Featured
-
-
Article
| Open Access Heritable DNA methylation marks associated with susceptibility to breast cancer
Heritable DNA methylation marks associated with susceptibility to breast cancer
DNA methylation is associated with breast cancer risk. Here the authors measure DNA methylation in the blood of individuals from 25 Australian families with multiple cases of breast cancer but not known mutations associated with breast cancer risk to identify possible heritable methylation markers.
- Jihoon E. Joo
- , James G. Dowty
- & Yoland Antill
-
Article
| Open Access Co-occurring expression and methylation QTLs allow detection of common causal variants and shared biological mechanisms
Co-occurring expression and methylation QTLs allow detection of common causal variants and shared biological mechanisms
Most expression QTLs (eQTLs) co-occur with a DNA methylation QTL (meQTL), suggesting a common causal variant. Here the authors analyse DNA and RNA from blood and identify eQTL-meQTL pairs likely to share a causal variant, finding that expression and methylation are often genetically co-regulated.
- Brandon L. Pierce
- , Lin Tong
- & Habibul Ahsan
-
Article
| Open Access Exploiting genetic variation to uncover rules of transcription factor binding and chromatin accessibility
Exploiting genetic variation to uncover rules of transcription factor binding and chromatin accessibility
Single nucleotide variants (SNVs) can affect chromatin occupancy by transcription factors (TF). Here the authors mine naturally occurring SNVs to probe cis elements and define contextual sequences that govern in vivo transcription factor chromatin occupancy and chromatin accessibility.
- Vivek Behera
- , Perry Evans
- & Gerd A. Blobel
-
Article
| Open Access scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells
scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells
Relationships between DNA methylation and transcription, and methylation and DNA accessibility can be probed but interrogating all three in the same single cells has not been possible. Here, the authors report the first single-cell method for parallel chromatin accessibility, DNA methylation and transcriptome profiling.
- Stephen J. Clark
- , Ricard Argelaguet
- & Wolf Reik
-
Article
| Open Access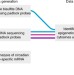 Cytosine modifications exhibit circadian oscillations that are involved in epigenetic diversity and aging
Cytosine modifications exhibit circadian oscillations that are involved in epigenetic diversity and aging
While epigenetic factors have been implicated in the circadian rhythm, the detection of circadian cytosine modifications has remained elusive. Here the authors identify a large number of epigenetically variable cytosines that show circadian oscillations in their modification status in mice.
- Gabriel Oh
- , Sasha Ebrahimi
- & Art Petronis
-
Article
| Open Access Neuropathic MORC2 mutations perturb GHKL ATPase dimerization dynamics and epigenetic silencing by multiple structural mechanisms
Neuropathic MORC2 mutations perturb GHKL ATPase dimerization dynamics and epigenetic silencing by multiple structural mechanisms
Microrchidia CW-type zinc finger protein 2 (MORC2) is an effector of epigenetic silencing by the human silencing hub (HUSH). Here the authors present the crystal structures of MORC2 and disease-causing MORC2 mutants and give mechanistic insights into how MORC2 mediates HUSH-dependent silencing.
- Christopher H. Douse
- , Stuart Bloor
- & Yorgo Modis
-
Article
| Open Access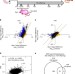 Epigenetic modulation of Fgf21 in the perinatal mouse liver ameliorates diet-induced obesity in adulthood
Epigenetic modulation of Fgf21 in the perinatal mouse liver ameliorates diet-induced obesity in adulthood
FGF21 exerts beneficial metabolic effects on multiple tissues. Here the authors show that the Fgf21 gene is demethylated during the postnatal suckling period, creating an epigenetic memory that determines the responsiveness of the Fgf21 gene to inducers such as PPARα activators or fasting in adulthood.
- Xunmei Yuan
- , Kazutaka Tsujimoto
- & Yoshihiro Ogawa
-
Article
| Open Access The plant-specific histone residue Phe41 is important for genome-wide H3.1 distribution
The plant-specific histone residue Phe41 is important for genome-wide H3.1 distribution
The canonical histone variant H3.1 of vascular plants contains a conserved Phe residue at position 41 that is unique to the plant kingdom. Here, Lu et al. provide evidence that H3.1Phe41 acts collaboratively with the H3.1 core domain to restrict H3.1 deposition to silent regions of the genome.
