Featured
-
-
Article
| Open Access DNA methylation-based high-resolution mapping of long-distance chromosomal interactions in nucleosome-depleted regions
DNA methylation-based high-resolution mapping of long-distance chromosomal interactions in nucleosome-depleted regions
Here, the authors present MTAC, a method to map chromosomal interactions in budding yeast. By applying MTAC to various viewpoints, they find that most of the long-distance chromosomal interactions detected by MTAC reflect tethering by the nuclear pore complexes.
- Yi Li
- , James Lee
- & Lu Bai
-
Article
| Open Access Two telomere-to-telomere gapless genomes reveal insights into Capsicum evolution and capsaicinoid biosynthesis
Two telomere-to-telomere gapless genomes reveal insights into Capsicum evolution and capsaicinoid biosynthesis
Chili pepper (Capsicum) is an important vegetables known for fruit pungency given by capsaicinoids. Here, the authors assemble the telomere-to-telomere genomes of a pungent pepper C. annuum and its non-pungent wild relative C. rhomboideum and reveal insights into Capsicum evolution and capsaicinoid biosynthesis.
- Weikai Chen
- , Xiangfeng Wang
- & Li Guo
-
Article
| Open Access Transfer learning enables identification of multiple types of RNA modifications using nanopore direct RNA sequencing
Transfer learning enables identification of multiple types of RNA modifications using nanopore direct RNA sequencing
Simultaneous profiling of multiple RNA modifications is a promising yet understudied field of research. Here, authors develop a transferable deep learning framework capable of detecting multiple types of RNA modifications in single nanopore sequencing sample.
- You Wu
- , Wenna Shao
- & Xiang Yu
-
Article
| Open Access tauFisher predicts circadian time from a single sample of bulk and single-cell pseudobulk transcriptomic data
tauFisher predicts circadian time from a single sample of bulk and single-cell pseudobulk transcriptomic data
There is a need to determine circadian time in gene expression datasets. Here, authors built tauFisher, a pipeline that predicts circadian time labels from single transcriptomic samples. tauFisher will be useful for determining body clock time in circadian medicine and for research.
- Junyan Duan
- , Michelle N. Ngo
- & Bogi Andersen
-
Article
| Open Access Determinants of mosaic chromosomal alteration fitness
Determinants of mosaic chromosomal alteration fitness
Here, the authors use passenger mutations to quantify expansion rate in ~6,000 people with mosaic chromosomal alterations in the NHLBI TOPMed cohort, finding associations between growth rate and blood counts along with germline genetic modulators of growth rate.
- Yash Pershad
- , Taralynn Mack
- & Alexander G. Bick
-
Article
| Open Access Genetic influence on within-person longitudinal change in anthropometric traits in the UK Biobank
Genetic influence on within-person longitudinal change in anthropometric traits in the UK Biobank
The availability of longitudinal data in large biobanks is increasing. Here, using data from the UK Biobank, the authors develop and apply analytical approaches to quantify genetic contributions to change over time for traits like height and weight.
- Kathryn E. Kemper
- , Julia Sidorenko
- & Peter M. Visscher
-
Article
| Open Access HIV transmission dynamics and population-wide drug resistance in rural South Africa
HIV transmission dynamics and population-wide drug resistance in rural South Africa
There is limited data on drug resistance in South African communities strongly affected by HIV. In this study, the authors observed low levels of resistance to newer drugs but widespread resistance to older HIV medications in a South African community. Resistance to rilpivirine was detected even in untreated individuals.
- Steven A. Kemp
- , Kimia Kamelian
- & Ravindra K. Gupta
-
Article
| Open Access Mitochondrial complex I deficiency stratifies idiopathic Parkinson’s disease
Mitochondrial complex I deficiency stratifies idiopathic Parkinson’s disease
Idiopathic Parkinson’s disease can be stratified according to the severity of neuronal respiratory complex I deficiency. The emerging disease subtypes show distinct molecular and clinical profiles.
- Irene H. Flønes
- , Lilah Toker
- & Charalampos Tzoulis
-
Article
| Open Access Introgression and disruption of migration routes have shaped the genetic integrity of wildebeest populations
Introgression and disruption of migration routes have shaped the genetic integrity of wildebeest populations
The evolutionary genetics of a keystone savannah species the blue wildebeest, and the related black wildebeest, remain largely unexplored. This study finds evidence for archaic introgression of black wildebeest to blue wildebeest and detrimental effects of human activities on migratory populations.
