Featured
-
-
Article
| Open Access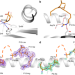 Challenging immunodominance of influenza-specific CD8+ T cell responses restricted by the risk-associated HLA-A*68:01 allomorph
Challenging immunodominance of influenza-specific CD8+ T cell responses restricted by the risk-associated HLA-A*68:01 allomorph
The HLA-A*68:01 allele has been associated with severe influenza disease during the 2009 influenza pandemic. Here, the authors analyze influenza nucleoprotein specific HLA-A*68:01-restricted CD8+ T cells from human donors and show immunodominance of these cells in approximately 35% of HLA-A*68:01-expressing donors.
- C. E. van de Sandt
- , E. B. Clemens
- & K. Kedzierska
-
Article
| Open Access The dynamic proteome of influenza A virus infection identifies M segment splicing as a host range determinant
The dynamic proteome of influenza A virus infection identifies M segment splicing as a host range determinant
Avian influenza A virus (IAV) strains replicate poorly in mammalian hosts, but mechanisms underlying species restriction are incompletely understood. Here, Bogdanow et al. show that avian and mammalian adapted IAV strains have evolved different RNA structure features for regulation of M segment RNA splicing.
- Boris Bogdanow
- , Xi Wang
- & Matthias Selbach
-
Article
| Open Access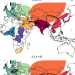 Disentangling the role of Africa in the global spread of H5 highly pathogenic avian influenza
Disentangling the role of Africa in the global spread of H5 highly pathogenic avian influenza
The role of Africa in the global spread of highly pathogenic avian influenza (HPAI) is not well understood. Here, using evolutionary analyses, the authors show that Africa mainly acts as ecological sink for HPAI H5, and reveal varying paths of HPAI incursions either through domestic or wild birds.
- Alice Fusaro
- , Bianca Zecchin
- & Isabella Monne
-
Article
| Open Access Influenza A virus M2 protein triggers mitochondrial DNA-mediated antiviral immune responses
Influenza A virus M2 protein triggers mitochondrial DNA-mediated antiviral immune responses
Cytosolic mitochondrial DNA (mtDNA) plays a role in innate antiviral immunity but how this is triggered during infection remains unclear. Here, the authors provide evidence that the Influenza virus protein M2 stimulates translocation of mtDNA into the cytosol in a MAVS-dependent manner.
- Miyu Moriyama
- , Takumi Koshiba
- & Takeshi Ichinohe
-
Article
| Open Access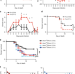 Circadian control of lung inflammation in influenza infection
Circadian control of lung inflammation in influenza infection
The circadian clock affects immune responses, but its role in influenza infection is not well understood. Here, Sengupta et al. show that time of infection and the circadian clock have no effect on lung virus titers, but affect inflammation, morbidity and mortality.
- Shaon Sengupta
- , Soon Y. Tang
- & Garret A. FitzGerald
-
Article
| Open Access Exposure of an occluded hemagglutinin epitope drives selection of a class of cross-protective influenza antibodies
Exposure of an occluded hemagglutinin epitope drives selection of a class of cross-protective influenza antibodies
Antibody cross-reactivity can help to prevent escape mutations from enabling viral escape, but underlying mechanisms are unclear. Here the authors identify influenza hemagglutinin epitopes that are exposed during viral replication and which result in the generation of a class of protective cross-reactive antibodies.
- Yu Adachi
- , Keisuke Tonouchi
- & Yoshimasa Takahashi
-
Article
| Open Access Incomplete influenza A virus genomes occur frequently but are readily complemented during localized viral spread
Incomplete influenza A virus genomes occur frequently but are readily complemented during localized viral spread
The genome of influenza is often incomplete in infected cells, but the implications for infection remain unclear. Here, Jacobs et al. show that an average of 3.6 particles is necessary for productive infection and that coinfection supports efficient complementation within a host but not upon transmission to a new host.
