Article
|
Open Access
Featured
-
-
Article
| Open Access Deep learning image segmentation reveals patterns of UV reflectance evolution in passerine birds
Deep learning image segmentation reveals patterns of UV reflectance evolution in passerine birds
Here, the authors develop software that uses photographs of birds to extract information on plumage UV reflectance. They use these data to show that UV reflectance is phylogenetically conserved and associated with the light environment.
- Yichen He
- , Zoë K. Varley
- & Christopher R. Cooney
-
Article
| Open Access Genome-wide mutational signatures in low-coverage whole genome sequencing of cell-free DNA
Genome-wide mutational signatures in low-coverage whole genome sequencing of cell-free DNA
Detection of mutational signatures in cell-free DNA (cfDNA) is challenging due to low sequence coverage and low mutant allele fractions. Here, the authors identify mutational signatures in plasma whole genome sequencing of cancer patients and use machine learning to distinguish them from healthy individuals.
- Jonathan C. M. Wan
- , Dennis Stephens
- & Luis A. Diaz Jr.
-
Comment
| Open Access Addressing fairness in artificial intelligence for medical imaging
Addressing fairness in artificial intelligence for medical imaging
A plethora of work has shown that AI systems can systematically and unfairly be biased against certain populations in multiple scenarios. The field of medical imaging, where AI systems are beginning to be increasingly adopted, is no exception. Here we discuss the meaning of fairness in this area and comment on the potential sources of biases, as well as the strategies available to mitigate them. Finally, we analyze the current state of the field, identifying strengths and highlighting areas of vacancy, challenges and opportunities that lie ahead.
- María Agustina Ricci Lara
- , Rodrigo Echeveste
- & Enzo Ferrante
-
Article
| Open Access Multi-cohort and longitudinal Bayesian clustering study of stage and subtype in Alzheimer’s disease
Multi-cohort and longitudinal Bayesian clustering study of stage and subtype in Alzheimer’s disease
Different types of atrophy in Alzheimer’s disease may reflect different disease stages or biologically distinct subtypes. Here the authors use longitudinal neuroimaging data to demonstrate five distinct patterns of atrophy with different demographical and cognitive characteristics.
- Konstantinos Poulakis
- , Joana B. Pereira
- & Eric Westman
-
Article
| Open Access ProtGPT2 is a deep unsupervised language model for protein design
ProtGPT2 is a deep unsupervised language model for protein design
Protein design aims to build novel proteins customized for specific purposes, thereby holding the potential to tackle many environmental and biomedical problems. Here the authors apply some of the latest advances in natural language processing, generative Transformers, to train ProtGPT2, a language model that explores unseen regions of the protein space while designing proteins with nature-like properties.
- Noelia Ferruz
- , Steffen Schmidt
- & Birte Höcker
-
Article
| Open Access Decoding kinase-adverse event associations for small molecule kinase inhibitors
Decoding kinase-adverse event associations for small molecule kinase inhibitors
Small molecule kinase inhibitors (SMKIs) are being approved at a fast pace under expedited programs for anticancer treatment. Here, the authors employ a machine-learning model to examine the relationships between kinase targets and adverse events in the trials of 16 FDA-approved SMKIs.
- Xiajing Gong
- , Meng Hu
- & Liang Zhao
-
Article
| Open Access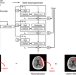 Emergency triage of brain computed tomography via anomaly detection with a deep generative model
Emergency triage of brain computed tomography via anomaly detection with a deep generative model
Triage is essential for the early diagnosis and reporting of emergency patients in the emergency department. Here, the authors develop an anomaly detection algorithm with a deep generative model that reprioritizes radiology worklists and provides lesion attention maps for brain CT images with critical findings.
- Seungjun Lee
- , Boryeong Jeong
- & Namkug Kim
-
Article
| Open Access Accurate somatic variant detection using weakly supervised deep learning
Accurate somatic variant detection using weakly supervised deep learning
Deep learning could be applied to the challenge of somatic variant calling in cancer by making use of large-scale genomic data. Here, the authors develop VarNet, a weakly supervised deep learning model for somatic variant calling in cancer with robust performance across multiple cancer genomics datasets.
