Featured
-
-
Article
| Open Access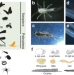 Genomic adaptations to aquatic and aerial life in mayflies and the origin of insect wings
Genomic adaptations to aquatic and aerial life in mayflies and the origin of insect wings
Genomic studies of paleopteran insects, such as mayflies, are needed to reconstruct early insect evolution. Here, Almudi and colleagues present the genome of the mayfly Cloeon dipterum and use transcriptomics to characterize its adaptations to distinct habitats and the origin of insect wings.
- Isabel Almudi
- , Joel Vizueta
- & Fernando Casares
-
Article
| Open Access Transcript isoform sequencing reveals widespread promoter-proximal transcriptional termination in Arabidopsis
Transcript isoform sequencing reveals widespread promoter-proximal transcriptional termination in Arabidopsis
Gene isoforms result from variable transcription start sites (TSSs) and polyadenylation sites (PASs) at the end of transcripts. Here, the authors perform transcript isoform sequencing and find widespread promoter- proximal transcriptional termination in Arabidopsis, suggesting this may represent a checkpoint that regulates plant gene expression.
- Quentin Angelo Thomas
- , Ryan Ard
- & Sebastian Marquardt
-
Article
| Open Access Determinants of transcription factor regulatory range
Determinants of transcription factor regulatory range
Characterization of the distance over which TF binding influences gene expression is important for inferring target genes. Here the authors study this relationship using thousands of genomic data sets, finding two classes of TFs with distinct ranges of regulatory influence modulated by chromatin states of topologically associated domains.
- Chen-Hao Chen
- , Rongbin Zheng
- & X. Shirley Liu
-
Article
| Open Access Tunable genetic devices through simultaneous control of transcription and translation
Tunable genetic devices through simultaneous control of transcription and translation
Synthetic genetic circuits are sensitive to their environment and host cell, requiring many rounds of physical reassembly to achieve a desired function. Here the authors use a multi-level regulatory motif to dynamically tune the function of genetic parts as a step towards robust adaptive circuits.
- Vittorio Bartoli
- , Grace A. Meaker
- & Thomas E. Gorochowski
-
Article
| Open Access Accurate estimation of cell composition in bulk expression through robust integration of single-cell information
Accurate estimation of cell composition in bulk expression through robust integration of single-cell information
Traditional methods for determining cell type composition lack scalability, while single-cell technologies remain costly and noisy compared to bulk RNA-seq. Here, the authors present a highly efficient tool to measure cellular heterogeneity in bulk expression through robust integration of single-cell information.
- Brandon Jew
- , Marcus Alvarez
- & Eran Halperin
-
Article
| Open Access Cysteine synthases CYSL-1 and CYSL-2 mediate C. elegans heritable adaptation to P. vranovensis infection
Cysteine synthases CYSL-1 and CYSL-2 mediate C. elegans heritable adaptation to P. vranovensis infection
Caenorhabditis elegans exhibits multigenerational adaptation to bacterial infection but the mechanisms remain unclear. Here, the authors show that C. elegans parental exposure to Pseudomonas vranovensis promotes offspring resistance to infection, a process mediated by the cysteine synthases CYSL-1 and CYSL-2.
- Nicholas O. Burton
- , Cristian Riccio
- & Eric A. Miska
-
Article
| Open Access Integrative differential expression and gene set enrichment analysis using summary statistics for scRNA-seq studies
Integrative differential expression and gene set enrichment analysis using summary statistics for scRNA-seq studies
Differential expression (DE) and gene set enrichment (GSE) analysis tend to be carried out separately. Here, the authors present iDEA (integrative Differential expression and gene set Enrichment Analysis) for the analysis of scRNAseq data which uses a Baysian approach to jointly model DE and GSE for improved power in both tasks.
