Featured
-
-
Article
| Open Access The genomic loci of specific human tRNA genes exhibit ageing-related DNA hypermethylation
The genomic loci of specific human tRNA genes exhibit ageing-related DNA hypermethylation
The epigenome has been shown to change with age, potentially impacting on ageing-related disease. Here the authors investigate the DNA methylation state of the genomic loci of human tRNA and observe enrichment for age-related DNA hypermethylation at tRNA loci.
- Richard J. Acton
- , Wei Yuan
- & Christopher G. Bell
-
Article
| Open Access The histone variant H2A.W and linker histone H1 co-regulate heterochromatin accessibility and DNA methylation
The histone variant H2A.W and linker histone H1 co-regulate heterochromatin accessibility and DNA methylation
T-DNA mutants have been widely used for Arabidopsis gene function characterization. Here, by characterizing a null mutant created by CRISPR, the authors show that previous reported function of H2A.W is confounded by a T-DNA insertion induced chromosomal rearrangement and reveal its role in regulating heterochromatin accessibility.
- Pierre Bourguet
- , Colette L. Picard
- & Olivier Mathieu
-
Article
| Open Access Comprehensive cell type decomposition of circulating cell-free DNA with CelFiE
Comprehensive cell type decomposition of circulating cell-free DNA with CelFiE
Tissue damage and turnover lead to the release of DNA in the blood and can be used to monitor changes in tissue state. Here, the authors developed a tool to accurately estimate the proportion of cell types contributing to cell-free DNA in the blood, with an application to pregnant women and ALS patients.
- Christa Caggiano
- , Barbara Celona
- & Noah Zaitlen
-
Article
| Open Access Systematic inference and comparison of multi-scale chromatin sub-compartments connects spatial organization to cell phenotypes
Systematic inference and comparison of multi-scale chromatin sub-compartments connects spatial organization to cell phenotypes
Computational algorithms to infer chromatin sub-compartments and compartment domains require high-resolution Hi-C maps. Here the authors present Calder, an algorithm that can infer sub-compartments and compartment domains with variable resolution Hi-C data, and they apply it to more than a hundred Hi-C experiments to study sub-compartment repositioning.
- Yuanlong Liu
- , Luca Nanni
- & Giovanni Ciriello
-
Article
| Open Access Interplay of two transcription factors for recruitment of the chromatin remodeling complex modulates fungal nitrosative stress response
Interplay of two transcription factors for recruitment of the chromatin remodeling complex modulates fungal nitrosative stress response
Plant and animal tissues produce nitric oxide and reactive nitrogen species that induce nitrosative stress in pathogens. Here, Jian et al. identify two transcriptional regulators in the phytopathogen Fusarium graminearum that control the nitrosative stress response by modulating the recruitment of a chromatin-remodelling complex at the promoters of the response genes.
- Yunqing Jian
- , Zunyong Liu
- & Zhonghua Ma
-
Article
| Open Access Differential chromatin binding of the lung lineage transcription factor NKX2-1 resolves opposing murine alveolar cell fates in vivo
Differential chromatin binding of the lung lineage transcription factor NKX2-1 resolves opposing murine alveolar cell fates in vivo
How transcription factors regulate cell fates in native tissues is unclear. Here, the authors report that differential chromatin binding of NKX2-1 determines opposing alveolar cell fates in the murine lung, showing loss of YAP/TAZ directs NKX2-1 to alternative binding sites leading to cell fate conversion.
- Danielle R. Little
- , Anne M. Lynch
- & Jichao Chen
-
Article
| Open Access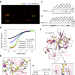 DNMT1 reads heterochromatic H4K20me3 to reinforce LINE-1 DNA methylation
DNMT1 reads heterochromatic H4K20me3 to reinforce LINE-1 DNA methylation
How histone modifications crosstalk with DNA methylation to regulate epigenomic patterning and genome stability in mammals remains elusive. Here, the authors show that DNA methyltransferase DNMT1 is a reader for histone H4K20 trimethylation via its BAH1 domain, which leads to optimal maintenance of DNA methylation at repetitive LINE-1 elements.