- Kiran Krishnamachari
- , Dylan Lu
- & Anders Jacobsen Skanderup
-
Article
| Open Access Identifying multicellular spatiotemporal organization of cells with SpaceFlow
Identifying multicellular spatiotemporal organization of cells with SpaceFlow
A critical task in spatial transcriptomics analysis is to understand inherently spatial relationships among cells. Here, the authors present a deep learning framework to integrate spatial and transcriptional information, spatially extending pseudotime and revealing spatiotemporal organization of cells.
- Honglei Ren
- , Benjamin L. Walker
- & Qing Nie
-
Article
| Open Access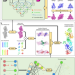 Model building of protein complexes from intermediate-resolution cryo-EM maps with deep learning-guided automatic assembly
Model building of protein complexes from intermediate-resolution cryo-EM maps with deep learning-guided automatic assembly
One challenge in cryo-EM is to build atomic models into intermediate resolution maps. Here, the authors present a deep learning-guided iterative assembling method by integrating AlphaFold, FFTbased fitting, and domain-based refinement.
- Jiahua He
- , Peicong Lin
- & Sheng-You Huang
-
Article
| Open Access Machine learning-based tissue of origin classification for cancer of unknown primary diagnostics using genome-wide mutation features
Machine learning-based tissue of origin classification for cancer of unknown primary diagnostics using genome-wide mutation features
The original tumor location can be unclear for metastatic tumors. Here, the authors show that DNA sequencing of whole genomes can be used to classify metastatic tumors using a machine learning model, Cancer of Unknown Primary Location Resolver, in order to improve diagnosis and inform treatment decisions.
- Luan Nguyen
- , Arne Van Hoeck
- & Edwin Cuppen
-
Article
| Open Access Deep learning from phylogenies to uncover the epidemiological dynamics of outbreaks
Deep learning from phylogenies to uncover the epidemiological dynamics of outbreaks
Widely applicable, accurate and fast inference methods in phylodynamics are needed to fully profit from the richness of genetic data in uncovering the dynamics of epidemics. Here, the authors develop a likelihood-free, simulation-based deep learning approach.
- J. Voznica
- , A. Zhukova
- & O. Gascuel
-
Article
| Open Access Self-evolving vision transformer for chest X-ray diagnosis through knowledge distillation
Self-evolving vision transformer for chest X-ray diagnosis through knowledge distillation
Although deep learning-based computer-aided diagnosis systems have recently achieved expert level performance, developing a robust model requires large, high-quality data with annotations. Here, the authors present a framework which can improve the performance of vision transformer simultaneously with self-supervision and self-training.
- Sangjoon Park
- , Gwanghyun Kim
- & Jong Chul Ye
-
Article
| Open Access A convolutional neural network highlights mutations relevant to antimicrobial resistance in Mycobacterium tuberculosis
A convolutional neural network highlights mutations relevant to antimicrobial resistance in Mycobacterium tuberculosis
Pathogen whole genome sequencing, coupled with statistical and machine learning models, offers a promising solution to multi-drug resistance diagnosis. Here, the authors present two deep convolutional neural networks that predict the antibiotic resistance phenotypes of M. tuberculosis isolates.
- Anna G. Green
- , Chang Ho Yoon
- & Maha Farhat
-
Article
| Open Access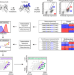 Co-optimization of therapeutic antibody affinity and specificity using machine learning models that generalize to novel mutational space
Co-optimization of therapeutic antibody affinity and specificity using machine learning models that generalize to novel mutational space
Optimising antibody properties such as affinity can be detrimental to other key properties. Here the authors use machine learning to simplify the identification of antibodies with co-optimal levels of affinity and specificity for a clinical-stage antibody that displays high levels of on- and off-target binding.
- Emily K. Makowski
- , Patrick C. Kinnunen
- & Peter M. Tessier
-
Article
| Open Access Deep learning to diagnose Hashimoto’s thyroiditis from sonographic images
Deep learning to diagnose Hashimoto’s thyroiditis from sonographic images
Hashimoto’s thyroiditis (HT) is the main cause of hypothyroidism. Here the authors develop a deep learning model for diagnosis of HT on a large multi-site dataset including image and video data.