- Ying Ma
- , Shiquan Sun
- & Xiang Zhou
-
Article
| Open Access Adar RNA editing-dependent and -independent effects are required for brain and innate immune functions in Drosophila
Adar RNA editing-dependent and -independent effects are required for brain and innate immune functions in Drosophila
Human RNA editing enzymes ADAR1 and ADAR2 are required for innate immune functions and neurological functions, respectively. Here, the authors show that Drosophila Adar has both innate immune and brain functions, despite being the homolog of mammalian ADAR2.
- Patricia Deng
- , Anzer Khan
- & Liam P. Keegan
-
Review Article
| Open Access Multiplexed CRISPR technologies for gene editing and transcriptional regulation
Multiplexed CRISPR technologies for gene editing and transcriptional regulation
Multiplexed CRISPR technologies have recently emerged as powerful approaches for genetic editing and transcriptional regulation. Here the authors review this emerging technology and discuss challenges and considerations for future studies.
- Nicholas S. McCarty
- , Alicia E. Graham
- & Rodrigo Ledesma-Amaro
-
Article
| Open Access Candidate silencer elements for the human and mouse genomes
Candidate silencer elements for the human and mouse genomes
Identification of silencer elements by computational or experimental approaches in a genome-wide manner is still challenging. Here authors define uncharacterized cis-regulatory elements (CREs) in human and mouse genomes likely containing silencer elements, and test them in cells using massively parallel reporter assays to identify silencer elements that showed silencer activity.
- Naresh Doni Jayavelu
- , Ajay Jajodia
- & R. David Hawkins
-
Article
| Open Access Collaborative interactions of heterogenous ribonucleoproteins contribute to transcriptional regulation of sterol metabolism in mice
Collaborative interactions of heterogenous ribonucleoproteins contribute to transcriptional regulation of sterol metabolism in mice
Heterogeneous nuclear ribonucleoproteins (hnRNPs) play critical roles in the biogenesis, localization and transport of RNA. Here authors investigate a role for hnRNPs in sterol metabolism in mice and provide insights into their role in selective promoter activation.
- Zhengyi Zhang
- , An-Chieh Feng
- & Tamer Sallam
-
Article
| Open Access Cellular deconvolution of GTEx tissues powers discovery of disease and cell-type associated regulatory variants
Cellular deconvolution of GTEx tissues powers discovery of disease and cell-type associated regulatory variants
Cellular heterogeneity can confound functional genomics analyses of bulk tissues. Here, Donovan et al. deconvolute bulk GTEx RNA-seq data utilizing scRNA-seq data from the Tabula Muris project and show the power of using cell populations in eQTL analyses to identify disease and cell-type-associated regulatory variants.
- Margaret K. R. Donovan
- , Agnieszka D’Antonio-Chronowska
- & Kelly A. Frazer
-
Article
| Open Access The COMET toolkit for composing customizable genetic programs in mammalian cells
The COMET toolkit for composing customizable genetic programs in mammalian cells
Engineering mammalian cellular functions requires a toolkit of orthogonal and well-characterized genetic components. Here the authors develop COMET: an ensemble of transcription factors, promoters, and accompanying models for the design and construction of genetic programs.
- Patrick S. Donahue
- , Joseph W. Draut
- & Joshua N. Leonard
-
Article
| Open Access High-coverage whole-genome analysis of 1220 cancers reveals hundreds of genes deregulated by rearrangement-mediated cis-regulatory alterations
High-coverage whole-genome analysis of 1220 cancers reveals hundreds of genes deregulated by rearrangement-mediated cis-regulatory alterations
In this study the authors consider the structural variants (SVs) present within cancer cases of the ICGC/TCGA Pan-Cancer Analysis of Whole Genomes (PCAWG) Consortium. They report hundreds of genes, including known cancer-associated genes for which the nearby presence of a SV breakpoint is associated with altered expression.
