Featured
-
-
Article |
 Amino acid coevolution reveals three-dimensional structure and functional domains of insect odorant receptors
Amino acid coevolution reveals three-dimensional structure and functional domains of insect odorant receptors
The structure of insect odorant receptors (ORs) has remained elusive due to their lack of homology to other proteins and the inability to obtain OR crystals. Here, the authors use amino acid evolutionary covariation patterns to fold these proteins de novoand generate the first three-dimensional models of insect ORs.
- Thomas A. Hopf
- , Satoshi Morinaga
- & Richard Benton
-
Article |
 Transcriptome meta-analysis of lung cancer reveals recurrent aberrations in NRG1 and Hippo pathway genes
Transcriptome meta-analysis of lung cancer reveals recurrent aberrations in NRG1 and Hippo pathway genes
Targeted cancer therapy requires knowledge of driver aberrations. Here the authors perform large-scale transcriptome analysis, and show that gene fusions in NRG1, NF1and Hippo pathway genes are recurrent mostly among lung cancers lacking known driver mutations.
- Saravana M. Dhanasekaran
- , O Alejandro Balbin
- & Arul M. Chinnaiyan
-
Article
| Open Access Decelerated genome evolution in modern vertebrates revealed by analysis of multiple lancelet genomes
Decelerated genome evolution in modern vertebrates revealed by analysis of multiple lancelet genomes
The lancelet, or amphioxus, is an extant basal chordate that diverged from other chordate lineages about 550 million years ago. Here the authors sequence and assemble the diploid genome of a male adult of the Chinese lancelet, B. belcheri, and highlight genomic features that may have played an important role in the origin and evolution of vertebrates.
- Shengfeng Huang
- , Zelin Chen
- & Anlong Xu
-
Article
| Open Access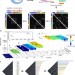 High-quality genome (re)assembly using chromosomal contact data
High-quality genome (re)assembly using chromosomal contact data
The correct assembly of genomes from sequencing data remains a challenge due to difficulties in correctly assigning the location of repeated DNA elements. Here the authors describe GRAAL, an algorithm that utilizes genome-wide chromosome contact data within a probabilistic framework to produce accurate genome assemblies.
- Hervé Marie-Nelly
- , Martial Marbouty
- & Romain Koszul
-
Article |
 Three-dimensional eukaryotic genomic organization is strongly correlated with codon usage expression and function
Three-dimensional eukaryotic genomic organization is strongly correlated with codon usage expression and function
The distribution of genes in eukaryotic genomes is not random. Here Diament et al.show, via a novel codon-based metric, that genes with shared function and similar expression levels tend to be close in the three-dimensional conformation of the yeast, a model plant species, mouse and human genomes.
- Alon Diament
- , Ron Y. Pinter
- & Tamir Tuller
-
Article
| Open Access Transcriptome analysis reveals dysregulation of innate immune response genes and neuronal activity-dependent genes in autism
Transcriptome analysis reveals dysregulation of innate immune response genes and neuronal activity-dependent genes in autism
Autism spectrum disorder (ASD) is a common, highly heritable neurodevelopmental condition characterized by marked genetic heterogeneity. In this study, the authors use RNA sequencing analyses to characterize differences in the transcriptome between autistic and typically developing brains.
- Simone Gupta
- , Shannon E. Ellis
- & Dan E. Arking
-
Article
| Open Access Khoisan hunter-gatherers have been the largest population throughout most of modern-human demographic history
Khoisan hunter-gatherers have been the largest population throughout most of modern-human demographic history
The expansion of Bantu agriculturalists 3,800 years ago in sub-Saharan Africa established first contact with Khoisan hunter-gatherers living in parts of Southern Africa. Sequencing the genomes of five Namibian-Khoisan hunter-gatherers and one Bantu individual tells a tale of admixture and isolation in the early history of modern human populations.
- Hie Lim Kim
- , Aakrosh Ratan
- & Stephan C. Schuster
-
Article |
 Somatic transcriptome priming gates lineage-specific differentiation potential of human-induced pluripotent stem cell states
Somatic transcriptome priming gates lineage-specific differentiation potential of human-induced pluripotent stem cell states
Molecular and functional differences between induced pluripotent stem cells (iPSCs) derived from distinct cell types have been described. Here the authors show, by comparing human iPSCs derived from fibroblasts or cord blood, that the competence in activating developmental genes upon differentiation is influenced by the donor cell of origin.