- Wendan Ren
- , Huitao Fan
- & Jikui Song
-
Article
| Open Access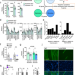 VPS39-deficiency observed in type 2 diabetes impairs muscle stem cell differentiation via altered autophagy and epigenetics
VPS39-deficiency observed in type 2 diabetes impairs muscle stem cell differentiation via altered autophagy and epigenetics
Insulin resistance and lower muscle strength in relation to mass are hallmarks of type 2 diabetes. Here, the authors report alterations in muscle stem cells from individuals with type 2 diabetes that may contribute to these phenotypes through VPS39 mediated effects on autophagy and epigenetics.
- Cajsa Davegårdh
- , Johanna Säll
- & Charlotte Ling
-
Article
| Open Access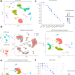 Single cell transcriptional and chromatin accessibility profiling redefine cellular heterogeneity in the adult human kidney
Single cell transcriptional and chromatin accessibility profiling redefine cellular heterogeneity in the adult human kidney
Single cell transcriptomic and epigenomic sequencing of human kidney highlight diverse cell types and states. These findings help characterize a novel population of injured proximal tubule cells and illustrate the power of multi-omic approaches to characterizing human tissue.
- Yoshiharu Muto
- , Parker C. Wilson
- & Benjamin D. Humphreys
-
Article
| Open Access B1a and B2 cells are characterized by distinct CpG modification states at DNMT3A-maintained enhancers
B1a and B2 cells are characterized by distinct CpG modification states at DNMT3A-maintained enhancers
B cell progenitors differentiate into multiple subsets with distinct functions. Here the authors analyze the epigenetic landscapes of sorted B cell subsets using multiple platforms and show that the epigenetic regulator, DNMT3A, is essential for modulating the activity of enhancers critical for B1 and B2 lineage-determining genes.
- Vinay S. Mahajan
- , Hamid Mattoo
- & Shiv Pillai
-
Article
| Open Access Epigenetic modulation of immune synaptic-cytoskeletal networks potentiates γδ T cell-mediated cytotoxicity in lung cancer
Epigenetic modulation of immune synaptic-cytoskeletal networks potentiates γδ T cell-mediated cytotoxicity in lung cancer
Gamma delta (γδ) T cells have potential for use in immunotherapy against tumours. Here, the authors demonstrate that treatment of tumours with DNA methyltransferase inhibitors modulates cytoskeleton arrangements, upregulates adhesion molecules and increases tumour killing by γδ T cells.
- Rueyhung R. Weng
- , Hsuan-Hsuan Lu
- & Hsing-Chen Tsai
-
Article
| Open Access Role of Hakai in m6A modification pathway in Drosophila
Role of Hakai in m6A modification pathway in Drosophila
Drosophila m6A writer complex regulates alternative splicing of the Sex-lethal gene. Here the authors show that a potential E3 ligase Hakai interacts with the fly m6A writer complex and that m6A level is reduced in Hakai mutant flies.
- Yanhua Wang
- , Lifeng Zhang
- & Dong Yan
-
Article
| Open Access Inactivating histone deacetylase HDA promotes longevity by mobilizing trehalose metabolism
Inactivating histone deacetylase HDA promotes longevity by mobilizing trehalose metabolism
Histone acetylations are important epigenetic marks for transcriptional activation and respond to metabolic changes. Here the authors develop a lifespan screen and show that inactivation of the histone deacetylase complex activates longevity and protects against stress via trehalose metabolism.
- Ruofan Yu
- , Xiaohua Cao
- & Weiwei Dang
-
Article
| Open Access The deubiquitinase Usp9x regulates PRC2-mediated chromatin reprogramming during mouse development
The deubiquitinase Usp9x regulates PRC2-mediated chromatin reprogramming during mouse development
During development, H3K27me3 is reallocated from large domains in preimplantation embryos to mark promoters of developmental genes. Here the authors show that the deubiquitinase Usp9x interacts with, deubiquitinates and stabilizes PRC2 and provide evidence that a Usp9x-PRC2 regulatory axis is critical at peri-implantation.
