Article
|
Open Access
Featured
-
-
Article
| Open Access Uncovering interpretable potential confounders in electronic medical records
Uncovering interpretable potential confounders in electronic medical records
Randomized clinical trials are often plagued by selection bias, and expert-selected covariates may insufficiently adjust for confounding factors. Here, the authors develop a framework based on natural language processing to uncover interpretable potential confounders from text.
- Jiaming Zeng
- , Michael F. Gensheimer
- & Ross D. Shachter
-
Article
| Open Access PyUUL provides an interface between biological structures and deep learning algorithms
PyUUL provides an interface between biological structures and deep learning algorithms
While artificial intelligence (AI) is quickly becoming ubiquitous, biology still suffers from the lack of interfaces connecting biological structures and modern AI methods. Here, the authors report PyUUL, a library to translate biological structures into 3D differentiable tensorial representations.
- Gabriele Orlando
- , Daniele Raimondi
- & Frederic Rousseau
-
Article
| Open Access A deep-learning system bridging molecule structure and biomedical text with comprehension comparable to human professionals
A deep-learning system bridging molecule structure and biomedical text with comprehension comparable to human professionals
To accelerate biomedical research process, deep-learning systems are developed to automatically acquire knowledge about molecule entities by reading large-scale biomedical data. Inspired by humans that learn deep molecule knowledge from both molecule structure and biomedical text information, the authors propose a machine reading system that bridges both types of information.
- Zheni Zeng
- , Yuan Yao
- & Maosong Sun
-
Article
| Open Access UINMF performs mosaic integration of single-cell multi-omic datasets using nonnegative matrix factorization
UINMF performs mosaic integration of single-cell multi-omic datasets using nonnegative matrix factorization
Single-cell genomic technologies present unique data integration challenges. Here the authors introduce an integrative nonnegative matrix factorization algorithm that incorporates features unshared between datasets when performing dataset integrations, improving integration results for spatial transcriptomic, cross-modality, and cross-species data.
- April R. Kriebel
- & Joshua D. Welch
-
Article
| Open Access Protein sequence design with a learned potential
Protein sequence design with a learned potential
Rational protein design to achieve a given protein backbone conformation is needed to engineer specific functions. Here Anand et al. describe a machine learning method using a learned neural network potential for fixed-backbone protein design.
- Namrata Anand
- , Raphael Eguchi
- & Po-Ssu Huang
-
Article
| Open Access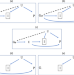 A robust method for collider bias correction in conditional genome-wide association studies
A robust method for collider bias correction in conditional genome-wide association studies
Genetic associations can be biased by conditioning on a phenotype. This study presents ‘Slope-Hunter’, a method which uses model-based clustering to correct this bias, even in the presence of genetic correlation, assuming the class of SNPs affecting only the collider explains more variation in the collider than any other class of SNPs.
- Osama Mahmoud
- , Frank Dudbridge
- & Kate Tilling
-
Article
| Open Access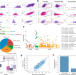 A pan-CRISPR analysis of mammalian cell specificity identifies ultra-compact sgRNA subsets for genome-scale experiments
A pan-CRISPR analysis of mammalian cell specificity identifies ultra-compact sgRNA subsets for genome-scale experiments
Context specificity confounds genetic analysis and prevents reproducible genome engineering. Here, the authors report a pan-CRISPR analysis of specificity in mammalian cells and identify ultra-compact sgRNA subsets for genome-scale screens.
- Boyang Zhao
- , Yiyun Rao
- & Justin R. Pritchard
-
Article
| Open Access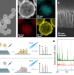 Multiplexed nanomaterial-assisted laser desorption/ionization for pan-cancer diagnosis and classification
Multiplexed nanomaterial-assisted laser desorption/ionization for pan-cancer diagnosis and classification
As cancer is increasingly considered a metabolic disorder, it is postulated that serum metabolite profiling can be a viable approach for detecting the presence of cancer. Here, the authors report a machine learning model using mass spectrometry-based liquid biopsy data for pan-cancer screening and classification.