- Nathan T. Jacobs
- , Nina O. Onuoha
- & Anice C. Lowen
-
Article
| Open Access Profiling host ANP32A splicing landscapes to predict influenza A virus polymerase adaptation
Profiling host ANP32A splicing landscapes to predict influenza A virus polymerase adaptation
Polymorphisms in the avian influenza A virus (IAV) polymerase restrict its host range during transmission from birds to mammals. Here, the authors investigate differences in the host chromatin regulator ANP32A regarding IAV polymerase adaptation, and profile ANP32A splicing to predict avian species associated with pre-adaptive human-signatures in the virus.
- Patricia Domingues
- , Davide Eletto
- & Benjamin G. Hale
-
Article
| Open Access Interaction between the nasal microbiota and S. pneumoniae in the context of live-attenuated influenza vaccine
Interaction between the nasal microbiota and S. pneumoniae in the context of live-attenuated influenza vaccine
Using a Streptococcus pneumoniae human challenge model, the authors here show that baseline nasal microbiota profiles are associated with pneumococcal acquisition and density, independent of live-attenuated influenza vaccine (LAIV) administration, and with mucosal cytokine responses to LAIV.
- Wouter A. A. de Steenhuijsen Piters
- , Simon P. Jochems
- & Debby Bogaert
-
Article
| Open Access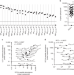 Quantification of epitope abundance reveals the effect of direct and cross-presentation on influenza CTL responses
Quantification of epitope abundance reveals the effect of direct and cross-presentation on influenza CTL responses
CTL responses are critical in protection against pathogens. Here, using mass spectrometry and flow cytometry, the authors characterize the kinetics of influenza A virus class I MHC epitopes cross-presented in professional antigen presenting cells and identify new epitopes that elicit T cell responses in infected mice.
- Ting Wu
- , Jing Guan
- & Nicole L. La Gruta
-
Article
| Open Access The role of platelets in mediating a response to human influenza infection
The role of platelets in mediating a response to human influenza infection
Influenza viremia is rare in human blood and not well studied. Here, the authors show that influenza can be found in human platelets and that platelet engulfment of influenza A results in TLR7-dependent C3 release, which in turn promotes neutrophil-DNA release and formation of platelet-DNA aggregates.
- Milka Koupenova
- , Heather A. Corkrey
- & Jane E. Freedman
-
Article
| Open Access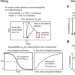 Age-specific differences in the dynamics of protective immunity to influenza
Age-specific differences in the dynamics of protective immunity to influenza
Protective immunity after influenza virus infection is poorly understood. Here, the authors quantify the dynamics of immunity against influenza A virus infections by fitting individual-level mechanistic models to longitudinal serology, and find that the form and dynamics of protection differ between children and adults.
- Sylvia Ranjeva
- , Rahul Subramanian
- & Sarah Cobey
-
Article
| Open Access Influenza A virus ribonucleoproteins form liquid organelles at endoplasmic reticulum exit sites
Influenza A virus ribonucleoproteins form liquid organelles at endoplasmic reticulum exit sites
Influenza A virus forms cytosolic inclusions containing viral ribonucleoproteins. Here, the authors show that viral inclusions form juxtaposed the endoplasmic reticulum and have liquid properties, likely constituting sites of assembly of epidemic and pandemic influenza genomes.
- Marta Alenquer
- , Sílvia Vale-Costa
- & Maria João Amorim
-
Article
| Open Access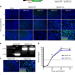 Non-lytic clearance of influenza B virus from infected cells preserves epithelial barrier function
Non-lytic clearance of influenza B virus from infected cells preserves epithelial barrier function
Infection of a cell with influenza B virus (IBV) often results in cell death and the role of surviving cells in pathogenesis is unclear. Here, Dumm et al. generate a recombinant IBV that activates a host-cell reporter to permanently label infected cells, and show that surviving cells are important to preserve epithelial barrier function.