- Li Lu
- , Xiangsong Chen
- & Xuehua Zhong
-
Article
| Open Access CARM1-expressing ovarian cancer depends on the histone methyltransferase EZH2 activity
CARM1-expressing ovarian cancer depends on the histone methyltransferase EZH2 activity
CARM1 is an arginine methyltransferase often overexpressed in human cancer. Here, the authors show that EZH2 inhibition suppresses growth in CARM1-expressing epithelial ovarian cancer, and examine the mechanism of how CARM1 promotes EZH2-mediated tumor suppressor gene silencing.
- Sergey Karakashev
- , Hengrui Zhu
- & Rugang Zhang
-
Article
| Open Access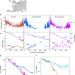 Absence of warmth permits epigenetic memory of winter in Arabidopsis
Absence of warmth permits epigenetic memory of winter in Arabidopsis
Plants use multiple cues to monitor seasonal temperatures. Here, the authors show that Arabidopsis requires not only prolonged cold, but the absence of temperature spikes above 15 °C to epigenetically silence FLC during winter.
- Jo Hepworth
- , Rea L. Antoniou-Kourounioti
- & Caroline Dean
-
Article
| Open Access Methylation profiling identifies two subclasses of squamous cell carcinoma related to distinct cells of origin
Methylation profiling identifies two subclasses of squamous cell carcinoma related to distinct cells of origin
Cutaneous squamous cell carcinoma (cSCC) is a skin cancer that normally progresses from UV-induced actinic keratosis (AK). Here, the authors investigate the epigenomics of cSCC and highlight two distinct subclasses of AK and cSCC originating from distinct keratinocyte differentiation stages.
- Manuel Rodríguez-Paredes
- , Felix Bormann
- & Frank Lyko
-
Article
| Open Access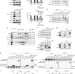 BMI1 regulates androgen receptor in prostate cancer independently of the polycomb repressive complex 1
BMI1 regulates androgen receptor in prostate cancer independently of the polycomb repressive complex 1
The polycomb group protein BMI1 is highly expressed in prostate cancer. Here, the authors demonstrate that BMI1 directly interacts with AR leading to increased AR signaling independently of PRC1 complex and that targeting BMI1 inhibits tumor growth of castration-resistant prostate cancer tumors.
- Sen Zhu
- , Dongyu Zhao
- & Qi Cao
-
Article
| Open Access A naturally occurring epiallele associates with leaf senescence and local climate adaptation in Arabidopsis accessions
A naturally occurring epiallele associates with leaf senescence and local climate adaptation in Arabidopsis accessions
Epigenetic variation underlies various aspects of phenotypic diversity of plants. Here, He et al show a naturally occurring epiallele controls Arabidopsis leaf senescence by regulating the expression of PHEOPHYTIN PHEOPHORBIDE HYDROLASE (PPH), and is associated with local climate adaptation.
- Li He
- , Wenwu Wu
- & Jian-Kang Zhu
-
Article
| Open Access Distinct epigenetic programs regulate cardiac myocyte development and disease in the human heart in vivo
Distinct epigenetic programs regulate cardiac myocyte development and disease in the human heart in vivo
How the cardiac myocyte epigenome is rearranged during development, postnatal maturation and disease is not well understood. Here, the authors investigate the human cardiac myocyte epigenome during development and chronic heart failure and identify distinct epigenetic programs regulating these processes.
- Ralf Gilsbach
- , Martin Schwaderer
- & Lutz Hein
-
Article
| Open Access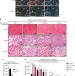 Lsd1 regulates skeletal muscle regeneration and directs the fate of satellite cells
Lsd1 regulates skeletal muscle regeneration and directs the fate of satellite cells
Satellite cells can differentiate both into myocytes and brown adipocytes. Here, the authors show that the histone demethylase Lsd1 prevents adipogenic differentiation of satellite cells by repressing expression of Glis1, and that its ablation changes satellite cell fate towards brown adipocytes and delays muscle regeneration in mice.