- Xiaodong Liu
- , Long Lin
- & Rasmus Heller
-
Article
| Open Access KSNP: a fast de Bruijn graph-based haplotyping tool approaching data-in time cost
KSNP: a fast de Bruijn graph-based haplotyping tool approaching data-in time cost
Haplotyping is the process of distinguishing alleles inherited together on a chromosome, a crucial step in assembling and interpreting genome sequences. Here, the authors present a computationally efficient haplotype assembly tool for long read sequencing data.
- Qian Zhou
- , Fahu Ji
- & Jue Ruan
-
Article
| Open Access Allopolyploid origin and diversification of the Hawaiian endemic mints
Allopolyploid origin and diversification of the Hawaiian endemic mints
Hawaiian endemic mints represent the second largest plant radiation in the archipelago. Here, the authors present a reference genome and numerous resequenced individuals to uncover evidence for polyploidy, geographic speciation and localized hybridization underlying diversification in this lineage
- Crystal M. Tomlin
- , Sitaram Rajaraman
- & Charlotte Lindqvist
-
Article
| Open Access DNA binding analysis of rare variants in homeodomains reveals homeodomain specificity-determining residues
DNA binding analysis of rare variants in homeodomains reveals homeodomain specificity-determining residues
Analysis of 92 human homeodomain mutants, including disease-associated variants and variants of uncertain significance, reveals variants with altered DNA binding affinity and/or specificity and specificity-determining positions.
- Kian Hong Kock
- , Patrick K. Kimes
- & Martha L. Bulyk
-
Article
| Open Access A chromosomal-scale genome assembly of modern cultivated hybrid sugarcane provides insights into origination and evolution
A chromosomal-scale genome assembly of modern cultivated hybrid sugarcane provides insights into origination and evolution
Modern sugarcane cultivars have complicated genome due to interspecific crosses and multiple backcrossing. Here, the authors report the haplotype-resolved, chromosome-level genome assembly of a modern hybrid sugarcane cultivar and reveal the expansion of genes related to sugar accumulation and smut resistance.
- Yixue Bao
- , Qing Zhang
- & Muqing Zhang
-
Article
| Open Access Early detection of emerging viral variants through analysis of community structure of coordinated substitution networks
Early detection of emerging viral variants through analysis of community structure of coordinated substitution networks
Rise of new viral strains is a major public health challenge, demanding advanced detection and forecasting methods. This study shows how examining communities within networks of viral mutations enables early detection of emerging strains.
- Fatemeh Mohebbi
- , Alex Zelikovsky
- & Pavel Skums
-
Article
| Open Access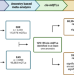 Genetic control of DNA methylation is largely shared across European and East Asian populations
Genetic control of DNA methylation is largely shared across European and East Asian populations
Most functional genomic resources have been based on individuals with European ancestry. Here, the authors perform DNA methylation quantitative trait locus (mQTL) analyses in 3,701 European and 2,099 East Asian individuals to identify thousands of genetic variants, a large degree of which are shared between the two populations.
- Alesha A. Hatton
- , Fei-Fei Cheng
- & Allan F. McRae
-
Article
| Open Access Developmental progression of DNA double-strand break repair deciphered by a single-allele resolution mutation classifier
Developmental progression of DNA double-strand break repair deciphered by a single-allele resolution mutation classifier
DNA double-strand breaks (DSBs) are repaired by a hierarchically regulated network of pathways. Here, authors develop ICP for deciphering somatic DSB repair patterns in multicellular organisms and discover developmental regulation in flies and mosquitoes, enabling tracking of mutant alleles and interhomolog copying of gene cassettes.
- Zhiqian Li
- , Lang You
- & Ethan Bier
-
Article
| Open Access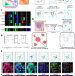 Mapping human tissues with highly multiplexed RNA in situ hybridization
Mapping human tissues with highly multiplexed RNA in situ hybridization
Application of multiplexed RNA in situ mapping techniques to human tissues remains challenging. Here, the authors report DART-FISH, a padlock probe-based technology capable of profiling large numbers of genes in centimetre-sized human tissue sections.