- Rebekah E. Dumm
- , Jessica K. Fiege
- & Nicholas S. Heaton
-
Article
| Open Access Cross-lineage protection by human antibodies binding the influenza B hemagglutinin
Cross-lineage protection by human antibodies binding the influenza B hemagglutinin
Immune recognition of Influenza B virus (IBV) is poorly understood. Here, Liu et al. use flow cytometry to characterize IBV-specific memory B cell responses following seasonal vaccination and show that elicited cross-reactive antibodies can protect against infection, providing a platform for vaccine design.
- Yi Liu
- , Hyon-Xhi Tan
- & Adam K. Wheatley
-
Article
| Open Access Virucidal nano-perforator of viral membrane trapping viral RNAs in the endosome
Virucidal nano-perforator of viral membrane trapping viral RNAs in the endosome
Membrane-disrupting agents that selectively target virus versus host membranes could potentially be potent antivirals. Here the authors incorporate a decoy virus receptor into a nanodisc and show that it ruptures the viral membrane at low pH and traps viral RNAs in the endolysosome for degradation.
- Byoungjae Kong
- , Seokoh Moon
- & Dae-Hyuk Kweon
-
Article
| Open Access Indirect protection from vaccinating children against influenza in households
Indirect protection from vaccinating children against influenza in households
Relevance of indirect protection of household members of vaccinees is unclear. Here, Tsang et al. quantify the direct and indirect protection of vaccination in a randomized controlled trial and show that benefits of individual vaccination remain important even when other household members are vaccinated.
- Tim K. Tsang
- , Vicky J. Fang
- & Simon Cauchemez
-
Article
| Open Access Broad CD8+ T cell cross-recognition of distinct influenza A strains in humans
Broad CD8+ T cell cross-recognition of distinct influenza A strains in humans
Mutations within immunological epitope containing regions of influenza A virus can impair the established immune response between influenza strains and could impact rational vaccine design. Here Grant et al. examine the presence, structural impact and cross reactivity of two human immunodominant influenza epitope variants.
- Emma J. Grant
- , Tracy M. Josephs
- & Katherine Kedzierska
-
Article
| Open Access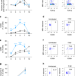 MAIT cells contribute to protection against lethal influenza infection in vivo
MAIT cells contribute to protection against lethal influenza infection in vivo
MAIT cells are abundant in the lungs and confer protection against bacterial pathogens. Whilst activation of these cells has been described during viral infections, here van Wilgenburg and colleagues show that in a murine model MAIT cells contribute to the protective host immune response to influenza virus infection.
- Bonnie van Wilgenburg
- , Liyen Loh
- & Timothy S. C. Hinks
-
Article
| Open Access A naturally protective epitope of limited variability as an influenza vaccine target
A naturally protective epitope of limited variability as an influenza vaccine target
Current influenza vaccine approaches largely focus on highly variable epitopes with high immunogenicity or epitopes of low variability that often have low immunogenicity. Here, Thompson et al. identify a highly immunogenic epitope of limited variability in the head domain of the H1 haemagglutinin and show protection from diverse H1N1 strains in mice.
- Craig P. Thompson
- , José Lourenço
- & Sunetra Gupta
-
Article
| Open Access Phosphoproteomic-based kinase profiling early in influenza virus infection identifies GRK2 as antiviral drug target
Phosphoproteomic-based kinase profiling early in influenza virus infection identifies GRK2 as antiviral drug target
Influenza A virus (IAV) causes annual epidemics and development of antivirals is needed. Here, the authors perform phosphoproteomics during IAV entry and identify GRK2 as drug target, inhibition of which decreases replication of seasonal and pandemic IAV in primary human cells and animal models.
- Emilio Yángüez
- , Annika Hunziker
- & Silke Stertz
-
Article
| Open Access LY6E mediates an evolutionarily conserved enhancement of virus infection by targeting a late entry step
LY6E mediates an evolutionarily conserved enhancement of virus infection by targeting a late entry step
The interferon-induced gene LY6E increases virus infection, but the underlying mechanism is poorly understood. Here, Mar et al. show that LY6E enhances uncoating of influenza A virus after endosomal escape and that viral enhancement by LY6E is conserved across evolution.