- Trisha A. Macrae
- & Miguel Ramalho-Santos
-
Article
| Open Access Epigenetically regulated digital signaling defines epithelial innate immunity at the tissue level
Epigenetically regulated digital signaling defines epithelial innate immunity at the tissue level
Fine tuning the immune response in line with the degree of threat is central to recognition of a pathogen versus commensal bacteria. Here the authors implicate a digital all-or-nothing cell response in control of the tissue level immune response.
- Helen R. Clark
- , Connor McKenney
- & Sergi Regot
-
Article
| Open Access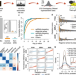 CRISPRi screens reveal a DNA methylation-mediated 3D genome dependent causal mechanism in prostate cancer
CRISPRi screens reveal a DNA methylation-mediated 3D genome dependent causal mechanism in prostate cancer
Prostate cancer risk-associated SNPs are enriched in noncoding CREs. Here the authors perform CRISPRi screens of CREs in prostate cancer cell lines to describe a causal mechanism synergistically driven by a risk SNP and DNA methylation-mediated 3D genome architecture.
- Musaddeque Ahmed
- , Fraser Soares
- & Housheng Hansen He
-
Article
| Open Access Interplay of BAF and MLL4 promotes cell type-specific enhancer activation
Interplay of BAF and MLL4 promotes cell type-specific enhancer activation
The SWI/SNF complex BAF and the histone H3K4 methyltransferase MLL4 (KMT2D) play critical roles in enhancer activation, however the interplay between them has remained unclear. Here the authors show that BAF and MLL4 are interdependent in promoting enhancer activation by lineage-determining transcription factors during adipogenesis.
- Young-Kwon Park
- , Ji-Eun Lee
- & Kai Ge
-
Article
| Open Access N6-methyladenosine RNA modification suppresses antiviral innate sensing pathways via reshaping double-stranded RNA
N6-methyladenosine RNA modification suppresses antiviral innate sensing pathways via reshaping double-stranded RNA
N6-methyladenosine (m6A) RNA modification regulates RNA metabolism, and has been implicated in immune regulation. Here, the authors show that the m6A methyltransferase, METTL3, translocates into the cytoplasm to increase viral RNA m6A modification, decreases viral ds RNA content, and thereby dampens the RIG/MDA5-induced anti-viral immunity.
- Weinan Qiu
- , Qingyang Zhang
- & Pengyuan Yang
-
Article
| Open Access The epigenetic pioneer EGR2 initiates DNA demethylation in differentiating monocytes at both stable and transient binding sites
The epigenetic pioneer EGR2 initiates DNA demethylation in differentiating monocytes at both stable and transient binding sites
DNA methylation turnover is an essential epigenetic process during development. Here, the authors look at the changes in DNA methylation during the differentiation of post-mitotic human monocytes (MO), and find that EGR2 interacts with TET2 and is required for DNA demethylation at its binding sites; revealing EGR2 as an epigenetic pioneer factor in human MO.
- Karina Mendes
- , Sandra Schmidhofer
- & Michael Rehli
-
Article
| Open Access Integrative pan cancer analysis reveals epigenomic variation in cancer type and cell specific chromatin domains
Integrative pan cancer analysis reveals epigenomic variation in cancer type and cell specific chromatin domains
The NCI-60 cancer cell line panel covers multiple cancer types but has not been extensively investigated at the epigenetic level. Here, the authors present H3K4me3, H3K27ac, H3K9me3, and H4K20me3 ChIP-Seq analysis of the cell lines, and describe features of chromatin states and integrative analyses of expression, epigenetic and genetic mutation data.
- Lijin K. Gopi
- & Benjamin L. Kidder
-
Article
| Open Access Strand-specific single-cell methylomics reveals distinct modes of DNA demethylation dynamics during early mammalian development
Strand-specific single-cell methylomics reveals distinct modes of DNA demethylation dynamics during early mammalian development
Erasure of DNA methylation from the parental genomes is critical to reset the methylome of differentiated gametes to pluripotent cells in the blastocyst. Here, the authors present a high-throughput single-cell method that enables strand-specific quantification of DNA methylation and identify distinct modes of DNA demethylation dynamics during early mammalian development.
