja Rockstroh
- & Debarka Sengupta
-
Article
| Open Access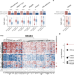 RAS oncogenic activity predicts response to chemotherapy and outcome in lung adenocarcinoma
RAS oncogenic activity predicts response to chemotherapy and outcome in lung adenocarcinoma
Mutations in RAS oncogenes and related pathways are frequent in lung cancers. Here, the authors derive a RAS gene expression signature and a machine learning classifier to predict drug response and clinical outcomes in lung adenocarcinoma and other solid tumours, with improved performance over KRAS mutations alone.
- Philip East
- , Gavin P. Kelly
- & Sophie de Carné Trécesson
-
Article
| Open Access Identification of spatially variable genes with graph cuts
Identification of spatially variable genes with graph cuts
Single-cell gene expression data with positional information is critical to dissect mechanisms and architectures of multicellular organisms, but the potential is limited by the scalability of current data analysis strategies. Here the authors develop a highly scalable method, scGCO, to identify genes whose expression values form spatial patterns from spatial transcriptomics data.
- Ke Zhang
- , Wanwan Feng
- & Peng Wang
-
Article
| Open Access Mutated processes predict immune checkpoint inhibitor therapy benefit in metastatic melanoma
Mutated processes predict immune checkpoint inhibitor therapy benefit in metastatic melanoma
Tumour mutational burden is a biomarker of immune checkpoint inhibitor response, but their association is not fully understood. Here, the authors train classifiers to identify key mutated processes which show stable predictive performance in multiple melanoma cohorts.
- Andrew Patterson
- & Noam Auslander
-
Matters Arising
| Open AccessMachine-learning prediction of hosts of novel coronaviruses requires caution as it may affect wildlife conservation
- Sophie Lund Rasmussen
- , Cino Pertoldi
- & David W. Macdonald
-
Matters Arising
| Open AccessReply to: Machine-learning prediction of hosts of novel coronaviruses requires caution as it may affect wildlife conservation
- Marcus S. C. Blagrove
- , Matthew Baylis
- & Maya Wardeh
-
Article
| Open Access Enhanced detection of threat materials by dark-field x-ray imaging combined with deep neural networks
Enhanced detection of threat materials by dark-field x-ray imaging combined with deep neural networks
Dark-field X-ray imaging is sensitive to the microstructure of a material. Here, the authors combine this with a neural network algorithm to provide efficient material discrimination, e.g., of explosives vs non-threat materials.
- T. Partridge
- , A. Astolfo
- & A. Olivo
-
Article
| Open Access Integrating and formatting biomedical data as pre-calculated knowledge graph embeddings in the Bioteque
Integrating and formatting biomedical data as pre-calculated knowledge graph embeddings in the Bioteque
Biomedical data is accumulating at a fast pace and integrating it into a unified framework is a major challenge. Here, the authors present a resource that contains pre-calculated biomedical descriptors derived from a very large knowledge graph.
- Adrià Fernández-Torras
- , Miquel Duran-Frigola
- & Patrick Aloy
-
Article
| Open Access Traject3d allows label-free identification of distinct co-occurring phenotypes within 3D culture by live imaging
Traject3d allows label-free identification of distinct co-occurring phenotypes within 3D culture by live imaging
There are currently a lack of tools to detect heterogeneity in 3D cultures. Here the authors report Traject3d as a framework to identify heterogeneous states in 3D culture and to understand how these give rise to distinct phenotypes using label-free multi-day time-lapse imaging.
- Eva C. Freckmann
- , Emma Sandilands
- & David M. Bryant
-
Article
| Open Access devCellPy is a machine learning-enabled pipeline for automated annotation of complex multilayered single-cell transcriptomic data
devCellPy is a machine learning-enabled pipeline for automated annotation of complex multilayered single-cell transcriptomic data
A major informatic challenge in single cell RNA-sequencing analysis is the precise annotation of datasets where cells exhibit complex multilayered identities or transitory states. Here the authors present devCellPy, a Python-based package that enables the automated prediction of cell types across complex cellular hierarchies, species, and experimental systems with high accuracy, particularly for developmental scRNA-seq datasets.
- Francisco X. Galdos
- , Sidra Xu
- & Sean M. Wu
-
Article
| Open Access Accounting for small variations in the tracrRNA sequence improves sgRNA activity predictions for CRISPR screening
Accounting for small variations in the tracrRNA sequence improves sgRNA activity predictions for CRISPR screening
Existing methods for generating sgRNA predictions do not account for the tracrRNA sequence. Here the authors report an on-target model, Rule Set 3, to generate optimal predictions for multiple tracrRNA variants, and validate this on a new dataset of sgRNAs showing improvement over prior prediction models.