- Yiqun Zhang
- , Fengju Chen
- & Christian von Mering
-
Article
| Open Access Transcriptomic and open chromatin atlas of high-resolution anatomical regions in the rhesus macaque brain
Transcriptomic and open chromatin atlas of high-resolution anatomical regions in the rhesus macaque brain
Non-human primates share many features with humans and are an important animal model in neuroscience. Here, the authors present a comprehensive transcriptomic and open chromatin atlas of the rhesus macaque brain.
- Senlin Yin
- , Keying Lu
- & Yang Yu
-
Article
| Open Access Dissecting transcriptomic signatures of neuronal differentiation and maturation using iPSCs
Dissecting transcriptomic signatures of neuronal differentiation and maturation using iPSCs
Here, authors present results of a hiPSC transcriptomics study on corticogenesis from multiple donors across four transitions in differentiation. They present a bulk data deconvolution method and show that co-culturing human NPCs with rodent astrocytes results in mutually synergistic maturation.
- Emily E. Burke
- , Joshua G. Chenoweth
- & Andrew E. Jaffe
-
Article
| Open Access Phenotypic changes of HER2-positive breast cancer during and after dual HER2 blockade
Phenotypic changes of HER2-positive breast cancer during and after dual HER2 blockade
HER2-enriched breast cancers within the HER2-positive subtype are addicted to the HER2 pathway. Here, the authors analyse gene expression before, during, and after treatment with anti-HER2-based therapies in the phase II PAMELA clinical trial, finding phenotypic changes induced by treatment.
- Fara Brasó-Maristany
- , Gaia Griguolo
- & Aleix Prat
-
Article
| Open Access Using regulatory variants to detect gene–gene interactions identifies networks of genes linked to cell immortalisation
Using regulatory variants to detect gene–gene interactions identifies networks of genes linked to cell immortalisation
For most human genes nearby regulatory variants explain only a small proportion of their expression variation between individuals. Here the authors show how the impact of a gene’s set of nearby regulatory variant is often linked to the expression levels of distal genes, providing insights into gene networks.
- D. Wragg
- , Q. Liu
- & J. G. D. Prendergast
-
Article
| Open Access A transcriptome-wide Mendelian randomization study to uncover tissue-dependent regulatory mechanisms across the human phenome
A transcriptome-wide Mendelian randomization study to uncover tissue-dependent regulatory mechanisms across the human phenome
Gene expression and how genetic variants can influence gene expression are tissue-specific processes with important implications for phenotypes. Here, Richardson et al. use eQTL data from GTEx and the eQTLGen project in a two-sample SMR + HEIDI framework for causal inference of gene expression associations with complex trait.
- Tom G. Richardson
- , Gibran Hemani
- & George Davey Smith
-
Article
| Open Access Accurate quantification of circular RNAs identifies extensive circular isoform switching events
Accurate quantification of circular RNAs identifies extensive circular isoform switching events
Quantification and characterization of circRNAs in sequencing data remains challenging, hindering efforts to understand their roles and regulation. The algorithm introduced here enables accurate circRNA quantification and permits insight into competitive splicing between linear and circular isoforms.
- Jinyang Zhang
- , Shuai Chen
- & Fangqing Zhao
-
Article
| Open Access De novo generation of hit-like molecules from gene expression signatures using artificial intelligence
De novo generation of hit-like molecules from gene expression signatures using artificial intelligence
High quality hit identification remains a considerable challenge in de novo drug design. Here, the authors train a generative adversarial network with transcriptome profiles induced by a large set of compounds, enabling it to design molecules that are likely to induce desired expression profiles.
- Oscar Méndez-Lucio
- , Benoit Baillif
- & Joerg Wichard
-
Article
| Open Access Single-nuclei RNA-seq on human retinal tissue provides improved transcriptome profiling
Single-nuclei RNA-seq on human retinal tissue provides improved transcriptome profiling
The retina is a heterogeneous tissue composed of multiple cell types. Via single-nuclei RNA sequencing on human neural retinal tissue, the authors characterise the transcriptome profile for individual cell types in the human retina.