- Hua Zhang
- , Lin Zhao
- & Xiangfeng Duan
-
Article
| Open Access Automatic mapping of multiplexed social receptive fields by deep learning and GPU-accelerated 3D videography
Automatic mapping of multiplexed social receptive fields by deep learning and GPU-accelerated 3D videography
High resolution descriptions of social interactions and their neural correlates are lacking. Here the authors report a pipeline enabling fully automatic multi-animal tracking during social encounters, together with simultaneous electrophysiological recordings, and show this works in low-light settings.
- Christian L. Ebbesen
- & Robert C. Froemke
-
Article
| Open Access Machine learning-informed and synthetic biology-enabled semi-continuous algal cultivation to unleash renewable fuel productivity
Machine learning-informed and synthetic biology-enabled semi-continuous algal cultivation to unleash renewable fuel productivity
Growth limitation caused by mutual shading and the high harvest cost hamper algal biofuel production. Here, the authors overcome these two problems by designing a semi-continuous algal cultivation system and an aggregation-based sedimentation strategy to achieve high levels production of biomass and limonene.
- Bin Long
- , Bart Fischer
- & Joshua S. Yuan
-
Article
| Open Access Algorithm-aided engineering of aliphatic halogenase WelO5* for the asymmetric late-stage functionalization of soraphens
Algorithm-aided engineering of aliphatic halogenase WelO5* for the asymmetric late-stage functionalization of soraphens
The late-stage functionalization of unactivated carbon–hydrogen bonds is a difficult but important task, which has been met with promising but limited success through synthetic organic chemistry. Here the authors use machine learning to engineer WelO5* halogenase variants, which led to regioselective chlorination of inert C–H bonds on a representative polyketide that is a non-natural substrate for the enzyme.
- Johannes Büchler
- , Sumire Honda Malca
- & Rebecca Buller
-
Article
| Open Access Harnessing protein folding neural networks for peptide–protein docking
Harnessing protein folding neural networks for peptide–protein docking
AlphaFold2 has originally been developed to provide highly accurate predictions of protein monomer structures. Here, the authors present a simple adaptation of AlphaFold2 that enables structural modeling of peptide–protein complexes, and explore the underlying mechanisms and limitations of this approach.
- Tomer Tsaban
- , Julia K. Varga
- & Ora Schueler-Furman
-
Article
| Open Access Mini-batch optimization enables training of ODE models on large-scale datasets
Mini-batch optimization enables training of ODE models on large-scale datasets
Ordinary differential equation (ODE) models are widely used to understand multiple processes. Here the authors show how the concept of mini-batch optimization can be transferred from the field of Deep Learning to ODE modelling.
- Paul Stapor
- , Leonard Schmiester
- & Jan Hasenauer
-
Article
| Open Access Topographic mapping of the glioblastoma proteome reveals a triple-axis model of intra-tumoral heterogeneity
Topographic mapping of the glioblastoma proteome reveals a triple-axis model of intra-tumoral heterogeneity
Gioblastoma tumours consist of different niches defined by histology. Here, the authors use proteomics and machine learning to assign protein expression programs to these niches, and reveal that KRAS and hypoxia are associated with drug resistance.
- K. H. Brian Lam
- , Alberto J. Leon
- & Phedias Diamandis
-
Article
| Open Access A machine and human reader study on AI diagnosis model safety under attacks of adversarial images
A machine and human reader study on AI diagnosis model safety under attacks of adversarial images
While active efforts are advancing medical AI model development and clinical translation, safety issues of medical AI models have emerged. Here, the authors investigate the effects on an AI model and on human experts of potential fake/adversarial images for breast cancer diagnosis.
- Qianwei Zhou
- , Margarita Zuley
- & Shandong Wu
-
Article
| Open Access DeepRank: a deep learning framework for data mining 3D protein-protein interfaces
DeepRank: a deep learning framework for data mining 3D protein-protein interfaces
The authors present DeepRank, a deep learning framework for the data mining of large sets of 3D protein-protein interfaces (PPI). They use DeepRank to address two challenges in structural biology: distinguishing biological versus crystallographic PPIs in crystal structures, and secondly the ranking of docking models.
- Nicolas Renaud
- , Cunliang Geng
- & Li C. Xue
-
Article
| Open Access A unified drug–target interaction prediction framework based on knowledge graph and recommendation system
A unified drug–target interaction prediction framework based on knowledge graph and recommendation system
Prediction of drug-target interactions (DTI) plays a vital role in drug development through applications in various areas, such as virtual screening for lead discovery, drug repurposing and identification of potential drug side effects. Here, the authors develop a unified framework for DTI prediction by combining a knowledge graph and a recommendation system.