- Qiang Zhang
- , Sheng Zhang
- & Xiangchun Li
-
Article
| Open Access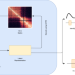 Learning representations of chromatin contacts using a recurrent neural network identifies genomic drivers of conformation
Learning representations of chromatin contacts using a recurrent neural network identifies genomic drivers of conformation
Despite the availability of chromatin conformation capture experiments, discerning the relationship between the 1D genome and 3D conformation remains a challenge. Here, the authors propose a method that produces low-dimensional latent representations that summarize intra-chromosomal Hi-C contacts.
- Kevin B. Dsouza
- , Alexandra Maslova
- & Maxwell W. Libbrecht
-
Article
| Open Access Network-based machine learning approach to predict immunotherapy response in cancer patients
Network-based machine learning approach to predict immunotherapy response in cancer patients
Identifying biomarkers for response to immunotherapy in cancer remains challenging. Here, the authors develop an approach based on network biology and machine learning -NetBio- to identify molecular biomarkers of response to immunotherapy across different cancer types and cohorts.
- JungHo Kong
- , Doyeon Ha
- & Sanguk Kim
-
Article
| Open Access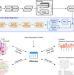 GenomicSuperSignature facilitates interpretation of RNA-seq experiments through robust, efficient comparison to public databases
GenomicSuperSignature facilitates interpretation of RNA-seq experiments through robust, efficient comparison to public databases
Many transcriptomic profiles have been deposited in public archives but are underused for the interpretation of experiments. Here the authors report GenomicSuperSignature for interpreting new transcriptomic datasets through comparison to public archives, without high-performance computing requirements.
- Sehyun Oh
- , Ludwig Geistlinger
- & Sean Davis
-
Article
| Open Access Forest Fire Clustering for single-cell sequencing combines iterative label propagation with parallelized Monte Carlo simulations
Forest Fire Clustering for single-cell sequencing combines iterative label propagation with parallelized Monte Carlo simulations
In the era of single-cell sequencing, there is a growing need to extract insights from data with clustering methods. Here, inspired by forest fire dynamics, the authors devise an algorithm that can cluster single-cell data with minimal prior assumptions and can compute a non-parametric posterior probability for each data point.
- Zhanlin Chen
- , Jeremy Goldwasser
- & Mark Gerstein
-
Article
| Open Access Deep neural network trained on gigapixel images improves lymph node metastasis detection in clinical settings
Deep neural network trained on gigapixel images improves lymph node metastasis detection in clinical settings
The pathological identification of lymph node metastasis in whole-slide images is demanding and tedious. Here, the authors design an artificial-intelligence-assisted assessment workflow to facilitate the routine counting of metastatic LNs.
- Shih-Chiang Huang
- , Chi-Chung Chen
- & Tse-Ching Chen
-
Article
| Open Access Genome-wide mapping of individual replication fork velocities using nanopore sequencing
Genome-wide mapping of individual replication fork velocities using nanopore sequencing
Theulot et al. introduce NanoForkSpeed, a nanopore sequencing-based method to map individual replication fork velocities on entire genomes. NFS shows that fork speed is uniform across yeast chromosomes except for a marked slowdown at pausing sites.
- Bertrand Theulot
- , Laurent Lacroix
- & Benoît Le Tallec
-
Article
| Open Access Deep learning enables reference-free isotropic super-resolution for volumetric fluorescence microscopy
Deep learning enables reference-free isotropic super-resolution for volumetric fluorescence microscopy
Volumetric fluorescence microscopy is often limited by anisotropic spatial resolution. Here, the authors present an unsupervised deep-learning approach that enhances axial resolution by learning from high-resolution lateral images, and demonstrate isotropic resolution and restoration of suppressed visual details.
- Hyoungjun Park
- , Myeongsu Na
- & Jong Chul Ye
-
Article
| Open Access Language models can learn complex molecular distributions
Language models can learn complex molecular distributions
Generative models for the novo molecular design attract enormous interest for exploring the chemical space. Here the authors investigate the application of chemical language models to challenging modeling tasks demonstrating their capability of learning complex molecular distributions.