- Qingnan Liang
- , Rachayata Dharmat
- & Rui Chen
-
Article
| Open Access Chaperone biomarkers of lifespan and penetrance track the dosages of many other proteins
Chaperone biomarkers of lifespan and penetrance track the dosages of many other proteins
In C. elegans the chaperone hsp-16.2 predicts the penetrance of mutations and lifespan after heat shock, but it is not known why cells express different amounts of hsp-16.2. Here the authors show hsp-16.2 tracks differences in global gene expression capacity, rather than differences in signaling or intrinsic noise in adult intestine cells.
- Nikolay Burnaevskiy
- , Bryan Sands
- & Alexander Mendenhall
-
Article
| Open Access CSB promoter downregulation via histone H3 hypoacetylation is an early determinant of replicative senescence
CSB promoter downregulation via histone H3 hypoacetylation is an early determinant of replicative senescence
Senescence of metabolically active cells is a process linked to ageing. Here the authors reveal that CSB is required to block replicative senescence, and epigenetic control of CSB downregulation triggers proliferative arrest in a p21-dependent manner.
- Clément Crochemore
- , Cristina Fernández-Molina
- & Miria Ricchetti
-
Article
| Open Access Genomic and transcriptomic insights into molecular basis of sexually dimorphic nuptial spines in Leptobrachium leishanense
Genomic and transcriptomic insights into molecular basis of sexually dimorphic nuptial spines in Leptobrachium leishanense
The basis of sexual dimorphism in non-model species may be elusive, in part due to a lack of genomic and molecular resources. Here, Li et al. report a high-quality anuran genome and reveal candidate genes and pathways associated with shaping sexually dimorphic nuptial spines in a moustache toad.
- Jun Li
- , Haiyan Yu
- & Hua Wu
-
Article
| Open Access Molecular insight into RNA polymerase I promoter recognition and promoter melting
Molecular insight into RNA polymerase I promoter recognition and promoter melting
RNA polymerase I (Pol I) catalyses the transcription of pre-ribosomal RNA and for transcription initiation Pol I assembles with core factor and Rrn3 on the rDNA core promoter. Here the authors provide insights into the molecular mechanism of promoter opening by Pol I by determining the cryo-EM structures of two closed, two intermediate and two open Pol I pre-initiation complexes.
- Yashar Sadian
- , Florence Baudin
- & Christoph W. Müller
-
Article
| Open Access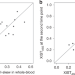 Heritability of skewed X-inactivation in female twins is tissue-specific and associated with age
Heritability of skewed X-inactivation in female twins is tissue-specific and associated with age
Skewing of X chromosome inactivation (XCI) occurs when the silencing of one parental X chromosome is non-random. Here, Zito et al. report XCI patterns in lymphoblastoid cell lines, blood, subcutaneous adipose tissue samples and skin samples of monozygotic and dizygotic twins and find XCI skew to associate with tissue and age.
- Antonino Zito
- , Matthew N. Davies
- & Kerrin S. Small
-
Article
| Open Access Chaperone-mediated ordered assembly of the SAGA and NuA4 transcription co-activator complexes in yeast
Chaperone-mediated ordered assembly of the SAGA and NuA4 transcription co-activator complexes in yeast
Transcription initiation involves the coordinated assembly and activity of large multimeric complexes. Here the authors report on the chaperone-mediated ordered assembly of the SAGA and NuA4 transcription co-activator complexes in fission yeast, providing insight into the de novo assembly of transcriptional complexes and the contribution of dedicated chaperones to this process.
- Alberto Elías-Villalobos
- , Damien Toullec
- & Dominique Helmlinger
-
Article
| Open Access The Firre locus produces a trans-acting RNA molecule that functions in hematopoiesis
The Firre locus produces a trans-acting RNA molecule that functions in hematopoiesis
LncRNA loci potentially contain multiple modes that can exert function, including DNA regulatory elements. Here, the authors generated genetic models in mice to dissect the role of the syntenically conserved lncRNA Firre in the context of hematopoiesis.