- Katrina B. Mar
- , Nicholas R. Rinkenberger
- & John W. Schoggins
-
Article
| Open Access Nucleoside-modified mRNA immunization elicits influenza virus hemagglutinin stalk-specific antibodies
Nucleoside-modified mRNA immunization elicits influenza virus hemagglutinin stalk-specific antibodies
The highly conserved influenza virus hemagglutinin (HA) stalk represents a potential target for a broadly protective vaccine. Here, the authors show that immunization with nucleoside-modified mRNA encoding full-length HA formulated in lipid nanoparticles elicits HA stalk-specific antibodies and protects from heterosubtypic virus infection.
- Norbert Pardi
- , Kaela Parkhouse
- & Drew Weissman
-
Article
| Open Access Nuclear-resident RIG-I senses viral replication inducing antiviral immunity
Nuclear-resident RIG-I senses viral replication inducing antiviral immunity
RIG-I senses cytoplasmic viral RNA, resulting in induction of an antiviral response. Here, the authors identify nuclear RIG-I and show that it binds nuclear influenza A virus RNA, resulting in a cooperative interferon induction along with its cytoplasmic counterpart.
- GuanQun Liu
- , Yao Lu
- & Yan Zhou
-
Article
| Open Access A multifunctional human monoclonal neutralizing antibody that targets a unique conserved epitope on influenza HA
A multifunctional human monoclonal neutralizing antibody that targets a unique conserved epitope on influenza HA
Broadly neutralizing antibodies are potential therapeutics and can aid rational vaccine development. Here, the authors show that the human monoclonal antibody H3v-47 recognizes a highly conserved epitope in HA of H3N2 viruses, inhibits virus replication by blocking egress and other mechanisms, and protects mice from disease.
- Sandhya Bangaru
- , Heng Zhang
- & James E. Crowe Jr.
-
Article
| Open Access Co-regulatory activity of hnRNP K and NS1-BP in influenza and human mRNA splicing
Co-regulatory activity of hnRNP K and NS1-BP in influenza and human mRNA splicing
Alternative splicing of influenza A virus (IAV) M transcript is regulated by hnRNP K and NS1-BP, but mechanistic details are unknown. Here, Thompson et al. show how hnRNP K and NS1-BP bind M mRNA and that these proteins regulate splicing of host transcripts in both the absence and presence of IAV infection.
- Matthew G. Thompson
- , Raquel Muñoz-Moreno
- & Kristen W. Lynch
-
Article
| Open Access Dual-functional peptide with defective interfering genes effectively protects mice against avian and seasonal influenza
Dual-functional peptide with defective interfering genes effectively protects mice against avian and seasonal influenza
A limited number of therapeutics is available to treat influenza A virus (IAV) infections. Here, the authors show that defective interfering genes, delivered with a dual-functional peptide that enables intracellular accumulation and prevents endosomal acidification, inhibit IAV replication in vitro and in vivo.
- Hanjun Zhao
- , Kelvin K. W. To
- & Kwok-Yung Yuen
-
Article
| Open Access Molecular mechanism of influenza A NS1-mediated TRIM25 recognition and inhibition
Molecular mechanism of influenza A NS1-mediated TRIM25 recognition and inhibition
NS1 of influenza A virus inhibits TRIM25 activity, which is an E3 ligase important for induction of the interferon response. Here, Koliopoulos et al. present structures of TRIM25 and NS1 and show how NS1 binding interferes with substrate recognition of TRIM25.
- Marios G. Koliopoulos
- , Mathilde Lethier
- & Katrin Rittinger
-
Article
| Open Access How single mutations affect viral escape from broad and narrow antibodies to H1 influenza hemagglutinin
How single mutations affect viral escape from broad and narrow antibodies to H1 influenza hemagglutinin
Influenza A virus can escape antibodies, but it is unclear how the ease of escape depends on the epitope targeted by an antibody. Here, the authors show that neutralization breadth is an imperfect indicator of the ease of viral escape by single mutations, and that antibodies targeting the stalk of hemagglutinin are harder to escape.