- Kian Kalhor
- , Chien-Ju Chen
- & Kun Zhang
-
Article
| Open Access A universal molecular control for DNA, mRNA and protein expression
A universal molecular control for DNA, mRNA and protein expression
Multi-omics analyses powerfully combine gene expression and translation, however no available controls can be used across these techniques. Here the authors develop pREF, a universal control construct designed for use in DNA, RNA and protein analyses.
- Helen M. Gunter
- , Scott E. Youlten
- & Tim R. Mercer
-
Article
| Open Access Exemestane plus everolimus and palbociclib in metastatic breast cancer: clinical response and genomic/transcriptomic determinants of resistance in a phase I/II trial
Exemestane plus everolimus and palbociclib in metastatic breast cancer: clinical response and genomic/transcriptomic determinants of resistance in a phase I/II trial
Intrinsic and acquired resistances to CDK4/6 inhibitors have been described in patients with breast cancer. Here the authors report the results from a phase I/II clinical trial of the aromatase inhibitor exemestane plus everolimus (mTOR inhibitor) and palbociclib (CDK4/6i) in patients with metastatic breast cancer, assessing safety, clinical efficacy, as well as genomic and transcriptomic determinants of resistance.
- Jorge Gómez Tejeda Zañudo
- , Romualdo Barroso-Sousa
- & Nikhil Wagle
-
Article
| Open Access Highly sensitive spatial transcriptomics using FISHnCHIPs of multiple co-expressed genes
Highly sensitive spatial transcriptomics using FISHnCHIPs of multiple co-expressed genes
Leveraging the fact that eukaryotic genomes are organized into gene modules, FISHnCHIPs images multiple co-expressed genes simultaneously for sensitive and high throughput profiling of gene programs and cell types in tissues.
- Xinrui Zhou
- , Wan Yi Seow
- & Kok Hao Chen
-
Article
| Open Access Charting cellular differentiation trajectories with Ricci flow
Charting cellular differentiation trajectories with Ricci flow
When stem cells develop into tissues intracellular signalling is rewired, errors in this process lead to cancer. Here, authors applied tools from differential geometry made by Albert Einstein’s General Relativity to understand and predict biological network rewiring in health and disease.
- Anthony Baptista
- , Ben D. MacArthur
- & Christopher R. S. Banerji
-
Article
| Open Access BASALT refines binning from metagenomic data and increases resolution of genome-resolved metagenomic analysis
BASALT refines binning from metagenomic data and increases resolution of genome-resolved metagenomic analysis
Binning is an essential step in genome-resolved metagenomic analysis in which assembled contigs originating from the same source population are clustered. However it is challenging, especially for low abundance microbial species. Here the authors introduce a toolkit that integrates multiple prominent binning tools and AI for efficient and high-resolution recovery of non-redundant bins from short- and long-read metagenomic sequencing datasets.
- Zhiguang Qiu
- , Li Yuan
- & Ke Yu
-
Article
| Open Access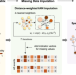 Interrogations of single-cell RNA splicing landscapes with SCASL define new cell identities with physiological relevance
Interrogations of single-cell RNA splicing landscapes with SCASL define new cell identities with physiological relevance
RNA splicing serves as a critical layer of gene expression regulation. Here, authors introduce SCASL for investigating the heterogeneity of RNA splicing landscapes at single-cell resolution, offering a novel scheme for classifying cell identities with physiological relevance.
- Xianke Xiang
- , Yao He
- & Xuerui Yang
-
Article
| Open Access The defensome of complex bacterial communities
The defensome of complex bacterial communities
Bacteria have evolved numerous innate and adaptive defence mechanisms. Here, Beavogui et al characterise the impact of biogeography, genetic mobility, and clustering in defense islands, on the defence systems of soil, marine, and human gut bacterial populations genomes.
- Angelina Beavogui
- , Auriane Lacroix
- & Pedro H. Oliveira
-
Article
| Open Access Statistical method scDEED for detecting dubious 2D single-cell embeddings and optimizing t-SNE and UMAP hyperparameters
Statistical method scDEED for detecting dubious 2D single-cell embeddings and optimizing t-SNE and UMAP hyperparameters
2D visualisation of single-cell data is highly impacted by the hyperparameter setting of the 2D embedding method, such as t-SNE and UMAP. Here, authors develop a statistical method scDEED to detect dubious cell embeddings and optimise the hyperparameter setting for trustworthy visualisation.