- Jordan P. Lewandowski
- , James C. Lee
- & John L. Rinn
-
Article
| Open Access An enhancer cluster controls gene activity and topology of the SCN5A-SCN10A locus in vivo
An enhancer cluster controls gene activity and topology of the SCN5A-SCN10A locus in vivo
Mutations associated with SCN5A, which encodes the major cardiac sodium channel, influence impulse conduction and are associated with arrhythmia disorders. Here, the authors identify an evolutionary conserved, super enhancer-like, regulatory cluster downstream of SCN5A and show that it controls the chromatin architecture of the locus and Scn5a expression.
- Joyce C. K. Man
- , Rajiv A. Mohan
- & Vincent M. Christoffels
-
Article
| Open Access A reference map of murine cardiac transcription factor chromatin occupancy identifies dynamic and conserved enhancers
A reference map of murine cardiac transcription factor chromatin occupancy identifies dynamic and conserved enhancers
Mapping transcription factors (TFs) occupancy is essential for understanding transcriptional programs. Here the authors use biotinylated knockin alleles of key cardiac TFs (GATA4, NKX2-5, MEF2A, MEF2C, SRF, TBX5, TEAD1) to map their genome-wide occupancy in the fetal and adult mouse heart, providing insight into the cardiac transcriptional regulatory network.
- Brynn N. Akerberg
- , Fei Gu
- & William T. Pu
-
Article
| Open Access A nucleotide resolution map of Top2-linked DNA breaks in the yeast and human genome
A nucleotide resolution map of Top2-linked DNA breaks in the yeast and human genome
Topoisomerase 2 (Top2) is known to resolve DNA topological stress through double strand breaks (DSBs), yet Top2 inhibition has been reported to result in a significant amount of single-strand breaks (SSBs). Here the authors develop CC-seq—a method that allows direct mapping of both Top2-linked SSBs and DSBs—and reveal a significant impact of primary DNA sequence on Top2 directed cleavage.
- William H. Gittens
- , Dominic J. Johnson
- & Matthew J. Neale
-
Article
| Open Access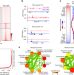 Super-enhancer-guided mapping of regulatory networks controlling mouse trophoblast stem cells
Super-enhancer-guided mapping of regulatory networks controlling mouse trophoblast stem cells
Trophectoderm lineage development is essential for implantation, placentation, and healthy pregnancy. Here the authors map super-enhancers (SEs) in trophoblast stem cells and find both TE-specific master regulators and 150 previous uncharacterised transcription factors that are SE-associated, providing insight into trophectoderm-specific regulatory networks.
- Bum-Kyu Lee
- , Yu jin Jang
- & Jonghwan Kim
-
Article
| Open Access Multi-strategic RNA-seq analysis reveals a high-resolution transcriptional landscape in cotton
Multi-strategic RNA-seq analysis reveals a high-resolution transcriptional landscape in cotton
In-depth functional characterization of genomes relies on comprehensive transcriptome data. Here, the authors employ four complementary RNA sequencing technologies to explore the transcription landscape across 16 tissues or different organ types in diploid A genome cotton using a newly developed computational pipeline.
- Kun Wang
- , Dehe Wang
- & Yuxian Zhu
-
Article
| Open Access Single cell transcriptome in aneuploidies reveals mechanisms of gene dosage imbalance
Single cell transcriptome in aneuploidies reveals mechanisms of gene dosage imbalance
Gene dosage anomalies such as those caused by aneuploidy underlie diseases including Down syndrome. Here, the authors perform allele-specific single cell transcriptome analysis to investigate the mechanisms of gene dosage imbalance in fibroblasts with trisomies T21, T18, T13 and T8.