- Milica Tosic
- , Anita Allen
- & Roland Schüle
-
Article
| Open Access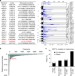 The protective role of DOT1L in UV-induced melanomagenesis
The protective role of DOT1L in UV-induced melanomagenesis
The interaction of DOT1L with MLL oncogenic fusion proteins has been implicated in leukemogenesis. Here, the authors show a contrasting role for DOT1L in protecting UVR-induced melanomagenesis by facilitating DNA repair through interaction with XPC.
- Bo Zhu
- , Shuyang Chen
- & Rutao Cui
-
Article
| Open Access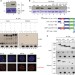 SIRT6-dependent cysteine monoubiquitination in the PRE-SET domain of Suv39h1 regulates the NF-κB pathway
SIRT6-dependent cysteine monoubiquitination in the PRE-SET domain of Suv39h1 regulates the NF-κB pathway
Sirtuins are involved in the regulation of responses to diverse types of cellular stress. Here the authors describe the SirT6-dependent cysteine monoubiquitination of the histone methyltransferase Suv39h1 as part of a regulatory circuit for the NF-κB pathway.
- Irene Santos-Barriopedro
- , Laia Bosch-Presegué
- & Alejandro Vaquero
-
Article
| Open Access Phf8 histone demethylase deficiency causes cognitive impairments through the mTOR pathway
Phf8 histone demethylase deficiency causes cognitive impairments through the mTOR pathway
Mutations in PHF8 gene are genetically associated with X-linked mental retardation. Here, Chen et al. show that Phf8 KO mouse have cognitive and synaptic plasticity impairment, and pharmacological inhibition of mTOR signaling can partially alleviate such defects.
- Xuemei Chen
- , Shuai Wang
- & Charlie Degui Chen
-
Article
| Open Access A PRDX1 mutant allele causes a MMACHC secondary epimutation in cblC patients
A PRDX1 mutant allele causes a MMACHC secondary epimutation in cblC patients
Inborn errors of vitamin B12 metabolism of the cblC class are caused by mutations in the MMACHC gene. Here, Guéant et al. report epi-cblC, a class of cblC in which patients are compound heterozygous for a genetic mutation and a secondary epimutation at the MMACHC locus.
- Jean-Louis Guéant
- , Céline Chéry
- & David S. Rosenblatt
-
Article
| Open Access Trithorax dependent changes in chromatin landscape at enhancer and promoter regions drive female puberty
Trithorax dependent changes in chromatin landscape at enhancer and promoter regions drive female puberty
Before the onset of puberty, Polycomb proteins repress the expression of Kiss1 in KNDy neurons of the arcuate nucleus. Here, by CRISPR-Cas9-directed epigenome editing and RNAi, the authors show that coordinated action of Mll proteins at the Kiss1 promoter and enhancer is required for correct timing of puberty.
- Carlos A. Toro
- , Hollis Wright
- & Alejandro Lomniczi
-
Article
| Open Access Diabetes impairs wound healing by Dnmt1-dependent dysregulation of hematopoietic stem cells differentiation towards macrophages
Diabetes impairs wound healing by Dnmt1-dependent dysregulation of hematopoietic stem cells differentiation towards macrophages
Type 2 diabetes is associated with impaired wound healing, which can lead to limb loss. Here, the authors show that in Type 2 diabetic mouse models, Dnmt1 is upregulated in hematopoietic stem cells, leading to impaired differentiation towards macrophages, reduced macrophage infiltration in the wound and skewed M1/M2 polarization.
- Jinglian Yan
- , Guodong Tie
- & Louis M. Messina
-
Article
| Open Access Chromatin state changes during neural development revealed by in vivo cell-type specific profiling
Chromatin state changes during neural development revealed by in vivo cell-type specific profiling
While transitions between active and repressive chromatin states are essential for differentiation, little is known regarding their role during development of the brain in Drosophila. Here, the authors investigate the large scale chromatin remodelling taking place during fly neural development.
- Owen J. Marshall
- & Andrea H. Brand
-
Article
| Open Access Brd4 binds to active enhancers to control cell identity gene induction in adipogenesis and myogenesis
Brd4 binds to active enhancers to control cell identity gene induction in adipogenesis and myogenesis
Despite being an important cancer drug target, the role of epigenetic reader Brd4 in cell differentiation and development remains unclear. Here, the authors provide evidence that Brd4 plays an important role in adipogenesis and myogenesis by binding to active enhancers to regulate gene expression.