- Jong-Hee Lee
- , Jung Bok Lee
- & Mickie Bhatia
-
Article
| Open Access Mudskipper genomes provide insights into the terrestrial adaptation of amphibious fishes
Mudskipper genomes provide insights into the terrestrial adaptation of amphibious fishes
Mudskippers are amphibious fishes that have adapted to live on mudflats. Here, the authors sequence the genomes of four different mudskipper species and highlight genetic changes that may have had an evolutionary role in the water-to-land transition of vertebrates.
- Xinxin You
- , Chao Bian
- & Qiong Shi
-
Article
| Open Access Multiple haplotype-resolved genomes reveal population patterns of gene and protein diplotypes
Multiple haplotype-resolved genomes reveal population patterns of gene and protein diplotypes
Knowing which genetic variants exist on either parental chromosome requires diploid human genomes to be phased. Here the authors generate haplotype-resolved genomes and identify a large diversity of haploid and diploid gene forms, a common diplotypic proteome, and an abundance of cisconfigurations of mutations, highlighting the functional importance of diploidy.
- Margret R. Hoehe
- , George M. Church
- & Thomas Huebsch
-
Article |
 Biology of a widespread uncultivated archaeon that contributes to carbon fixation in the subsurface
Biology of a widespread uncultivated archaeon that contributes to carbon fixation in the subsurface
Research on microbes that inhabit the Earth's subsurface is mostly based on metagenomic information only. Here, Probst et al. combine metagenomics with ultrastructural and functional analyses to study the biology of a group of uncultivated subsurface archaea, the SM1 Euryarchaeon lineage.
- Alexander J. Probst
- , Thomas Weinmaier
- & Christine Moissl-Eichinger
-
Article |
 Regulatory network decoded from epigenomes of surface ectoderm-derived cell types
Regulatory network decoded from epigenomes of surface ectoderm-derived cell types
Epigenomes are thought to retain molecular memories of their developmental history. Here, by comparing differentially methylated regions of genomes from different cells, the authors reveal an epigenetic signature that underlies a shared gene regulatory network with a common developmental origin.
- Rebecca F. Lowdon
- , Bo Zhang
- & Jeffrey B. Cheng
-
Article |
 TALEN and CRISPR/Cas9-mediated genome editing in the early-branching metazoan Nematostella vectensis
TALEN and CRISPR/Cas9-mediated genome editing in the early-branching metazoan Nematostella vectensis
Genome editing has yet to be performed in non-bilaterian phyla. Here, Ikmi et al. develop techniques to use both TALEN and CRISPR/Cas9 in the sea anemone, Nematostella vectensis, and further leverage a locus expressing an endogenous fluorescent protein as a landing site for homologous recombination-mediated transgenesis.
- Aissam Ikmi
- , Sean A. McKinney
- & Matthew C. Gibson
-
Article
| Open Access The draft genome of the large yellow croaker reveals well-developed innate immunity
The draft genome of the large yellow croaker reveals well-developed innate immunity
The large yellow croaker, Larimichthys crocea, is an economically important marine fish in China. Here, the authors sequence the draft genome of a wild large yellow croaker and highlight genes that may have played a role in the development of innate immunity in this species.
- Changwen Wu
- , Di Zhang
- & Yun Liu
-
Article
| Open Access Genome sequence of mungbean and insights into evolution within Vigna species
Genome sequence of mungbean and insights into evolution within Vigna species
Mungbean is a fast-growing and warm-season legume crop, cultivated mainly in Asia. Here, the authors sequence the genomes of both wild and domesticated mungbean varieties and, together with detailed transcriptome data, provide insight into mungbean domestication, polyploidization and speciation.
- Yang Jae Kang
- , Sue K. Kim
- & Suk-Ha Lee
-
Article
| Open Access Endogenous florendoviruses are major components of plant genomes and hallmarks of virus evolution
Endogenous florendoviruses are major components of plant genomes and hallmarks of virus evolution
Endogenous viral elements have been extensively described in animals but their significance in plants is less well understood. Here, Geering et al. describe a new group of endogenous pararetroviruses, called florendoviruses, which have colonized the genomes of many important crop species.