- Georgios Stamoulis
- , Marco Garieri
- & Stylianos E. Antonarakis
-
Article
| Open Access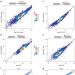 Distinct transcriptional roles for Histone H3-K56 acetylation during the cell cycle in Yeast
Distinct transcriptional roles for Histone H3-K56 acetylation during the cell cycle in Yeast
H3K56Ac promotes nucleosome turnover at promoter-proximal locations and assembly of new nucleosomes during S phase. Here the authors find that H3K56Ac is both a genome-wide activator of transcription in yeast while also repressing promiscuous transcription that occurs after replication fork passage.
- Salih Topal
- , Pauline Vasseur
- & Craig L. Peterson
-
Article
| Open Access Functional interpretation of single cell similarity maps
Functional interpretation of single cell similarity maps
The increasing accessibility of single cell RNA sequencing demands tools that enable data visualization and interpretation. Here, the authors introduce Vision, a flexible annotation tool that operates directly on the manifold of cell-cell similarity and aids interpretation of cellular heterogeneity.
- David DeTomaso
- , Matthew G. Jones
- & Nir Yosef
-
Article
| Open Access Catalytically inactive Dnmt3b rescues mouse embryonic development by accessory and repressive functions
Catalytically inactive Dnmt3b rescues mouse embryonic development by accessory and repressive functions
The family of DNA methyltransferases (Dnmts) consists of four catalytically active enzymes that catalyze DNA methylation and also play a role as transcriptional regulators. Here the authors show that catalytically inactive Dnmt3b rescues a majority of methylation and expression changes in the absence of Dnmt3b during mouse embryonic development.
- Pawel Nowialis
- , Katarina Lopusna
- & Rene Opavsky
-
Article
| Open Access BORDER proteins protect expression of neighboring genes by promoting 3′ Pol II pausing in plants
BORDER proteins protect expression of neighboring genes by promoting 3′ Pol II pausing in plants
In plants, 3′ Pol II pausing helps prevent transcriptional interference. Here the authors provide evidence that BORDER proteins are enriched in intergenic regions and inhibit transcriptional interference between closely spaced genes by preventing RNA polymerase from intruding into the promoters of nearby downstream genes.
- Xuhong Yu
- , Pascal G. P. Martin
- & Scott D. Michaels
-
Article
| Open Access Ageing affects DNA methylation drift and transcriptional cell-to-cell variability in mouse muscle stem cells
Ageing affects DNA methylation drift and transcriptional cell-to-cell variability in mouse muscle stem cells
Age-related tissue alterations have been associated with a decline in stem cell number and function. Here the authors report a single cell multi-omics study of mouse muscle stem cells, combining single cell transcriptome and DNA methylome profiling and find that aged cells have a global increase of uncoordinated transcriptional heterogeneity biased towards genes regulating cell-niche interactions.
- Irene Hernando-Herraez
- , Brendan Evano
- & Wolf Reik
-
Article
| Open Access Maternal pluripotency factors initiate extensive chromatin remodelling to predefine first response to inductive signals
Maternal pluripotency factors initiate extensive chromatin remodelling to predefine first response to inductive signals
Embryonic development produces different cell types in response to a small number of inductive signals. Here, the authors characterise how maternal factors modify chromatin to specify initial competence in Xenopus tropicalis, finding that the pioneering activity of the pluripotency factors Pou5f3 and Sox3 establishes competence for germ layer formation by remodelling chromatin before the onset of signalling.
- George E. Gentsch
- , Thomas Spruce
- & James C. Smith
-
Article
| Open Access Conformational heterogeneity in human interphase chromosome organization reconciles the FISH and Hi-C paradox
Conformational heterogeneity in human interphase chromosome organization reconciles the FISH and Hi-C paradox
Studies comparing Hi-C and FISH data show that in some cases the distance between one pair of loci is paradoxically larger compared to another pair with a smaller value of the contact probability. Here the authors use a theory based on a Generalized Rouse Model for Chromosomes to resolve this paradox.