- Daniel Flam-Shepherd
- , Kevin Zhu
- & Alán Aspuru-Guzik
-
Article
| Open Access A federated graph neural network framework for privacy-preserving personalization
A federated graph neural network framework for privacy-preserving personalization
Mainstream personalization methods rely on centralized Graph Neural Network learning on global graphs, which have considerable privacy risks due to the privacy-sensitive nature of user data. Here, the authors present a federated GNN framework for both effective and privacy-preserving personalization.
- Chuhan Wu
- , Fangzhao Wu
- & Xing Xie
-
Article
| Open Access Estimating tumor mutational burden from RNA-sequencing without a matched-normal sample
Estimating tumor mutational burden from RNA-sequencing without a matched-normal sample
The identification of somatic point mutations in tumor samples is of high clinical value, such as for the development of targeted therapies. Here the authors develop a machine learning pipeline for detecting somatic point mutations from RNA sequencing without a matched-normal sample, and utilize the model's prediction for computing the tumor mutational burden.
- Rotem Katzir
- , Noam Rudberg
- & Keren Yizhak
-
Article
| Open Access Machine learning aided construction of the quorum sensing communication network for human gut microbiota
Machine learning aided construction of the quorum sensing communication network for human gut microbiota
Microbes communicate with each other by Quorum sensing (QS) languages. Here the authors construct a QS database and the QS communication network to decipher intricate QSbased communications and form one of the key knowledge maps for human gut microbiota.
- Shengbo Wu
- , Jie Feng
- & Jianjun Qiao
-
Article
| Open Access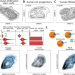 Cell cycle gene regulation dynamics revealed by RNA velocity and deep-learning
Cell cycle gene regulation dynamics revealed by RNA velocity and deep-learning
Single-cell RNA-sequencing technology gives access to cell cycle dynamics without externally perturbing the cell. Here the authors present DeepCycle,a robust deep learning method to infer the cell cycle state in single cells from scRNA-seq data.
- Andrea Riba
- , Attila Oravecz
- & Nacho Molina
-
Article
| Open Access Automated next-generation profiling of genomic alterations in human cancers
Automated next-generation profiling of genomic alterations in human cancers
The genomic profiling of tumours has not been widely adopted in the clinic due to technical and practical hurdles. Here, the authors develop PGDx elio tissue complete, a scalable, standardised and FDA-cleared test comprising a targeted gene panel and automated machine-learning analysis, which detects clinically relevant sequence biomarkers in cancer samples.
- Laurel A. Keefer
- , James R. White
- & Mark Sausen
-
Article
| Open Access A deep learning model and human-machine fusion for prediction of EBV-associated gastric cancer from histopathology
A deep learning model and human-machine fusion for prediction of EBV-associated gastric cancer from histopathology
Epstein–Barr virus-associated gastric cancer shows a robust response to immune checkpoint inhibitors. Here the authors introduce a deep convolutional neural network and its fusion with pathologists for predicting it from histopathology.
- Xueyi Zheng
- , Ruixuan Wang
- & Muyan Cai
-
Article
| Open Access Reconstruct high-resolution 3D genome structures for diverse cell-types using FLAMINGO
Reconstruct high-resolution 3D genome structures for diverse cell-types using FLAMINGO
High-resolution reconstruction of spatial chromosome organisation is in demand. Here the authors report FLAMINGO, for reconstructing high-resolution 3D Genome Organisation from HiC data which they use to generate both 5 kb and 1 kb-resolution 3D chromosomal structures for the human genome.
- Hao Wang
- , Jiaxin Yang
- & Jianrong Wang
-
Article
| Open Access Deep learning of a bacterial and archaeal universal language of life enables transfer learning and illuminates microbial dark matter
Deep learning of a bacterial and archaeal universal language of life enables transfer learning and illuminates microbial dark matter
Computational methods to analyse microbial systems rely on reference databases which do not capture their full functional diversity. Here the authors develop a deep learning model and apply it using transfer learning, creating biologically useful models for multiple different tasks.
- A. Hoarfrost
- , A. Aptekmann
- & Y. Bromberg
-
Article
| Open Access Knowledge integration and decision support for accelerated discovery of antibiotic resistance genes
Knowledge integration and decision support for accelerated discovery of antibiotic resistance genes
Here the authors present KIDS, a knowledge graph integration and phenotypic prediction framework. When applied on antibiotic data, it identifies 6 novel antibiotic resistant E. coli genes that the authors subsequently validate.