- Maya Sen
- , Dylan Mooijman
- & Alexander van Oudenaarden
-
Article
| Open Access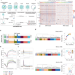 Single-cell multiomics sequencing reveals the functional regulatory landscape of early embryos
Single-cell multiomics sequencing reveals the functional regulatory landscape of early embryos
Extensive epigenetic reprogramming occurs during preimplantation embryo development. Here the authors develop a single cell multiomics sequencing technology that enables profiling of genome-wide chromatin accessibility, DNA methylation and RNA expression in the same individual cell and apply this method to study mouse preimplantation embryos.
- Yang Wang
- , Peng Yuan
- & Liying Yan
-
Article
| Open Access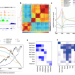 Non-CG methylation and multiple histone profiles associate child abuse with immune and small GTPase dysregulation
Non-CG methylation and multiple histone profiles associate child abuse with immune and small GTPase dysregulation
Early-life adversity is thought to increase the risk of psychopathology through epigenetic mechanisms. Here, the authors profile 6 histone marks, chromatin states and DNA methylation in the lateral amygdala in subjects with a history of early-life adversity.
- Pierre-Eric Lutz
- , Marc-Aurèle Chay
- & Gustavo Turecki
-
Article
| Open Access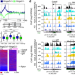 Positioning of nucleosomes containing γ-H2AX precedes active DNA demethylation and transcription initiation
Positioning of nucleosomes containing γ-H2AX precedes active DNA demethylation and transcription initiation
The order of DNA methylation and histone modifications during transcription remained unclear. Here the authors show that HMGA2 induces DNA nicks at TGFB1-responsive genes, promoting nucleosome incorporation containing γ-H2AX, which is required for repair-mediated DNA demethylation and transcription.
- Stephanie Dobersch
- , Karla Rubio
- & Guillermo Barreto
-
Article
| Open Access Chromosomal coordination and differential structure of asynchronous replicating regions
Chromosomal coordination and differential structure of asynchronous replicating regions
Most regions of the mammalian genome replicate both alleles in a synchronous manner, but some loci have been found to replicate asynchronously and the time of replication of each allele is different. Here the authors, by employing clonal mouse cells from a hybrid strain chart replication timing over the entire genome, using polymorphisms to distinguish between the paternal and maternal alleles.
- Britny Blumenfeld
- , Hagit Masika
- & Itamar Simon
-
Article
| Open Access Targeting NSD2-mediated SRC-3 liquid–liquid phase separation sensitizes bortezomib treatment in multiple myeloma
Targeting NSD2-mediated SRC-3 liquid–liquid phase separation sensitizes bortezomib treatment in multiple myeloma
The mechanisms behind acquired resistance to the proteasome inhibitor bortezomib in multiple myeloma remain to be elucidated. Here, the authors show that the histone methyltransferase NSD2 stabilized SRC-3 protein levels, promotes its phase separation and alters H3K36me2 at certain gene promoters resulting in a transcriptional profile that favors resistance of myeloma cells to bortezomib.
- Jing Liu
- , Ying Xie
- & Zhiqiang Liu
-
Article
| Open Access CTCF loss has limited effects on global genome architecture in Drosophila despite critical regulatory functions
CTCF loss has limited effects on global genome architecture in Drosophila despite critical regulatory functions
Although the Drosophila genome has widespread contact domains and CTCF, it remains unclear whether CTCF-dependent domains exist in flies. Here, the authors ablate CTCF in Drosophila and find that CTCF is required to form a small fraction of all domain boundaries, suggesting differences in the role of CTCF for genome folding in flies and vertebrates.
- Anjali Kaushal
- , Giriram Mohana
- & Maria Cristina Gambetta
-
Article
| Open Access PRMT5 inhibition disrupts splicing and stemness in glioblastoma
PRMT5 inhibition disrupts splicing and stemness in glioblastoma
The arginine methyltransferase PRMT5 is over-expressed in cancer and has a role in the maintenance of stem cells. Here, the authors show that PRMT5 inhibitors can block the growth of patient derived glioblastoma stem cell cultures in vitro and in vivo, suggesting that PRMT5 inhibition may be a useful therapeutic strategy
- Patty Sachamitr
- , Jolene C. Ho
- & Peter B. Dirks
-
Article
| Open Access Structural mechanism of bivalent histone H3K4me3K9me3 recognition by the Spindlin1/C11orf84 complex in rRNA transcription activation
Structural mechanism of bivalent histone H3K4me3K9me3 recognition by the Spindlin1/C11orf84 complex in rRNA transcription activation
Spindlin1 is an epigenetic reader that facilitates ribosomal RNA transcription. Here the authors reveal in vitro and structural evidence suggesting that Spindlin1 acts together with C11orf84 to recognize noncanonical bivalent mark of trimethylated lysine 4 and lysine 9 present on histone H3 tail (H3K4me3K9me3).