- Qing Ye
- , Chang-Yu Hsieh
- & Tingjun Hou
-
Article
| Open Access DeepPhospho accelerates DIA phosphoproteome profiling through in silico library generation
DeepPhospho accelerates DIA phosphoproteome profiling through in silico library generation
The coverage and throughput of data-independent acquisition (DIA)-based phosphoproteomics is limited by its dependence on experimental spectral libraries. Here the authors develop a DIA workflow based on in silico spectral libraries generated by a novel deep neural network to expand phosphoproteome coverage.
- Ronghui Lou
- , Weizhen Liu
- & Wenqing Shui
-
Article
| Open Access Machine learning of genomic features in organotropic metastases stratifies progression risk of primary tumors
Machine learning of genomic features in organotropic metastases stratifies progression risk of primary tumors
The location and timing of metastasis are still fundamentally unpredictable. Here the authors present the Metastatic Network model, a machine learning framework that integrates clinical data and DNA alterations to predict the risk of metastasis to specific organs as well as clinical outcomes
- Biaobin Jiang
- , Quanhua Mu
- & Jiguang Wang
-
Article
| Open Access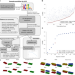 Local DNA shape is a general principle of transcription factor binding specificity in Arabidopsis thaliana
Local DNA shape is a general principle of transcription factor binding specificity in Arabidopsis thaliana
Methods to predict transcription factor binding sites typically focus on sequence motifs without considering DNA shape. Here the authors use a random forest machine learning approach that incorporates DNA shape and improves binding site prediction for Arabidopsis thaliana transcription factors.
- Janik Sielemann
- , Donat Wulf
- & Andrea Bräutigam
-
Article
| Open Access Co-evolution based machine-learning for predicting functional interactions between human genes
Co-evolution based machine-learning for predicting functional interactions between human genes
With the rise in number of eukaryotic species being fully sequenced, large scale phylogenetic profiling can give insights on gene function, Here, the authors describe a machine-learning approach that integrates co-evolution across eukaryotic clades to predict gene function and functional interactions among human genes.
- Doron Stupp
- , Elad Sharon
- & Yuval Tabach
-
Article
| Open Access Physics-informed deep learning characterizes morphodynamics of Asian soybean rust disease
Physics-informed deep learning characterizes morphodynamics of Asian soybean rust disease
Deep learning (DL) can be used to automatically extract complex features from dynamic systems. Here, the authors combine high-content imaging, DL and mechanistic models to extract and explain drug-induced morphological changes in the growth of the fungus responsible for Asian soybean rust.
- Henry Cavanagh
- , Andreas Mosbach
- & Robert G. Endres
-
Article
| Open Access Massively parallel interrogation of protein fragment secretability using SECRiFY reveals features influencing secretory system transit
Massively parallel interrogation of protein fragment secretability using SECRiFY reveals features influencing secretory system transit
The exact protein features that control passage through the eukaryotic secretory system remain largely unknown. Here the authors report SECRiFY which they use to evaluate the secretory potential of polypeptides on a proteome-wide scale in yeast, revealing a role for flexibility and intrinsic disorder.
- Morgane Boone
- , Pathmanaban Ramasamy
- & Nico Callewaert
-
Article
| Open Access Chromatin-accessibility estimation from single-cell ATAC-seq data with scOpen
Chromatin-accessibility estimation from single-cell ATAC-seq data with scOpen
scATAC-Seq yields data that is extremely sparse. Here, the authors present a computationally efficient imputation method called scOpen that improves the downstream analyses of scATAC-Seq data and use it to identify transcriptional regulators of kidney fibrosis.
- Zhijian Li
- , Christoph Kuppe
- & Ivan G. Costa
-
Article
| Open Access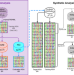 The generative capacity of probabilistic protein sequence models
The generative capacity of probabilistic protein sequence models
Generative models have become increasingly popular in protein design, yet rigorous metrics that allow the comparison of these models are lacking. Here, the authors propose a set of such metrics and use them to compare three popular models.