- Lucy Xia
- , Christy Lee
- & Jingyi Jessica Li
-
Article
| Open Access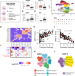 Complex regulatory networks influence pluripotent cell state transitions in human iPSCs
Complex regulatory networks influence pluripotent cell state transitions in human iPSCs
Stem cells exist in vitro in a spectrum of interconvertible pluripotent states. Here, authors show that pluripotency and self-renewal processes have a high level of regulatory complexity and suggest that genetic factors contribute to cell state transitions in human iPSC lines.
- Timothy D. Arthur
- , Jennifer P. Nguyen
- & Kelly A. Frazer
-
Article
| Open Access Genomic evidence for rediploidization and adaptive evolution following the whole-genome triplication
Genomic evidence for rediploidization and adaptive evolution following the whole-genome triplication
Polyploidization-rediploidization process plays an important role in plant adaptive evolution. Here, the authors assemble the genomes of mangrove species Sonneratia alba and its inland relative Lagerstroemia speciosa, and reveal genomic evidence for rediploidization and adaptive evolution after the whole-genome triplication.
- Xiao Feng
- , Qipian Chen
- & Ziwen He
-
Article
| Open Access Converging and evolving immuno-genomic routes toward immune escape in breast cancer
Converging and evolving immuno-genomic routes toward immune escape in breast cancer
Immune response during breast cancer progression remains to be explored. Here, the characterisation of sequential and parallel multiregion samples of an index patient and a cohort of metastatic triple-negative breast cancers reveals convergent immune evasion mechanisms and an increase in tumor genomic heterogeneity.
- Juan Blanco-Heredia
- , Carla Anjos Souza
- & Leticia De Mattos-Arruda
-
Article
| Open Access Cases of trisomy 21 and trisomy 18 among historic and prehistoric individuals discovered from ancient DNA
Cases of trisomy 21 and trisomy 18 among historic and prehistoric individuals discovered from ancient DNA
Information on the occurrence of aneuploidies in prehistory human populations are rare. Here, from a large screen of ancient human genomes and osteological examination, the authors find genetic evidence for six cases of trisomy 21 (Down syndrome) and one case of trisomy 18 (Edwards syndrome) in historic and prehistoric infants.
- Adam Benjamin Rohrlach
- , Maïté Rivollat
- & Kay Prüfer
-
Article
| Open Access Cepharanthine analogs mining and genomes of Stephania accelerate anti-coronavirus drug discovery
Cepharanthine analogs mining and genomes of Stephania accelerate anti-coronavirus drug discovery
Cepharanthine is a secondary metabolite isolated from Stephania with a variety of medicinal properties. Here, the authors assembled three Stephania genomes, propose cepharanthine biosynthetic pathway, and assess the antiviral potential of cepharanthine-related metabolites.
- Liang Leng
- , Zhichao Xu
- & Shilin Chen
-
Article
| Open Access A genome and gene catalog of the aquatic microbiomes of the Tibetan Plateau
A genome and gene catalog of the aquatic microbiomes of the Tibetan Plateau
The Tibetan Plateau is the largest plateau in the world and hosts a variety of aquatic ecosystems. Here, the authors present a gene and genome catalogue of Tibetan Plateau aquatic microbiomes, greatly expanding known taxonomic and functional diversity for the region and giving insights into its microbial biogeography.
- Mingyue Cheng
- , Shuai Luo
- & Kang Ning
-
Article
| Open Access Matrin3 mediates differentiation through stabilizing chromatin loop-domain interactions and YY1 mediated enhancer-promoter interactions
Matrin3 mediates differentiation through stabilizing chromatin loop-domain interactions and YY1 mediated enhancer-promoter interactions
Alterations in proteins within nuclear compartments often lead to changes in chromosomal architecture. Here, using acute targeted protein degradation, the authors reveal that the nuclear complex protein Matrin3 directly mediates differentiation through stabilizing chromatin loop domain interactions.
- Tianxin Liu
- , Qian Zhu
- & Stuart H. Orkin
-
Article
| Open Access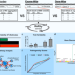 A method to estimate the contribution of rare coding variants to complex trait heritability
A method to estimate the contribution of rare coding variants to complex trait heritability
The contribution of rare variants to complex traits has not been well studied. Here, the authors present RARity, a method to assess rare variant heritability without assuming a particular genetic architecture and enabling both gene-level and exome-wide heritability estimation of continuous traits.