- Yongming Du
- , Yinxia Yan
- & Chengmin Qian
-
Article
| Open Access The genome-wide impact of trisomy 21 on DNA methylation and its implications for hematopoiesis
The genome-wide impact of trisomy 21 on DNA methylation and its implications for hematopoiesis
Down syndrome has a high co-morbidity with immune and hematopoietic disorders. Here, the authors perform an epigenome-wide association study in newborns with and without Down syndrome to find differential methylation across the genome, including in hematopoietic regulators RUNX1 and FLI1.
- Ivo S. Muskens
- , Shaobo Li
- & Adam J. de Smith
-
Article
| Open Access An all-to-all approach to the identification of sequence-specific readers for epigenetic DNA modifications on cytosine
An all-to-all approach to the identification of sequence-specific readers for epigenetic DNA modifications on cytosine
Identifying readers of epigenetic marks is a critical step for understanding the role of epigenetic marks in biology. Here, the authors applied DAPPL, an all-to-all approach to profile the interactions between TFs and epigenetic modified DNA libraries.
- Guang Song
- , Guohua Wang
- & Heng Zhu
-
Article
| Open Access De novo DNA methyltransferase activity in colorectal cancer is directed towards H3K36me3 marked CpG islands
De novo DNA methyltransferase activity in colorectal cancer is directed towards H3K36me3 marked CpG islands
Aberrant gain of DNA methylation at CpG islands is frequently observed in colorectal tumours. Here the authors use ectopically integrated CpG islands in colorectal cancer cells and find that aberrantly methylated CpG islands are subject to low levels of de novo DNA methylation, and that instead de novo DNA methylation activity is targeted primarily to CpG islands marked by the histone modification H3K36me3.
- Roza H. Ali Masalmeh
- , Francesca Taglini
- & Duncan Sproul
-
Article
| Open Access H3K27me3-rich genomic regions can function as silencers to repress gene expression via chromatin interactions
H3K27me3-rich genomic regions can function as silencers to repress gene expression via chromatin interactions
Mechanisms underlying gene repression and silencers remain poorly understood. Here the authors investigate the role of H3K27me3-rich regions in the genome, as defined from clusters of H3K27me3 peaks, in regulating gene expression via looping.
- Yichao Cai
- , Ying Zhang
- & Melissa Jane Fullwood
-
Article
| Open Access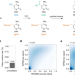 Subtraction-free and bisulfite-free specific sequencing of 5-methylcytosine and its oxidized derivatives at base resolution
Subtraction-free and bisulfite-free specific sequencing of 5-methylcytosine and its oxidized derivatives at base resolution
Specific and quantitative sequencing of cytosine modifications is challenging at base-resolution. Here the authors present TAPSβ and CAPS for subtraction-free whole genome sequencing of 5mC and 5hmC.
- Yibin Liu
- , Zhiyuan Hu
- & Chun-Xiao Song
-
Article
| Open Access LEAFY is a pioneer transcription factor and licenses cell reprogramming to floral fate
LEAFY is a pioneer transcription factor and licenses cell reprogramming to floral fate
Pioneer transcription factors access their DNA binding motifs in closed chromatin and often act in cell fate reprogramming. Here, Jin et al. present biochemical evidence for a pioneer factor in plants and show that LFY promotes floral cell fate and locally unlocks chromatin by displacing histone H1 and recruiting SWI/SNF chromatin remodelers.