- Francisco McGee
- , Sandro Hauri
- & Allan Haldane
-
Article
| Open Access Accurate recognition of colorectal cancer with semi-supervised deep learning on pathological images
Accurate recognition of colorectal cancer with semi-supervised deep learning on pathological images
Machine-assisted recognition of colorectal cancer has been mainly focused on supervised deep learning that suffers from a significant bottleneck of requiring massive amounts of labeled data. Here, the authors propose a semi-supervised model based on the mean teacher architecture that provides pathological predictions at both patch- and patient-levels.
- Gang Yu
- , Kai Sun
- & Hong-Wen Deng
-
Article
| Open Access Precise measurements of chromatin diffusion dynamics by modeling using Gaussian processes
Precise measurements of chromatin diffusion dynamics by modeling using Gaussian processes
Although much effort has been devoted to determine the 3D structure of chromatin, there is a need for new experimental and computational methods. Here the authors present GP-FBM to extract chromatin diffusion parameters with high precision and apply it to live-imaging of embryonic stem cells, revealing that the diffusive properties of mitotic and interphase chromatin do not differ significantly.
- Guilherme M. Oliveira
- , Attila Oravecz
- & Nacho Molina
-
Article
| Open Access Bayesian log-normal deconvolution for enhanced in silico microdissection of bulk gene expression data
Bayesian log-normal deconvolution for enhanced in silico microdissection of bulk gene expression data
Deconvolution methods reveal individual cell types in complex tissues profiled by bulk methods. Here the authors present a Bayesian deconvolution method that outperforms existing methods when benchmarked on >700 datasets, especially in estimating cell-type-specific gene expression profiles.
- Bárbara Andrade Barbosa
- , Saskia D. van Asten
- & Yongsoo Kim
-
Article
| Open Access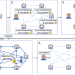 Truly privacy-preserving federated analytics for precision medicine with multiparty homomorphic encryption
Truly privacy-preserving federated analytics for precision medicine with multiparty homomorphic encryption
Existing approaches to sharing of distributed medical data either provide only limited protection of patients’ privacy or sacrifice the accuracy of results. Here, the authors propose a federated analytics system, based on multiparty homomorphic encryption (MHE), to overcome these issues.
- David Froelicher
- , Juan R. Troncoso-Pastoriza
- & Jean-Pierre Hubaux
-
Article
| Open Access Annotation-efficient deep learning for automatic medical image segmentation
Annotation-efficient deep learning for automatic medical image segmentation
Existing high-performance deep learning methods typically rely on large training datasets with high-quality manual annotations, which are difficult to obtain in many clinical applications. Here, the authors introduce an open-source framework to handle imperfect training datasets.
- Shanshan Wang
- , Cheng Li
- & Hairong Zheng
-
Article
| Open Access DUBStepR is a scalable correlation-based feature selection method for accurately clustering single-cell data
DUBStepR is a scalable correlation-based feature selection method for accurately clustering single-cell data
Cell-type-specific genes are often strongly correlated in expression - an informative yet underexplored property of single-cell data. Here, the authors leverage gene expression correlations to develop DUBStepR, a feature selection method for accurately clustering single-cell data.
- Bobby Ranjan
- , Wenjie Sun
- & Shyam Prabhakar
-
Article
| Open Access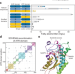 Machine learning-guided acyl-ACP reductase engineering for improved in vivo fatty alcohol production
Machine learning-guided acyl-ACP reductase engineering for improved in vivo fatty alcohol production
Fatty acyl reductases (FARs) are critical enzymes in the biosynthesis of fatty alcohols and have the ability to directly acces acyl-ACP substrates. Here, authors couple machine learning-based protein engineering framework with gene shuffling to optimize a FAR for the activity on acyl-ACP and improve fatty alcohol production.
- Jonathan C. Greenhalgh
- , Sarah A. Fahlberg
- & Philip A. Romero
-
Article
| Open Access Efficient generative modeling of protein sequences using simple autoregressive models
Efficient generative modeling of protein sequences using simple autoregressive models
Deep learning is a powerful tool for the design of novel protein sequences, yet can be computationally very inefficient. Here the authors propose using simple forecasting models to efficiently generate a large number of novel protein structures.