- Guang Shi
- & D. Thirumalai
-
Article
| Open Access Mapping eGFR loci to the renal transcriptome and phenome in the VA Million Veteran Program
Mapping eGFR loci to the renal transcriptome and phenome in the VA Million Veteran Program
Persistently low levels of estimated glomerular filtration rate (eGFR) are a biomarker of chronic kidney disease. Here, the authors reinterpret the genetic architecture of kidney function across ancestries, to identify not only genes, but the tissue and anatomical contexts of renal homeostasis.
- Jacklyn N. Hellwege
- , Digna R. Velez Edwards
- & Adriana M. Hung
-
Article
| Open Access Diurnal rhythms in gene expression in the prefrontal cortex in schizophrenia
Diurnal rhythms in gene expression in the prefrontal cortex in schizophrenia
Sleep disturbance is common in psychiatric disease, and this may contribute to altered circadian rhythm in gene expression. Here the authors show that rhythms in gene expression in the dorsolateral prefrontal cortex in schizophrenia are different to that seen in healthy controls.
- Marianne L. Seney
- , Kelly Cahill
- & Colleen A. McClung
-
Article
| Open Access Comprehensive transcriptomic analysis of cell lines as models of primary tumors across 22 tumor types
Comprehensive transcriptomic analysis of cell lines as models of primary tumors across 22 tumor types
Cell lines are used ubiquitously in cancer research but how well they represent the tumor type they were derived from is variable. Here, the authors compare transcriptomic profiles of 22 tumor types and cell lines and propose a new comprehensive cell line panel for pan-cancer studies.
- K. Yu
- , B. Chen
- & M. Sirota
-
Article
| Open Access MAPCap allows high-resolution detection and differential expression analysis of transcription start sites
MAPCap allows high-resolution detection and differential expression analysis of transcription start sites
The position, shape and number of transcription start sites (TSS) regulate gene expression. Here authors present MAPCap, a method for high-resolution detection and differential expression analysis of TSS, and apply MAPCap to early fly development, detecting stage and sex-specific promoter and enhancer activity.
- Vivek Bhardwaj
- , Giuseppe Semplicio
- & Asifa Akhtar
-
Article
| Open Access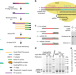 Single-cell CAS-seq reveals a class of short PIWI-interacting RNAs in human oocytes
Single-cell CAS-seq reveals a class of short PIWI-interacting RNAs in human oocytes
PIWI-interacting RNAs (piRNAs) are ~25–33 nt small RNAs expressed in animal germ cells. Here, the authors develop a single-cell small RNA sequencing method and report that a class of ~20-nt piRNAs lacking 3′ end 2′-O-methylation are associated with PIWIL3 protein and predominantly expressed in human and monkey oocytes.
- Qiyuan Yang
- , Ronghong Li
- & Ligang Wu
-
Article
| Open Access Mendelian randomization integrating GWAS and eQTL data reveals genetic determinants of complex and clinical traits
Mendelian randomization integrating GWAS and eQTL data reveals genetic determinants of complex and clinical traits
Many genetic variants identified in genome-wide association studies are associated with gene expression. Here, Porcu et al. propose a transcriptome-wide summary statistics-based Mendelian randomization approach (TWMR) that, applied to 43 human traits, uncovers hundreds of previously unreported gene–trait associations.
- Eleonora Porcu
- , Sina Rüeger
- & Zoltán Kutalik
-
Article
| Open Access Genetic mapping of cell type specificity for complex traits
Genetic mapping of cell type specificity for complex traits
Tissue- and cell type-specific information helps to interpret findings from genome-wide association studies. Here, the authors leverage multiple single cell expression datasets to infer cell type specificity of traits.
- Kyoko Watanabe
- , Maša Umićević Mirkov
- & Danielle Posthuma

 Genome and single-cell RNA-sequencing of the earthworm Eisenia andrei identifies cellular mechanisms underlying regeneration
Genome and single-cell RNA-sequencing of the earthworm Eisenia andrei identifies cellular mechanisms underlying regeneration