- Andrew D. W. Geering
- , Florian Maumus
- & Pierre-Yves Teycheney
-
Article
| Open Access The complex jujube genome provides insights into fruit tree biology
The complex jujube genome provides insights into fruit tree biology
The jujube is a major dry fruit crop in China and is commonly used for medicinal purposes. Here the authors sequence the genome and transcriptome of the most widely cultivated jujube cultivar, Dongzao, and highlight the genetic and molecular basis of agronomically important jujube traits, such as vitamin C content.
- Meng-Jun Liu
- , Jin Zhao
- & Long-Hai Luo
-
Article
| Open Access Genome flux and stasis in a five millennium transect of European prehistory
Genome flux and stasis in a five millennium transect of European prehistory
Recent advances in high-throughput sequencing techniques have enabled the analysis of ancient human genomes. Here the authors sequence ancient human genomes that span a period of 5,000 years, to understand the ancestral influence on Europe's genetic landscape.
- Cristina Gamba
- , Eppie R. Jones
- & Ron Pinhasi
-
Article |
 Camelid genomes reveal evolution and adaptation to desert environments
Camelid genomes reveal evolution and adaptation to desert environments
Comparative genomics can provide valuable insights on adaptations to hostile environments. Here, the authors sequence the genomes and transcriptomes of the Bactrian camel, dromedary and alpaca, to reveal the demographic history of the group as well as metabolic adaptations to the desert environment.
- Huiguang Wu
- , Xuanmin Guang
- & Jun Wang
-
Article
| Open Access The cavefish genome reveals candidate genes for eye loss
The cavefish genome reveals candidate genes for eye loss
Populations of the cave fish Astyanax mexicanus exhibit a variety of traits that evolved repeatedly and independently from its surface counterparts. Here the authors present a de novo genome assembly for A. mexicanusand identify candidate genes for eye loss and reduced pigmentation.
- Suzanne E. McGaugh
- , Joshua B. Gross
- & Wesley C. Warren
-
Article |
 Morphological and population genomic evidence that human faces have evolved to signal individual identity
Morphological and population genomic evidence that human faces have evolved to signal individual identity
The evolution of facial identity is poorly understood. Here, the authors show that human faces have elevated phenotypic variation as well as low between-trait correlations and that the regions surrounding face-associated genes show elevated diversity, which is consistent with frequency-dependent selection.
- Michael J. Sheehan
- & Michael W. Nachman
-
Article |
 The plastid ancestor originated among one of the major cyanobacterial lineages
The plastid ancestor originated among one of the major cyanobacterial lineages
Chloroplasts originate from endosymbiosis between a cyanobacterium and a eukaryotic mitochondriate ancestor. Here, the authors show that the plastid ancestor is related to a cyanobacterial lineage that include N2-fixing filamentous cyanobacteria and species with specialized nitrogen-fixing cells.
- Jesús A. G. Ochoa de Alda
- , Rocío Esteban
- & Jean Houmard
-
Article
| Open Access Comparative genome sequencing reveals genomic signature of extreme desiccation tolerance in the anhydrobiotic midge
Comparative genome sequencing reveals genomic signature of extreme desiccation tolerance in the anhydrobiotic midge
The African chironomid midge, Polypedilum vanderplanki, is able to withstand extreme desiccation. Here the authors sequence the genomes of a desiccation-tolerant and desiccation-sensitive species of chironomid midge and pinpoint genes that may have a role in conferring resistance to desiccation.
- Oleg Gusev
- , Yoshitaka Suetsugu
- & Takahiro Kikawada
-
Article
| Open Access The landscape of kinase fusions in cancer
The landscape of kinase fusions in cancer
Kinases activated by gene fusions represent potentially important targets for the development of cancer drugs. Here, the authors develop a method for detecting gene fusion events in RNA sequencing data from The Cancer Genome Atlas and identify several novel recurrent fusions involving kinases.
- Nicolas Stransky
- , Ethan Cerami
- & Christoph Lengauer
-
Article
| Open Access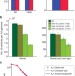 Sequencing an Ashkenazi reference panel supports population-targeted personal genomics and illuminates Jewish and European origins
Sequencing an Ashkenazi reference panel supports population-targeted personal genomics and illuminates Jewish and European origins
Ashkenazi Jews are a genetically isolated population with distinct patterns of genetic diversity. Here, the authors sequence the genomes of 128 Ashkenazi Jewish individuals and use the sequence information to provide insight into the population's European and Middle Eastern origins.