- Ji-Eun Lee
- , Young-Kwon Park
- & Kai Ge
-
Article
| Open Access RAS-pathway mutation patterns define epigenetic subclasses in juvenile myelomonocytic leukemia
RAS-pathway mutation patterns define epigenetic subclasses in juvenile myelomonocytic leukemia
Juvenile myelomonocytic leukemia (JMML) is an aggressive disease with limited options for treatment. Here, the authors analyse the DNA methylome and mutational profile of JMML to define three subgroups with unique molecular and clinical characteristics.
- Daniel B. Lipka
- , Tania Witte
- & Christoph Plass
-
Article
| Open Access Stable transgenerational epigenetic inheritance requires a DNA methylation-sensing circuit
Stable transgenerational epigenetic inheritance requires a DNA methylation-sensing circuit
DNA methylation patterns are inherited over many generations in plants. Here, Williams and Gehring show that the 5-methylcytosine DNA glycosylase ROS1 functions as part of a methylation-sensitive circuit that ensures long-term epigenetic fidelity in Arabidopsis.
- Ben P. Williams
- & Mary Gehring
-
Article
| Open Access H3K14ac is linked to methylation of H3K9 by the triple Tudor domain of SETDB1
H3K14ac is linked to methylation of H3K9 by the triple Tudor domain of SETDB1
SETDB1 is a histone methyltransferase that generates H3K9me3 marks in euchromatic regions. Here the authors show that the triple Tudor domain (3TD) of SETDB1 binds histone H3 tails containing K14 acetylation combined with K9 methylation, and that the K9me–K14ac modification defines a novel chromatin state enriched at SETDB1 binding sites.
- Renata Z. Jurkowska
- , Su Qin
- & Albert Jeltsch
-
Article
| Open Access Ancestral perinatal obesogen exposure results in a transgenerational thrifty phenotype in mice
Ancestral perinatal obesogen exposure results in a transgenerational thrifty phenotype in mice
Early life exposure to endocrine disrupting chemicals has been linked to increased adiposity during adulthood. Here Chamorro-García et al. show that ancestral exposure to the obesogen tributyltin causes obesity in untreated F4 generation male descendants by inducing heritable changes in genome architecture that promote a thrifty phenotype.
- Raquel Chamorro-Garcia
- , Carlos Diaz-Castillo
- & Bruce Blumberg
-
Article
| Open Access Epigenetic targeting of bromodomain protein BRD4 counteracts cancer cachexia and prolongs survival
Epigenetic targeting of bromodomain protein BRD4 counteracts cancer cachexia and prolongs survival
Cachexia is a metabolic syndrome leading to muscle and adipose tissue loss in majority of cancer patients. Here the authors show that, in a mouse model, BET inhibitor JQ1 counteracts muscle and adipose tissue wasting tempering cachexia and prolonging survival through a mechanism unrelated to tumour growth.
- Marco Segatto
- , Raffaella Fittipaldi
- & Giuseppina Caretti
-
Article
| Open Access DNA methylation signatures follow preformed chromatin compartments in cardiac myocytes
DNA methylation signatures follow preformed chromatin compartments in cardiac myocytes
Chromatin is organized in higher order A and B compartments, reflecting active and inactive chromatin. Here, the authors provide evidence that in cardiac myocytes DNA methylation is established in preformed chromatin compartments and may be dispensable for higher order chromatin organization.
- Stephan Nothjunge
- , Thomas G. Nührenberg
- & Ralf Gilsbach
-
Article
| Open Access Mrg15 stimulates Ash1 H3K36 methyltransferase activity and facilitates Ash1 Trithorax group protein function in Drosophila
Mrg15 stimulates Ash1 H3K36 methyltransferase activity and facilitates Ash1 Trithorax group protein function in Drosophila
Ash1 is an H3K36 methyltransferase, Trithorax group protein and is critical in antagonizing Polycomb silencing. Here, the authors purify the Drosophila Ash1 complex and identify Mrg15 and Nurf55 as subunits, finding that Mrg15 is recruited by Ash1 and reinforces Ash1 activity.