- Nazia Pathan
- , Wei Q. Deng
- & Guillaume Paré
-
Article
| Open Access Unraveling the genetic architecture of congenital vertebral malformation with reference to the developing spine
Unraveling the genetic architecture of congenital vertebral malformation with reference to the developing spine
Congenital vertebral malformation has a complex genetic architecture that isn’t fully understood. Here, the authors explore the genetic architecture of congenital vertebral malformation through case-control rare variant genetic analyses and embryonic transcriptome analyses of the developing spine.
- Sen Zhao
- , Hengqiang Zhao
- & Nan Wu
-
Article
| Open Access Synchrony of Bird Migration with Global Dispersal of Avian Influenza Reveals Exposed Bird Orders
Synchrony of Bird Migration with Global Dispersal of Avian Influenza Reveals Exposed Bird Orders
Highly pathogenic avian influenza virus subtype H5 is an important pathogen of wild birds and poultry that has also caused infection in humans and other mammals. Here the authors use wild bird movement tracking data and virus genome sequences to quantify how seasonal bird migration facilitates global dispersal of the virus.
- Qiqi Yang
- , Ben Wang
- & Bryan Grenfell
-
Article
| Open Access Unzipped genome assemblies of polyploid root-knot nematodes reveal unusual and clade-specific telomeric repeats
Unzipped genome assemblies of polyploid root-knot nematodes reveal unusual and clade-specific telomeric repeats
Telomeres protect the extremities of linear chromosomes and are involved in ageing, senescence and genome stability. Here, the authors have identified peculiar and specific telomeric DNA repeats in the genomes of devastating plant-parasitic nematodes, opening new perspectives for their control.
- Ana Paula Zotta Mota
- , Georgios D. Koutsovoulos
- & Etienne G. J. Danchin
-
Article
| Open Access Orchestrating chromosome conformation capture analysis with Bioconductor
Orchestrating chromosome conformation capture analysis with Bioconductor
The Bioconductor project aims to develop R packages for analysis of genomic datasets. Here the authors show the HiCExperiment package suite and its companion online book (https://bioconductor.org/books/OHCA/) which present data structures, computational methods and visualization tools available in Bioconductor to investigate chromatin conformation capture (3C) data in R.
- Jacques Serizay
- , Cyril Matthey-Doret
- & Romain Koszul
-
Article
| Open Access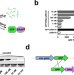 Engineering stringent genetic biocontainment of yeast with a protein stability switch
Engineering stringent genetic biocontainment of yeast with a protein stability switch
Comprehensive safety measures are lacking to employ engineered microorganisms in open-environment applications. Here the authors introduce a genetically encoded biocontainment system for engineered microorganisms based on conditional protein stability.
- Stefan A. Hoffmann
- & Yizhi Cai
-
Article
| Open Access The impacts of active and self-supervised learning on efficient annotation of single-cell expression data
The impacts of active and self-supervised learning on efficient annotation of single-cell expression data
Cell type annotation for single-cell data is challenging. Here, authors explore active and self-supervised learning and introduce adaptive reweighting as a tailored heuristic, demonstrating competitive performance and showing that incorporating prior knowledge enhances cell type annotation accuracy.
- Michael J. Geuenich
- , Dae-won Gong
- & Kieran R. Campbell
-
Article
| Open Access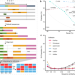 A sequence-aware merger of genomic structural variations at population scale
A sequence-aware merger of genomic structural variations at population scale
Existing tools for structural variations (SVs) calling and merging often lead to fragmented SVs and the potential of introducing unnecessary errors. Here, the authors report the PanPop pipeline to address these issues by implementing sequence-aware SV merging algorithm to efficiently merge SVs of various types.
- Zeyu Zheng
- , Mingjia Zhu
- & Yongzhi Yang
-
Article
| Open Access Multiplexed screening reveals how cancer-specific alternative polyadenylation shapes tumor growth in vivo
Multiplexed screening reveals how cancer-specific alternative polyadenylation shapes tumor growth in vivo
Dysregulation of alternative polyadenylation (APA) is associated with poor prognosis in cancer but its functional role is less clear. Here, the authors develop a CRISPR-Cas9- based screen to determine the effects of different APA events on melanoma growth in mouse models.