- Peter C. DeWeirdt
- , Abby V. McGee
- & John G. Doench
-
Article
| Open Access Automated model-predictive design of synthetic promoters to control transcriptional profiles in bacteria
Automated model-predictive design of synthetic promoters to control transcriptional profiles in bacteria
Transcription rates are regulated by the interactions between RNA polymerase, sigma factor, and promoter DNA sequences in bacteria. Here the authors combine massively parallel experiments & machine learning to develop a predictive biophysical model of transcription, validated across 22132 bacterial promoters, and apply it to the design and debugging of genetic circuits.
- Travis L. LaFleur
- , Ayaan Hossain
- & Howard M. Salis
-
Article
| Open Access Controlling gene expression with deep generative design of regulatory DNA
Controlling gene expression with deep generative design of regulatory DNA
Design of de novo synthetic regulatory DNA is a promising avenue to control gene expression in biotechnology and medicine. Here the authors present EspressionGAN, a generative adversarial network that uses genomic and transcriptomic data to generate regulatory sequences.
- Jan Zrimec
- , Xiaozhi Fu
- & Aleksej Zelezniak
-
Article
| Open Access Deep learning image segmentation reveals patterns of UV reflectance evolution in passerine birds
Deep learning image segmentation reveals patterns of UV reflectance evolution in passerine birds
Here, the authors develop software that uses photographs of birds to extract information on plumage UV reflectance. They use these data to show that UV reflectance is phylogenetically conserved and associated with the light environment.
- Yichen He
- , Zoë K. Varley
- & Christopher R. Cooney
-
Article
| Open Access Genome-wide mutational signatures in low-coverage whole genome sequencing of cell-free DNA
Genome-wide mutational signatures in low-coverage whole genome sequencing of cell-free DNA
Detection of mutational signatures in cell-free DNA (cfDNA) is challenging due to low sequence coverage and low mutant allele fractions. Here, the authors identify mutational signatures in plasma whole genome sequencing of cancer patients and use machine learning to distinguish them from healthy individuals.
- Jonathan C. M. Wan
- , Dennis Stephens
- & Luis A. Diaz Jr.
-
Comment
| Open Access Addressing fairness in artificial intelligence for medical imaging
Addressing fairness in artificial intelligence for medical imaging
A plethora of work has shown that AI systems can systematically and unfairly be biased against certain populations in multiple scenarios. The field of medical imaging, where AI systems are beginning to be increasingly adopted, is no exception. Here we discuss the meaning of fairness in this area and comment on the potential sources of biases, as well as the strategies available to mitigate them. Finally, we analyze the current state of the field, identifying strengths and highlighting areas of vacancy, challenges and opportunities that lie ahead.
- María Agustina Ricci Lara
- , Rodrigo Echeveste
- & Enzo Ferrante
-
Article
| Open Access Multi-cohort and longitudinal Bayesian clustering study of stage and subtype in Alzheimer’s disease
Multi-cohort and longitudinal Bayesian clustering study of stage and subtype in Alzheimer’s disease
Different types of atrophy in Alzheimer’s disease may reflect different disease stages or biologically distinct subtypes. Here the authors use longitudinal neuroimaging data to demonstrate five distinct patterns of atrophy with different demographical and cognitive characteristics.
- Konstantinos Poulakis
- , Joana B. Pereira
- & Eric Westman
-
Article
| Open Access ProtGPT2 is a deep unsupervised language model for protein design
ProtGPT2 is a deep unsupervised language model for protein design
Protein design aims to build novel proteins customized for specific purposes, thereby holding the potential to tackle many environmental and biomedical problems. Here the authors apply some of the latest advances in natural language processing, generative Transformers, to train ProtGPT2, a language model that explores unseen regions of the protein space while designing proteins with nature-like properties.
- Noelia Ferruz
- , Steffen Schmidt
- & Birte Höcker
-
Article
| Open Access Decoding kinase-adverse event associations for small molecule kinase inhibitors
Decoding kinase-adverse event associations for small molecule kinase inhibitors
Small molecule kinase inhibitors (SMKIs) are being approved at a fast pace under expedited programs for anticancer treatment. Here, the authors employ a machine-learning model to examine the relationships between kinase targets and adverse events in the trials of 16 FDA-approved SMKIs.
- Xiajing Gong
- , Meng Hu
- & Liang Zhao
-
Article
| Open Access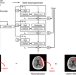 Emergency triage of brain computed tomography via anomaly detection with a deep generative model
Emergency triage of brain computed tomography via anomaly detection with a deep generative model
Triage is essential for the early diagnosis and reporting of emergency patients in the emergency department. Here, the authors develop an anomaly detection algorithm with a deep generative model that reprioritizes radiology worklists and provides lesion attention maps for brain CT images with critical findings.
- Seungjun Lee
- , Boryeong Jeong
- & Namkug Kim

 A framework for clinical cancer subtyping from nucleosome profiling of cell-free DNA
A framework for clinical cancer subtyping from nucleosome profiling of cell-free DNA