- Shai Carmi
- , Ken Y. Hui
- & Itsik Pe’er
-
Article
| Open Access The Glanville fritillary genome retains an ancient karyotype and reveals selective chromosomal fusions in Lepidoptera
The Glanville fritillary genome retains an ancient karyotype and reveals selective chromosomal fusions in Lepidoptera
Butterflies and moths (Lepidoptera) vary in chromosome number. Here, the authors sequence the genome of the Glanville fritillary butterfly, Melitaea cinxia, show it has the ancestral lepidopteran karyotype and provide insight into how chromosomal fusions have shaped karyotype evolution in butterflies and moths.
- Virpi Ahola
- , Rainer Lehtonen
- & Ilkka Hanski
-
Article
| Open Access Genome dynamics of the human embryonic kidney 293 lineage in response to cell biology manipulations
Genome dynamics of the human embryonic kidney 293 lineage in response to cell biology manipulations
The human embryonic kidney 293 (HEK293) cell lineage is widely used in cell biology and biotechnology. Here, the authors apply whole genome resequencing methods to characterise genomic variation in six HEK293 cell lines and suggest that this variation could affect experiments using these cell lines.
- Yao-Cheng Lin
- , Morgane Boone
- & Nico Callewaert
-
Article
| Open Access A robust SNP barcode for typing Mycobacterium tuberculosis complex strains
A robust SNP barcode for typing Mycobacterium tuberculosis complex strains
Genetic variation in Mycobacterium tuberculosiscomplex (MTBC) bacteria is responsible for differences in factors such as virulence and transmissibility. Here, the authors analyse the genomes of 1,601 MTBC isolates from diverse geographic locations and identify 62 SNPs that may be used to resolve lineages and sublineages of these strains.
- Francesc Coll
- , Ruth McNerney
- & Taane G. Clark
-
Article |
 A smart and versatile theranostic nanomedicine platform based on nanoporphyrin
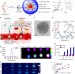
A smart and versatile theranostic nanomedicine platform based on nanoporphyrin
Nanoparticles can be used for therapeutic and diagnostic purposes. Here, the authors report that nanoparticles made of a single chemical building block, called nanoporphyrins, incorporate eight different functionalities, including various types of imaging, drug delivery and cancer therapy.
- Yuanpei Li
- , Tzu-yin Lin
- & Kit S. Lam
-
Article
| Open Access Strong effects of genetic and lifestyle factors on biomarker variation and use of personalized cutoffs
Strong effects of genetic and lifestyle factors on biomarker variation and use of personalized cutoffs
Protein biomarkers could play an important role in the diagnosis and management of diseases. Here the authors investigate the impact of genetic, clinical and lifestyle factors on 92 protein biomarkers for cancer and inflammation and suggest that personalized biomarker thresholds should be used in cancer management.
- Stefan Enroth
- , Åsa Johansson
- & Ulf Gyllensten
-
Article
| Open Access A designer cell-based histamine-specific human allergy profiler
A designer cell-based histamine-specific human allergy profiler
The advancement of sensitive, accurate and non-invasive methods to identify the allergen that drives allergic disease in an individual remains a challenge. Here, the authors develop a synthetic biology approach using human designer cells to profile allergic reactions against an array of allergens measuring histamine release from whole blood.
- David Ausländer
- , Benjamin Eggerschwiler
- & Martin Fussenegger
-
Article |
 Dispersed cells represent a distinct stage in the transition from bacterial biofilm to planktonic lifestyles
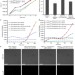
Dispersed cells represent a distinct stage in the transition from bacterial biofilm to planktonic lifestyles
Bacteria can grow as free living planktonic cells or as part of surface-associated biofilms. Here the authors show, for the opportunistic pathogen Pseudomonas aeruginosa, that cells recently dispersed from biofilms are physiologically different from, and more virulent than, planktonic and biofilm cells.