- Run Jin
- , Samantha Klasfeld
- & Doris Wagner
-
Article
| Open Access Metabolic regulation of telomere silencing by SESAME complex-catalyzed H3T11 phosphorylation
Metabolic regulation of telomere silencing by SESAME complex-catalyzed H3T11 phosphorylation
Pyruvate kinase phosphorylates histone H3T11 (H3pT11) and represses gene expression by forming a large complex SESAME (Serine-responsive SAM-containing Metabolic Enzyme). Here the authors show that SESAME-catalyzed H3pT11 regulates telomere silencing by promoting Sir2 binding at telomeres and preventing autophagy-mediated Sir2 degradation.
- Shihao Zhang
- , Xilan Yu
- & Shanshan Li
-
Article
| Open Access Nanobody-mediated control of gene expression and epigenetic memory
Nanobody-mediated control of gene expression and epigenetic memory
Targeting chromatin regulators to a gene is emerging as powerful tool to control transcription. Here the authors demonstrate the use of nanobodies against chromatin regulators to control gene expression and epigenetic memory.
- Mike V. Van
- , Taihei Fujimori
- & Lacramioara Bintu
-
Article
| Open Access SOX-5 activates a novel RORγt enhancer to facilitate experimental autoimmune encephalomyelitis by promoting Th17 cell differentiation
SOX-5 activates a novel RORγt enhancer to facilitate experimental autoimmune encephalomyelitis by promoting Th17 cell differentiation
T helper 17 (Th17) cells are important mediators of inflammatory diseases and fungal infection protection, and are critically regulated by the transcription factor (TF), RORγt. Here the authors identify a new enhancer for RORγt, RORCE2, which synergizes with another TF, Sox5, for binding with RORγt promoter and thereby modulation of RORγt expression.
- Yi Tian
- , Chao Han
- & Yuzhang Wu
-
Article
| Open Access Male fertility in Arabidopsis requires active DNA demethylation of genes that control pollen tube function
Male fertility in Arabidopsis requires active DNA demethylation of genes that control pollen tube function
Active DNA demethylation is required for sexual reproduction in plants, but the underlying mechanisms are unknown. Here, the authors show that the DNA glycosylases DEMETER and REPRESSOR OF SILENCING 1 enable the DNA demethylation-dependent activation of genes involved in pollen tube progression.
- Souraya Khouider
- , Filipe Borges
- & Daniel Bouyer
-
Article
| Open Access Targeting aberrant DNA methylation in mesenchymal stromal cells as a treatment for myeloma bone disease
Targeting aberrant DNA methylation in mesenchymal stromal cells as a treatment for myeloma bone disease
Mesenchymal stromal cells (MSCs) have been shown to support multiple myeloma (MM) development. Here, MSCs isolated from the bone marrow of MM patients are shown to have altered DNA methylation patterns and a methyltransferase inhibitor reverts MM-associated bone loss and reduces tumour burden in MM murine models.
- Antonio Garcia-Gomez
- , Tianlu Li
- & Esteban Ballestar
-
Article
| Open Access Core transcription regulatory circuitry orchestrates corneal epithelial homeostasis
Core transcription regulatory circuitry orchestrates corneal epithelial homeostasis
Corneal epithelium shares similar molecular signatures to other stratified epithelia. Here, the authors map super-enhancers and accessible chromatin in corneal epithelium, identifying a transcription regulatory circuit, including RUNX1, PAX6, and SMAD3, required for corneal epithelial identity and homeostasis.
- Mingsen Li
- , Huaxing Huang
- & Hong Ouyang
-
Article
| Open Access Cellular Heterogeneity–Adjusted cLonal Methylation (CHALM) improves prediction of gene expression
Cellular Heterogeneity–Adjusted cLonal Methylation (CHALM) improves prediction of gene expression
Here, the authors introduce Cell Heterogeneity–Adjusted cLonal Methylation (CHALM) as a methylation quantification method that considers the heterogeneity of sequenced bulk cells. They apply CHALM to methylation datasets to detect differentially methylated genes that exhibit distinct biological functions supporting underlying mechanisms.