- Michael B. Doud
- , Juhye M. Lee
- & Jesse D. Bloom
-
Article
| Open Access A complex epistatic network limits the mutational reversibility in the influenza hemagglutinin receptor-binding site
A complex epistatic network limits the mutational reversibility in the influenza hemagglutinin receptor-binding site
The receptor-binding site (RBS) of influenza A viruses evolves to evade immune pressure, while maintaining efficient attachment to the host receptor. Wu et al. here identify the complex epistatic network in RBS of H3N2 viruses that limits reversibility of naturally occurring mutations to retain infectivity.
- Nicholas C. Wu
- , Andrew J. Thompson
- & Ian A. Wilson
-
Article
| Open Access Clonally diverse CD38+HLA-DR+CD8+ T cells persist during fatal H7N9 disease
Clonally diverse CD38+HLA-DR+CD8+ T cells persist during fatal H7N9 disease
Virus-specific CD8+ T cells are crucial during H7N9 influenza infection, but CD8+ T cell dysfunction is associated with poor prognosis. Here, the authors use molecular and phenotypic analysis to establish persistence of clonally diverse CD8+ T cell populations during fatal infection.
- Zhongfang Wang
- , Lingyan Zhu
- & Katherine Kedzierska
-
Article
| Open Access Nucleotide resolution mapping of influenza A virus nucleoprotein-RNA interactions reveals RNA features required for replication
Nucleotide resolution mapping of influenza A virus nucleoprotein-RNA interactions reveals RNA features required for replication
Influenza A virus packaging depends on interactions between nucleoprotein (NP) and viral RNA (vRNA), but the pattern of NP binding is unclear. Using PAR-CLIP, Williams et al. here show that NP binds vRNA non-uniformly and that RNA structures in low-NP binding regions are important for packaging.
- Graham D. Williams
- , Dana Townsend
- & Adrianus C. M. Boon
-
Article
| Open Access Double-layered protein nanoparticles induce broad protection against divergent influenza A viruses
Double-layered protein nanoparticles induce broad protection against divergent influenza A viruses
Relatively well conserved domains of influenza A virus (IAV) proteins are potential candidates for the development of a universal IAV vaccine. Here, Deng et al. combine two such conserved antigens (M2e and HA stalk) in a double-layered protein nanoparticle and show that it protects against divergent IAVs in mice.
- Lei Deng
- , Teena Mohan
- & Bao-Zhong Wang
-
Article
| Open Access Importance of the 1+7 configuration of ribonucleoprotein complexes for influenza A virus genome packaging
Importance of the 1+7 configuration of ribonucleoprotein complexes for influenza A virus genome packaging
Influenza A virus (IAV) packages its eight genomic RNA segments in a specific “1+7” pattern. Here, the authors generate IAV that lack one RNA segment and show that ribosomal RNA is packaged in place of the eighth segment, suggesting that the 1+7 pattern is important for particle production.
- Takeshi Noda
- , Shin Murakami
- & Yoshihiro Kawaoka
-
Article
| Open Access Pandemic H1N1 influenza A viruses suppress immunogenic RIPK3-driven dendritic cell death
Pandemic H1N1 influenza A viruses suppress immunogenic RIPK3-driven dendritic cell death
The differences in virus-host interactions resulting in distinct pathogenicity of seasonal and pandemic influenza A viruses (IAV) are not well understood. Here, the authors show that the hemagglutinin segment from pandemic, but not seasonal, IAV suppresses RIPK3-mediated dendritic cell death, thereby reducing T cell activation.
- Boris M. Hartmann
- , Randy A. Albrecht
- & Stuart C. Sealfon
-
Article
| Open Access Influenza virus genome reaches the plasma membrane via a modified endoplasmic reticulum and Rab11-dependent vesicles
Influenza virus genome reaches the plasma membrane via a modified endoplasmic reticulum and Rab11-dependent vesicles
Transport of neo-synthesized influenza A virus viral ribonucleoproteins (vRNPs) from the nucleus to the plasma membrane involves Rab 11 but the mechanism is unclear. Here the authors show that vRNPs are transported through a modified Rab11-positive endoplasmic reticulum and Rab11-dependent vesicles.