- Jeanne Trinquier
- , Guido Uguzzoni
- & Martin Weigt
-
Article
| Open Access ECNet is an evolutionary context-integrated deep learning framework for protein engineering
ECNet is an evolutionary context-integrated deep learning framework for protein engineering
Protein engineering is an active area of research in which machine learning has proven quite powerful. Here, the authors present a deep learning method that integrates both general and protein-specific sequence representations to improve the engineering of one’s protein of interest.
- Yunan Luo
- , Guangde Jiang
- & Jian Peng
-
Article
| Open Access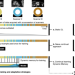 Dynamic memory to alleviate catastrophic forgetting in continual learning with medical imaging
Dynamic memory to alleviate catastrophic forgetting in continual learning with medical imaging
In clinical practice, the continuous progress of image acquisition technology or diagnostic procedures and evolving imaging protocols hamper the utility of machine learning, as prediction accuracy on new data deteriorates. Here, the authors propose a continual learning approach to deal with such domain shifts occurring at unknown time points.
- Matthias Perkonigg
- , Johannes Hofmanninger
- & Georg Langs
-
Article
| Open Access VEGA is an interpretable generative model for inferring biological network activity in single-cell transcriptomics
VEGA is an interpretable generative model for inferring biological network activity in single-cell transcriptomics
Developing interpretable models is a major challenge in single cell deep learning. Here we show that the VEGA variational autoencoder model, whose decoder wiring mirrors gene modules, can provide direct interpretability to the latent space further enabling the inference of biological module activity.
- Lucas Seninge
- , Ioannis Anastopoulos
- & Joshua Stuart
-
Article
| Open Access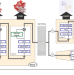 Mapping the glycosyltransferase fold landscape using interpretable deep learning
Mapping the glycosyltransferase fold landscape using interpretable deep learning
Glycosyltransferases (GT) are proteins that display extensive sequence and functional variation on a subset of 3D folds. Here, the authors use interpretable deep learning to predict 3D folds from sequence without the need for sequence alignment, which also enables the prediction of GTs with new folds.
- Rahil Taujale
- , Zhongliang Zhou
- & Natarajan Kannan
-
Article
| Open Access Robust whole slide image analysis for cervical cancer screening using deep learning
Robust whole slide image analysis for cervical cancer screening using deep learning
Computer-assisted diagnosis is key for scaling up cervical cancer screening, but current algorithms perform poorly on whole slide image analysis and generalization. Here, the authors present a WSI classification and top lesion cell recommendation system using deep learning, and achieve comparable results with cytologists.
- Shenghua Cheng
- , Sibo Liu
- & Xiuli Liu
-
Article
| Open Access Evolutionarily informed machine learning enhances the power of predictive gene-to-phenotype relationships
Evolutionarily informed machine learning enhances the power of predictive gene-to-phenotype relationships
Predicting complex phenotypes from genomic information is still a challenge. Here, the authors use an evolutionarily informed machine learning approach within and across species to predict genes affecting nitrogen utilization in crops, and show their approach is also useful in mammalian systems.
- Chia-Yi Cheng
- , Ying Li
- & Gloria M. Coruzzi
-
Article
| Open Access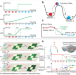 Dissecting transition cells from single-cell transcriptome data through multiscale stochastic dynamics
Dissecting transition cells from single-cell transcriptome data through multiscale stochastic dynamics
How to infer transient cells and cell-fate transitions from snap-shot single cell transcriptome dataset remains a major challenge. Here the authors present a multiscale approach to construct single-cell dynamical manifold, quantify cell stability, and compute transition trajectory and probability between cell states.
- Peijie Zhou
- , Shuxiong Wang
- & Qing Nie
-
Article
| Open Access Leveraging the Cell Ontology to classify unseen cell types
Leveraging the Cell Ontology to classify unseen cell types
Classifying cells into unseen cell types remains challenging in scRNA-seq analysis. Here we show that Cell Ontology enables an accurate classification of unseen cell types through considering the cell type relationships in the Cell Ontology graph.