- Jianfeng Xu
- , Jiejun Shi
- & Wei Li
-
Article
| Open Access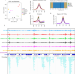 H2AK121ub in Arabidopsis associates with a less accessible chromatin state at transcriptional regulation hotspots
H2AK121ub in Arabidopsis associates with a less accessible chromatin state at transcriptional regulation hotspots
Polycomb Group complexes maintain gene repression through the incorporation of H2AK121ub and H3K27me3. Here, the authors show that H2AK121ub marks less accessible but transcriptionally permissive chromatin, while H3K27me3 enforces a repressed transcriptionally less-permissive state.
- Xiaochang Yin
- , Francisco J. Romero-Campero
- & Yue Zhou
-
Article
| Open Access BET inhibition disrupts transcription but retains enhancer-promoter contact
BET inhibition disrupts transcription but retains enhancer-promoter contact
The role of BRD4 and Mediator in regulating enhancer-promoter interactions is poorly understood. Here the authors find that treatment with BET inhibitors or pharmacological degradation of BRD4 disrupts transcription while having very little effect on enhancer-promoter interactions.
- Nicholas T. Crump
- , Erica Ballabio
- & Thomas A. Milne
-
Article
| Open Access Order and stochasticity in the folding of individual Drosophila genomes
Order and stochasticity in the folding of individual Drosophila genomes
Genomes are partitioned into topologically associating domains (TADs). Here the authors present single-nucleus Hi-C maps in Drosophila at 10 kb resolution, demonstrating the presence of chromatin compartments in individual nuclei, and partitioning of the genome into non-hierarchical TADs at the scale of 100 kb, which resembles population TAD profiles.
- Sergey V. Ulianov
- , Vlada V. Zakharova
- & Sergey V. Razin
-
Article
| Open Access Long first exons and epigenetic marks distinguish conserved pachytene piRNA clusters from other mammalian genes
Long first exons and epigenetic marks distinguish conserved pachytene piRNA clusters from other mammalian genes
The pachytene piRNA loci are transcribed by RNA polymerase II in the male germline of placental mammals. Here the authors show that a long first exon or a long unspliced transcript correlates with germline-specific production of piRNA precursor transcripts and mature piRNAs.
- Tianxiong Yu
- , Kaili Fan
- & Zhiping Weng
-
Article
| Open Access PRC2 and EHMT1 regulate H3K27me2 and H3K27me3 establishment across the zygote genome
PRC2 and EHMT1 regulate H3K27me2 and H3K27me3 establishment across the zygote genome
Dynamic arrangement of epigenetic modifications such as repressive H3K27 methylation is essential for zygote development. Here the authors show that establishment of genome-wide H3K27me3 in zygotes requires EZH2, that EZH1 partially compensates for EZH2 loss, and that EHMT1 is involved in H3K27me2 establishment.
- Tie-Gang Meng
- , Qian Zhou
- & Qing-Yuan Sun
-
Article
| Open Access The molecular basis for recognition of 5′-NNNCC-3′ PAM and its methylation state by Acidothermus cellulolyticus Cas9
The molecular basis for recognition of 5′-NNNCC-3′ PAM and its methylation state by Acidothermus cellulolyticus Cas9
Acidothermus cellulolyticus CRISPR-Cas9 (AceCas9) is a Type II-C enzyme that cleaves DNA in a Protospacer Adjacent Motif (PAM) methylation sensitive fashion. Biochemical analysis and crystal structures of AceCas9 in complex with sgRNA and DNA bearing the correct and incorrect PAM offer insight into the structural basis for the recognition of PAM and its methylation.
- Anuska Das
- , Travis H. Hand
- & Hong Li
-
Article
| Open Access Primary effusion lymphoma enhancer connectome links super-enhancers to dependency factors
Primary effusion lymphoma enhancer connectome links super-enhancers to dependency factors
Primary effusion lymphoma (PEL) has a very poor prognosis. Here, the authors perform H3K27ac HiChIP in PEL cells and generate the PEL enhancer connectome, linking enhancers and promoters in PEL, as well as super-enhancers to dependency factors.
- Chong Wang
- , Luyao Zhang
- & Bo Zhao

 Contrasting epigenetic control of transgenes and endogenous genes promotes post-transcriptional transgene silencing in Arabidopsis
Contrasting epigenetic control of transgenes and endogenous genes promotes post-transcriptional transgene silencing in Arabidopsis