- Song Lin Chua
- , Yang Liu
- & Liang Yang
-
Article
| Open Access Identification of a novel salt tolerance gene in wild soybean by whole-genome sequencing
Identification of a novel salt tolerance gene in wild soybean by whole-genome sequencing
The identification of genes that control economically important traits is an essential step towards crop improvement. Here the authors sequence the genome of the wild soybean and, through a combined genetic and functional approach, identify a new gene affecting salt tolerance in soybean.
- Xinpeng Qi
- , Man-Wah Li
- & Hon-Ming Lam
-
Article
| Open Access Ancestral repeats have shaped epigenome and genome composition for millions of years in Arabidopsis thaliana
Ancestral repeats have shaped epigenome and genome composition for millions of years in Arabidopsis thaliana
Repeated sequences are common in genomes, yet little is known about the long-term evolution of repeats in plants. Here, Maumus and Quesneville show that most of the repeated sequences in the model plant, Arabidopsis thaliana, are old and that many small RNAs correspond to old repeats.
- Florian Maumus
- & Hadi Quesneville
-
Article |
 A mutation burst during the acute phase of Helicobacter pylori infection in humans and rhesus macaques
A mutation burst during the acute phase of Helicobacter pylori infection in humans and rhesus macaques
Helicobacter pylori chronically infects humans, and this is associated with high mutation and recombination rates in the bacterium. Here the authors provide evidence that genome evolution in H. pyloriduring acute infection of the host is orders of magnitude faster than any previously determined mutation rates in bacteria.
- Bodo Linz
- , Helen M. Windsor
- & Barry J. Marshall
-
Article |
 Integrating sequence and array data to create an improved 1000 Genomes Project haplotype reference panel
Integrating sequence and array data to create an improved 1000 Genomes Project haplotype reference panel
1000 Genomes imputation can increase the power of genome-wide association studies to detect genetic variants associated with human traits and diseases. Here, the authors develop a method to integrate and analyse low-coverage sequence data and SNP array data, and show that it improves imputation performance.
- Olivier Delaneau
- , Jonathan Marchini
- & Leena Peltonenz
-
Article |
 Genome-wide adaptive complexes to underground stresses in blind mole rats Spalax
Genome-wide adaptive complexes to underground stresses in blind mole rats Spalax
The blind mole rat (BMR), Spalax galili, is perfectly adapted to life underground. Here, the authors sequence the BMR genome and transcriptome and highlight genomic features that may have played a role in adaptation to extreme underground stressors, such as darkness hypercapnia and hypoxia.
- Xiaodong Fang
- , Eviatar Nevo
- & Jun Wang
-
Article
| Open Access Klebsormidium flaccidum genome reveals primary factors for plant terrestrial adaptation
Klebsormidium flaccidum genome reveals primary factors for plant terrestrial adaptation
Plant colonization of land is an important evolutionary event. Here, the authors sequence the genome of a filamentous terrestrial alga and, through a comparative analysis with related algae and land plant species, provide insight into how aquatic algae adapted to terrestrial environments.
- Koichi Hori
- , Fumito Maruyama
- & Hiroyuki Ohta
-
Article
| Open Access The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes
The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes
Brassica oleracea is plant species comprising economically important vegetable crops. Here, the authors report the draft genome sequence of B. oleracea and, through a comparative analysis with the closely related B. rapa, reveal insights into Brassicaevolution and divergence of interspecific genomes and intraspecific subgenomes.
- Shengyi Liu
- , Yumei Liu
- & Andrew H Paterson
-
Article
| Open Access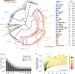 Mobile elements drive recombination hotspots in the core genome of Staphylococcus aureus
Mobile elements drive recombination hotspots in the core genome of Staphylococcus aureus
Horizontal gene transfer occurs in most bacteria, yet it is unclear whether it happens in clonal species. Here, Everitt et al. show widespread within-species recombination, driven by mobile elements, in the genome of the pathogen Staphylococcus aureus, but no recombination between closely related strains.
- Richard G. Everitt
- , Xavier Didelot
- & Daniel J. Wilson
-
Article
| Open Access Specific adaptation of Ustilaginoidea virens in occupying host florets revealed by comparative and functional genomics
Specific adaptation of Ustilaginoidea virens in occupying host florets revealed by comparative and functional genomics
Rice false smut, caused by the pathogenic ascomycete fungus Ustilaginoidea virens (Cooke) Takah, has a significant economic impact on crop production. Here, Zhang et al. report the draft genome sequence of U. virensand provide insight into the evolution of genes involved in pathogenicity and adaptation to a biotrophic and floret-infecting lifestyle.