- Jason Youn
- , Navneet Rai
- & Ilias Tagkopoulos
-
Article
| Open Access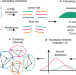 A deep siamese neural network improves metagenome-assembled genomes in microbiome datasets across different environments
A deep siamese neural network improves metagenome-assembled genomes in microbiome datasets across different environments
Here, the authors present SemiBin, a siamese deep neural network framework that incorporates information from reference genomes, able to extract better metagenome-assembled genomes (MAGs) in several host-associated and environmental habitats.
- Shaojun Pan
- , Chengkai Zhu
- & Luis Pedro Coelho
-
Article
| Open Access Leveraging omic features with F3UTER enables identification of unannotated 3’UTRs for synaptic genes
Leveraging omic features with F3UTER enables identification of unannotated 3’UTRs for synaptic genes
3’ untranslated regions (3’UTRs) play a crucial role in regulating gene expression, but our 3’UTR catalogue is incomplete. Here, the authors develop a machine learning-based framework to predict previously unannotated 3’UTRs in 39 human tissues.
- Siddharth Sethi
- , David Zhang
- & Juan A. Botia
-
Article
| Open Access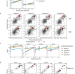 Machine learning-coupled combinatorial mutagenesis enables resource-efficient engineering of CRISPR-Cas9 genome editor activities
Machine learning-coupled combinatorial mutagenesis enables resource-efficient engineering of CRISPR-Cas9 genome editor activities
Screening combinatorial mutants is too massive for wet-lab experiment alone. Here the authors present a machine learning-coupled combinatorial mutagenesis approach to vastly reduce experimental burden for engineering Cas9 genome editing enzymes.
- Dawn G. L. Thean
- , Hoi Yee Chu
- & Alan S. L. Wong
-
Article
| Open Access Communication-efficient federated learning via knowledge distillation
Communication-efficient federated learning via knowledge distillation
This work presents a communication-efficient federated learning method that saves a major fraction of communication cost. It reveals the advantage of reciprocal learning in machine knowledge transfer and the evolutional low-rank properties of deep model updates.
- Chuhan Wu
- , Fangzhao Wu
- & Xing Xie
-
Article
| Open Access Connecting high-resolution 3D chromatin organization with epigenomics
Connecting high-resolution 3D chromatin organization with epigenomics
While large-scale 3D genome architecture is well studied, the limits of resolution have hindered our understanding on the fine scale. Here the authors mapped 1D epigenomic profiles to fine-scale 3D chromatin structures with their deep learning model CAESAR. The model predicted fine-scale structures, such as short-range chromatin loops and stripes, that Hi-C datasets fail to detect.
- Fan Feng
- , Yuan Yao
- & Jie Liu
-
Article
| Open Access Membrane marker selection for segmenting single cell spatial proteomics data
Membrane marker selection for segmenting single cell spatial proteomics data
Cell segmentation of single-cell spatial proteomics data remains a challenge and often relies on the selection of a membrane marker, which is not always known. Here, the authors introduce RAMCES, a method that selects the optimal membrane markers to use for more accurate cell segmentation.
- Monica T. Dayao
- , Maigan Brusko
- & Ziv Bar-Joseph
-
Article
| Open Access Using deep learning to predict abdominal age from liver and pancreas magnetic resonance images
Using deep learning to predict abdominal age from liver and pancreas magnetic resonance images
Approaches to both determine abdominal age and identify risk factors for accelerated abdominal age will help delay the onset of several diseases. Here, the authors build an abdominal age predictor by training convolutional neural networks to predict abdominal age from liver and pancreas MRIs.
- Alan Le Goallec
- , Samuel Diai
- & Chirag J. Patel
-
Article
| Open Access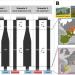 Congruent evolutionary responses of European steppe biota to late Quaternary climate change
Congruent evolutionary responses of European steppe biota to late Quaternary climate change
Quaternary climatic oscillations had a large impact on European biogeography. Using genomic data, machine learning, and approximate Bayesian computation, this study outlines a general scenario in which Quaternary climatic oscillations shaped the evolution of European steppe biota in a congruent way, emphasizing the role of climate underlying patterns of genetic variance at the biome level.