- Isabel Fernández de Castro Martin
- , Guillaume Fournier
- & Nadia Naffakh
-
Article
| Open Access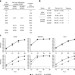 Role of influenza A virus NP acetylation on viral growth and replication
Role of influenza A virus NP acetylation on viral growth and replication
Post-translational modifications of influenza A virus proteins can regulate virus replication, but the effect of nucleoprotein (NP) acetylation is not known. Here, Giese et al. identify four NP lysine residues that are acetylated in infected cells and study their role in polymerase activity and virion release.
- Sebastian Giese
- , Kevin Ciminski
- & Martin Schwemmle
-
Article
| Open Access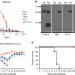 Alveolar macrophages are critical for broadly-reactive antibody-mediated protection against influenza A virus in mice
Alveolar macrophages are critical for broadly-reactive antibody-mediated protection against influenza A virus in mice
Broadly reactive antibodies that recognize influenza A virus HA can be protective, but the mechanism is not completely understood. Here, He et al. show that the inflammatory response and phagocytosis mediated by the interaction between protective antibodies and macrophages are essential for protection.
- Wenqian He
- , Chi-Jene Chen
- & Gene S. Tan
-
Article
| Open Access Epitope-associated and specificity-focused features of EV71-neutralizing antibody repertoires from plasmablasts of infected children
Epitope-associated and specificity-focused features of EV71-neutralizing antibody repertoires from plasmablasts of infected children
Enterovirus 71 is a leading cause of hand-foot-and-mouth disease and herpangina. Here, the authors characterize a large panel of plasmablast-derived IgG mAbs that target the capsid of EV71 to identify neutralizing antibodies induced by natural infection.
- Kuan-Ying Arthur Huang
- , Mei-Feng Chen
- & Tzou-Yien Lin
-
Article
| Open Access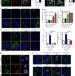 Endosomal NOX2 oxidase exacerbates virus pathogenicity and is a target for antiviral therapy
Endosomal NOX2 oxidase exacerbates virus pathogenicity and is a target for antiviral therapy
Production of reactive oxygen species is an ancient antimicrobial mechanism, but its role in antiviral defense in mammals is unclear. Here, To et al. show that virus infection activates endosomal NOX2 oxidase and restricts TLR7 signaling, and that an endosomal NOX2 inhibitor decreases viral pathogenicity.
- Eunice E. To
- , Ross Vlahos
- & Stavros Selemidis
-
Article
| Open Access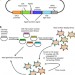 In vitro evolution of an influenza broadly neutralizing antibody is modulated by hemagglutinin receptor specificity
In vitro evolution of an influenza broadly neutralizing antibody is modulated by hemagglutinin receptor specificity
Broadly neutralizing antibodies (bnAbs) against influenza hemagglutinin (HA) have yielded insights for antiviral development. Here, the authors employ saturated mutagenesis of the paratope region of a bnAb combined with yeast display screening using H1 and H3 HAs, and find that a tradeoff exists between Ab affinity and breadth that influenced by disparate modes of receptor binding.
- Nicholas C. Wu
- , Geramie Grande
- & Ian A. Wilson
-
Article
| Open Access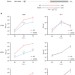 An NS-segment exonic splicing enhancer regulates influenza A virus replication in mammalian cells
An NS-segment exonic splicing enhancer regulates influenza A virus replication in mammalian cells
Some circulating avian influenza A viruses can infect humans, but the mechanism enabling species jump is poorly understood. Here, Huanget al. identify a nucleotide in NEP of avian H7N9 viruses that affects splicing efficiency of the NS segment and supports virus replication in avian and mammalian cells.