- Austin M. Gabel
- , Andrea E. Belleville
- & Robert K. Bradley
-
Comment
| Open Access Implementing community-engaged pharmacogenomics in Indigenous communities
Implementing community-engaged pharmacogenomics in Indigenous communities
Innovative pharmacogenomic approaches (genetic variation related to medication response) are needed to reduce disease and disparities in Indigenous communities. We support community-based pharmacogenomics research, inclusive of Indigenous values and priorities, to improve the health and well-being of Indigenous peoples.
- Katrina G. Claw
- , Casey R. Dorr
- & Erica L. Woodahl
-
Article
| Open Access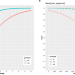 Utility of long-read sequencing for All of Us
Utility of long-read sequencing for All of Us
Using All of Us pilot data, the authors compared short- and long-read performance across medically relevant genes and showcased the utility of long reads to improve variant detection and phasing in easy and hard to resolve medically relevant genes.
- M. Mahmoud
- , Y. Huang
- & F. J. Sedlazeck
-
Article
| Open Access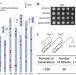 Large-scale genomic rearrangements boost SCRaMbLE in Saccharomyces cerevisiae
Large-scale genomic rearrangements boost SCRaMbLE in Saccharomyces cerevisiae
Synthetic Chromosome Rearrangement and Modification by LoxP-mediated Evolution (SCRaMbLE) is a promising tool to study genomic rearrangements. Here the authors present an engineered yeast strain with 83 sparsely distributed loxPsym sites across the genome can genrerate large-scale genomic rearrangements, which benefits cell fitness under stress and boosts the SCRaMbLE system when combined with synthetic chromosomes.
- Li Cheng
- , Shijun Zhao
- & Junbiao Dai
-
Article
| Open Access SiFT: uncovering hidden biological processes by probabilistic filtering of single-cell data
SiFT: uncovering hidden biological processes by probabilistic filtering of single-cell data
Cells simultaneously encode multiple signals, some harder to recover. Here, authors introduce SiFT (Signal FilTering), a kernel-based projection method, revealing underlying biological processes in single-cell data.
- Zoe Piran
- & Mor Nitzan
-
Article
| Open Access Investigating the etiologies of non-malarial febrile illness in Senegal using metagenomic sequencing
Investigating the etiologies of non-malarial febrile illness in Senegal using metagenomic sequencing
Non-malarial febrile illnesses have a range of potential aetiologies which are difficult to diagnose and therefore treat. Here, the authors investigate the causes of acute febrile illness in a peri-urban area of Senegal with low malaria incidence using untargeted and targeted sequencing methods.
- Zoë C. Levine
- , Aita Sene
- & Katherine J. Siddle
-
Article
| Open Access A chromosome-scale assembly reveals chromosomal aberrations and exchanges generating genetic diversity in Coffea arabica germplasm
A chromosome-scale assembly reveals chromosomal aberrations and exchanges generating genetic diversity in Coffea arabica germplasm
Coffea arabica is an allotetraploid hybrid of C. eugenioides and C. canephora and contributes to approximately 60% of world coffee production. Here, the authors report its chromosome-level genome assembly and identify that chromosomal abnormalities and introgression from C. canephora may contribute to diversity and pathogen resistance.
- Simone Scalabrin
- , Gabriele Magris
- & Michele Morgante
-
Article
| Open Access Systematic detection of co-infection and intra-host recombination in more than 2 million global SARS-CoV-2 samples
Systematic detection of co-infection and intra-host recombination in more than 2 million global SARS-CoV-2 samples
SARS-CoV-2 coinfections may lead to recombination events which could be important in the emergence of new variants. Here, the authors develop an automated bioinformatics pipeline to identify coinfections in genomic data and test it on >2 million publicly available raw read data sets collected globally.
- Orsolya Anna Pipek
- , Anna Medgyes-Horváth
- & István Csabai
-
Article
| Open Access DiffDomain enables identification of structurally reorganized topologically associating domains
DiffDomain enables identification of structurally reorganized topologically associating domains
Topologically associating domains (TADs) are critical structural units in 3D genome organization, and their reorganization between health and disease states is associated with essential genome functions. However, computational methods for identifying reorganized TADs are still in the early stages of development. Here, the authors present an algorithm leveraging random matrix theory to identify reorganized TADs.
- Dunming Hua
- , Ming Gu
- & Dechao Tian

 Far-East Asian Toxoplasma isolates share ancestry with North and South/Central American recombinant lineages
Far-East Asian Toxoplasma isolates share ancestry with North and South/Central American recombinant lineages