- Sheng Wang
- , Angela Oliveira Pisco
- & Russ B. Altman
-
Article
| Open Access Peak learning of mass spectrometry imaging data using artificial neural networks
Peak learning of mass spectrometry imaging data using artificial neural networks
The high dimensional and complex nature of mass spectrometry imaging (MSI) data poses challenges to downstream analyses. Here the authors show an application of artificial intelligence in mining MSI data revealing biologically relevant metabolomic and proteomic information from data acquired on different mass spectrometry platforms.
- Walid M. Abdelmoula
- , Begona Gimenez-Cassina Lopez
- & Nathalie Y. R. Agar
-
Article
| Open Access Cross-species behavior analysis with attention-based domain-adversarial deep neural networks
Cross-species behavior analysis with attention-based domain-adversarial deep neural networks
Comparing changes in behaviour across various species is not always trivial, especial across significantly divergent species. Here, the authors develop a deep learning framework that allows them to map changes in locomotion demonstrated on dopamine-deficient humans, mice and worms.
- Takuya Maekawa
- , Daiki Higashide
- & Susumu Takahashi
-
Article
| Open Access Automated bone mineral density prediction and fracture risk assessment using plain radiographs via deep learning
Automated bone mineral density prediction and fracture risk assessment using plain radiographs via deep learning
Dual-energy X-ray absorptiometry and the Fracture Risk Assessment Tool are recommended tools for osteoporotic fracture risk evaluation, but are underutilized. Here, the authors present an opportunistic tool to identify fractures, predict bone mineral density and evaluate fracture risk using plain pelvis and lumbar spine radiographs.
- Chen-I Hsieh
- , Kang Zheng
- & Chang-Fu Kuo
-
Article
| Open Access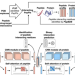 A deep-learning framework for multi-level peptide–protein interaction prediction
A deep-learning framework for multi-level peptide–protein interaction prediction
Peptide-protein interactions play fundamental roles in cellular processes and are crucial for designing peptide therapeutics. Here, the authors present a deep learning framework for simultaneously predicting peptide-protein interactions and identifying peptide binding residues involved in the interactions.
- Yipin Lei
- , Shuya Li
- & Jianyang Zeng
-
Article
| Open Access Unified AI framework to uncover deep interrelationships between gene expression and Alzheimer’s disease neuropathologies
Unified AI framework to uncover deep interrelationships between gene expression and Alzheimer’s disease neuropathologies
The molecular basis of Alzheimer’s Disease has been obscured by heterogeneity and scarcity of brain gene expression data, which limit effectiveness in complex models. Here, the authors introduce a multi-task deep learning framework to learn generalizable and nuanced relationships between gene expression and neuropathology.
- Nicasia Beebe-Wang
- , Safiye Celik
- & Su-In Lee
-
Article
| Open Access CloneSig can jointly infer intra-tumor heterogeneity and mutational signature activity in bulk tumor sequencing data
CloneSig can jointly infer intra-tumor heterogeneity and mutational signature activity in bulk tumor sequencing data
Intratumour heterogeneity (ITH) and mutational signatures are typically analysed separately, even though they are not necessarily independent. Here, the authors present CloneSig, a tool for the joint estimation of ITH and mutational signatures, with which they analyse the TCGA and PCAWG datasets.
- Judith Abécassis
- , Fabien Reyal
- & Jean-Philippe Vert
-
Article
| Open Access Automatically disambiguating medical acronyms with ontology-aware deep learning
Automatically disambiguating medical acronyms with ontology-aware deep learning
Disambiguating abbreviations is important for automated clinical note processing; however, deploying machine learning for this task is restricted by lack of good training data. Here, the authors show novel data augmentation methods that use biomedical ontologies to improve abbreviation disambiguation in many datasets.
- Marta Skreta
- , Aryan Arbabi
- & Michael Brudno
-
Article
| Open Access Learning interpretable cellular and gene signature embeddings from single-cell transcriptomic data
Learning interpretable cellular and gene signature embeddings from single-cell transcriptomic data
Computational single-cell RNA-seq analyses often face challenges in scalability, model interpretability, and confounders. Here, we show a new model to address these challenges by learning meaningful embeddings from the data that simultaneously refine gene signatures and cell functions in diverse conditions.
- Yifan Zhao
- , Huiyu Cai
- & Yue Li

 Active label cleaning for improved dataset quality under resource constraints
Active label cleaning for improved dataset quality under resource constraints