- Yong Zhang
- , Kang Zhang
- & Wenxian Sun
-
Article
| Open Access The tobacco genome sequence and its comparison with those of tomato and potato
The tobacco genome sequence and its comparison with those of tomato and potato
Common tobacco (Nicotiana tabacum) is a widely cultivated and economically important non-food crop. Here, the authors report the draft genome sequences for three of the most common tobacco varieties and provide insights into the evolution of tobacco through a comparative analysis with closely related species.
- Nicolas Sierro
- , James N.D. Battey
- & Nikolai V. Ivanov
-
Article
| Open Access Spider genomes provide insight into composition and evolution of venom and silk
Spider genomes provide insight into composition and evolution of venom and silk
Spiders use self-produced venom and silk for their daily survival. Here, the authors report the assembled genome of the social velvet spider and a draft assembly of the tarantula genome and, together with proteomic data, provide insights into the evolution of genes that affect venom and silk production.
- Kristian W. Sanggaard
- , Jesper S. Bechsgaard
- & Jun Wang
-
Article
| Open Access The rainbow trout genome provides novel insights into evolution after whole-genome duplication in vertebrates
The rainbow trout genome provides novel insights into evolution after whole-genome duplication in vertebrates
Although whole-genome duplications (WGDs) are rare events, they have an important role in shaping vertebrate evolution. Here, the authors sequence the rainbow trout genome and show that rediploidization after WGD occurs in a slow and stepwise manner.
- Camille Berthelot
- , Frédéric Brunet
- & Yann Guiguen
-
Article |
 Forensic genomics as a novel tool for identifying the causes of mass mortality events
Forensic genomics as a novel tool for identifying the causes of mass mortality events
Mass mortality events of fish and invertebrates are increasingly frequent in coastal zones, yet it is often difficult to identify their causes. Here, the authors provide evidence that a combined field and genomics approach could help identifying the specific cause of mass mortality events.
- Pierre De Wit
- , Laura Rogers-Bennett
- & Stephen R. Palumbi
-
Article
| Open Access Population genomics supports baculoviruses as vectors of horizontal transfer of insect transposons
Population genomics supports baculoviruses as vectors of horizontal transfer of insect transposons
Horizontal transfer of DNA is common among eukaryotes but the vectors involved remain elusive. Here, Gilbert et al. show high frequency of in vivotransposition from the cabbage looper moth into genomes of a baculovirus, suggesting that viruses can act as vectors of horizontal transfer between animals.
- Clément Gilbert
- , Aurélien Chateigner
- & Richard Cordaux
-
Article |
 Dynamic reassortments and genetic heterogeneity of the human-infecting influenza A (H7N9) virus
Dynamic reassortments and genetic heterogeneity of the human-infecting influenza A (H7N9) virus
H7N9 influenza A viruses capable of infecting humans have recently emerged in China. Here, the authors show that these viruses remain genetically diverse, suggesting that they are still in the process of adapting to human hosts.
- Lunbiao Cui
- , Di Liu
- & George F. Gao
-
Article
| Open Access The locust genome provides insight into swarm formation and long-distance flight
The locust genome provides insight into swarm formation and long-distance flight
Locusts are destructive agricultural pests and serve as a model organism for studies of insects. Here, the authors report a draft genome sequence of the migratory locust, Locusta migratoria, and provide insight into genes associated with key survival traits such as phase-change, long-distance migration and feeding.
- Xianhui Wang
- , Xiaodong Fang
- & Le Kang
-
Article |
 Genomic insights into salt adaptation in a desert poplar
Genomic insights into salt adaptation in a desert poplar
Little is known about the genes that confer salt tolerance in trees. Here, Ma et al. report the genome sequence of the desert poplar, Populus euphratica, and provide insight into the genetic architecture and adaptation of this salt tolerant desert poplar.
- Tao Ma
- , Junyi Wang
- & Jianquan Liu

 Enhanced transcriptome maps from multiple mouse tissues reveal evolutionary constraint in gene expression
Enhanced transcriptome maps from multiple mouse tissues reveal evolutionary constraint in gene expression