- Xiaofeng Huang
- , Min Zheng
- & Honglin Chen
-
Article
| Open Access Comparative influenza protein interactomes identify the role of plakophilin 2 in virus restriction
Comparative influenza protein interactomes identify the role of plakophilin 2 in virus restriction
Protein interaction networks can identify host proteins that affect virus replication. Here, the authors compare the protein interactomes of several influenza A virus strains and identify plakophilin 2 as a restriction factor that inhibits formation of the viral polymerase complex.
- Lingyan Wang
- , Bishi Fu
- & Shitao Li
-
Article
| Open Access A broadly protective therapeutic antibody against influenza B virus with two mechanisms of action
A broadly protective therapeutic antibody against influenza B virus with two mechanisms of action
Influenza B virus (IBV) co-circulates with influenza A virus to cause annual epidemics. Here, Chaiet al. isolate a human monoclonal antibody that binds to a conserved epitope in the viral HA protein, neutralizes IBV strains in vitro, and protects mice against IBV infection.
- Ning Chai
- , Lee R. Swem
- & Man-Wah Tan
-
Article
| Open Access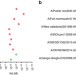 Human antibody 3E1 targets the HA stem region of H1N1 and H5N6 influenza A viruses
Human antibody 3E1 targets the HA stem region of H1N1 and H5N6 influenza A viruses
Treatment of influenza A viruses with broadly neutralizing monoclonal antibodies is an area of active research. Here, the authors characterise a human monoclonal antibody called 3E1 that was reactive against both H1 and H5 viruses in vitroand demonstrated some treatment efficacy in mice.
- Wenshuai Wang
- , Xiaoyu Sun
- & Bing Sun
-
Article
| Open Access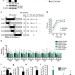 A conserved influenza A virus nucleoprotein code controls specific viral genome packaging
A conserved influenza A virus nucleoprotein code controls specific viral genome packaging
The nucleotide sequence of the eight genomic RNA segments of influenza A virus provides essential packaging signals, but how these sequences are recognized is unknown. Here, Moreira et al. identify conserved amino acids in the viral nucleoprotein that regulate packaging of RNA segments.
- Étori Aguiar Moreira
- , Anna Weber
- & Mindaugas Juozapaitis
-
Article
| Open Access A broadly neutralizing anti-influenza antibody reveals ongoing capacity of haemagglutinin-specific memory B cells to evolve
A broadly neutralizing anti-influenza antibody reveals ongoing capacity of haemagglutinin-specific memory B cells to evolve
A major goal of vaccine design is to protect against a broad range of pathogen strains. Here the authors isolate a new broadly neutralizing antibody against influenza haemagglutinin from human memory B cells, and identify mutations that increase and broaden the neutralization towards H5 HA subtype.
- Ying Fu
- , Zhen Zhang
- & Wayne A. Marasco
-
Article
| Open Access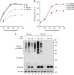 Influenza B virus non-structural protein 1 counteracts ISG15 antiviral activity by sequestering ISGylated viral proteins
Influenza B virus non-structural protein 1 counteracts ISG15 antiviral activity by sequestering ISGylated viral proteins
The ubiquitin-like protein ISG15 can be covalently linked to cellular and viral proteins, but the consequences of this ‘ISGylation’ remain largely unknown. Here, Zhao et al.show that ISGylation of the influenza B virus nucleoprotein inhibits formation of a functional viral replication complex.
- Chen Zhao
- , Haripriya Sridharan
- & Robert M. Krug
-
Article
| Open Access Influenza A virus targets a cGAS-independent STING pathway that controls enveloped RNA viruses
Influenza A virus targets a cGAS-independent STING pathway that controls enveloped RNA viruses
Stimulator of interferon genes (STING) is known to be involved in defence against DNA viruses, but its role in the control of RNA viruses remains poorly explored. Here the authors show that STING participates in an innate immune response to RNA virus infection in a cGAS-independent manner.
- Christian K. Holm
- , Stine H. Rahbek
- & Søren R. Paludan

 Genome-wide CRISPR screen identifies host dependency factors for influenza A virus infection
Genome-wide CRISPR screen identifies host dependency factors for influenza A virus infection