- Philipp Kirschner
- , Manolo F. Perez
- & Peter Schönswetter
-
Article
| Open Access A universal deep neural network for in-depth cleaning of single-cell RNA-Seq data
A universal deep neural network for in-depth cleaning of single-cell RNA-Seq data
Single cell RNA sequencing (scRNA-Seq) is widely used in biomedical research. Here the authors develop a novel AI model-AutoClass, which effectively cleans a wide range of noise and artifacts in scRNA-Seq data and improves downstream analyses.
- Hui Li
- , Cory R. Brouwer
- & Weijun Luo
-
Review Article
| Open Access Current progress and open challenges for applying deep learning across the biosciences
Current progress and open challenges for applying deep learning across the biosciences
Deep learning has enabled advances in understanding biology. In this review, the authors outline advances, and limitations of deep learning in five broad areas and the future challenges for the biosciences.
- Nicolae Sapoval
- , Amirali Aghazadeh
- & Todd J. Treangen
-
Article
| Open Access Knowledge graph-based recommendation framework identifies drivers of resistance in EGFR mutant non-small cell lung cancer
Knowledge graph-based recommendation framework identifies drivers of resistance in EGFR mutant non-small cell lung cancer
Resistance to EGFR inhibitors presents a major obstacle in treating non-small cell lung cancer. Here, the authors develop a recommender system ranking genes based on trade-offs between diverse types of evidence linking them to potential mechanisms of EGFRi resistance.
- Anna Gogleva
- , Dimitris Polychronopoulos
- & Krishna C. Bulusu
-
Article
| Open Access Integrating deep learning and unbiased automated high-content screening to identify complex disease signatures in human fibroblasts
Integrating deep learning and unbiased automated high-content screening to identify complex disease signatures in human fibroblasts
By coupling robotic cell culture systems with artificial intelligence–powered image analysis, Schiff et al. identify previously unseen characteristics of Parkinson’s disease in patient skin cells that distinguish them from healthy controls.
- Lauren Schiff
- , Bianca Migliori
- & Bjarki Johannesson
-
Article
| Open Access A deep-learning approach for online cell identification and trace extraction in functional two-photon calcium imaging
A deep-learning approach for online cell identification and trace extraction in functional two-photon calcium imaging
Processing of two-photon calcium imaging data is generally time-consuming, especially for large fields of view. Here, the authors present CITE-On, a tool based on a convolutional neural network, enabling online automatic cell identification, segmentation, identity tracking, and trace extraction.
- Luca Sità
- , Marco Brondi
- & Tommaso Fellin
-
Article
| Open Access Rapid age-grading and species identification of natural mosquitoes for malaria surveillance
Rapid age-grading and species identification of natural mosquitoes for malaria surveillance
Knowing the age of malaria-transmitting mosquitoes is important to understand transmission risk as only old mosquitoes can transmit the disease. Here, the authors develop a method based on mid-infrared spectra of mosquito cuticle that can rapidly identify the species and age class of main malaria vectors.
- Doreen J. Siria
- , Roger Sanou
- & Francesco Baldini
-
Article
| Open Access All-fiber high-speed image detection enabled by deep learning
All-fiber high-speed image detection enabled by deep learning
Here, the authors demonstrate high-speed imaging through multimode optical fibers by using the high intermodal dispersion to transform 2D spatial information into 1D temporal pulsed signal streams. Deep learning is used to reconstruct the images of micron-scale objects at high frame rates.
- Zhoutian Liu
- , Lele Wang
- & Qirong Xiao
-
Article
| Open Access Machine learning discovery of missing links that mediate alternative branches to plant alkaloids
Machine learning discovery of missing links that mediate alternative branches to plant alkaloids
Producing plant secondary metabolites by microbes is limited by the known enzymatic reactions. Here, the authors apply machine learning to predict missing link enzymes of benzylisoquinoline alkaloid (BIA) biosynthesis in Papaver somniferum, and validate the specialized activities through heterologous production.
- Christopher J. Vavricka
- , Shunsuke Takahashi
- & Tomohisa Hasunuma

 Controlling gene expression with deep generative design of regulatory DNA
Controlling gene expression with deep generative design of regulatory DNA