- Chang Huang
- , Fu Yang
- & Bing Zhu
-
Article
| Open Access FBXO32 promotes microenvironment underlying epithelial-mesenchymal transition via CtBP1 during tumour metastasis and brain development
FBXO32 promotes microenvironment underlying epithelial-mesenchymal transition via CtBP1 during tumour metastasis and brain development
Epithelial-to-mesenchymal transition (EMT) regulates both processes of organism development and changes in cell state causing disease. Here, the authors show that an E3 ubiquitin ligase, FBXO32, regulates EMT via CtBP1 and the transcriptional program, and also mediates cancer metastatic burden and neurogenesis.
- Sanjeeb Kumar Sahu
- , Neha Tiwari
- & Vijay K. Tiwari
-
Article
| Open Access Hit-and-run epigenetic editing prevents senescence entry in primary breast cells from healthy donors
Hit-and-run epigenetic editing prevents senescence entry in primary breast cells from healthy donors
“Although aberrant promoter DNA hypermethylation is a hallmark of cancer, it is not clear whether it is sufficient to drive transformation. Here, the authors use CRISPR-dCas9 to perform hit-and-run epigenetic editing, which prevents senescence entry in primary breast cells from healthy donors.”
- Emily A. Saunderson
- , Peter Stepper
- & Gabriella Ficz
-
Article
| Open Access Genome-wide genetic and epigenetic analyses of pancreatic acinar cell carcinomas reveal aberrations in genome stability
Genome-wide genetic and epigenetic analyses of pancreatic acinar cell carcinomas reveal aberrations in genome stability
Pancreatic acinar cell carcinoma (ACC) is an aggressive exocrine tumor with largely unknown biology. Here, the authors perform genome- and epigenome-wide analyses from normal and ACC pancreatic tissue that identify aberrations in genome stability and cell cycle control.
- Cornelia Jäkel
- , Frank Bergmann
- & Peter Schmezer
-
Article
| Open Access High-frequency recombination between members of an LTR retrotransposon family during transposition bursts
High-frequency recombination between members of an LTR retrotransposon family during transposition bursts
Retrotransposons are abundant in eukaryotic genomes. Here, Sanchez et al. show evidence of high-frequency recombination between members of an LTR retrotransposon family during transposition bursts in Arabidopsis.
- Diego H. Sanchez
- , Hervé Gaubert
- & Jerzy Paszkowski
-
Article
| Open Access Epigenome-wide association studies identify DNA methylation associated with kidney function
Epigenome-wide association studies identify DNA methylation associated with kidney function
Genome-wide association studies of kidney function show enrichment of associated genetic variants in regulatory regions. Here, the authors perform epigenome-wide association studies of kidney function and disease, identifying 19 CpG sites significantly associated with these.
- Audrey Y. Chu
- , Adrienne Tin
- & Anna Köttgen
-
Article
| Open Access Contribution of epigenetic landscapes and transcription factors to X-chromosome reactivation in the inner cell mass
Contribution of epigenetic landscapes and transcription factors to X-chromosome reactivation in the inner cell mass
X-chromosome inactivation is reversed in the mouse inner cell mass (ICM) through a mechanism that is not fully understood. Here, the authors investigate this process and characterize the contributions of the epigenetic landscape and transcription factors in X-linked gene reactivation dynamics.
- Maud Borensztein
- , Ikuhiro Okamoto
- & Edith Heard
-
Article
| Open Access The Polycomb group protein CBX6 is an essential regulator of embryonic stem cell identity
The Polycomb group protein CBX6 is an essential regulator of embryonic stem cell identity
In mammals, five different CBX proteins can be part of Polycomb repressive complex 1 (PRC1). Here, the authors provide evidence that CBX6 plays an essential role in regulating pluripotency in embryonic stem cells and that CBX6 functions as part of both canonical and non-canonical PRC1 complexes.
- Alexandra Santanach
- , Enrique Blanco
- & Luciano Di Croce
-
Article
| Open Access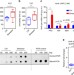 DNA N6-methyladenine is dynamically regulated in the mouse brain following environmental stress
DNA N6-methyladenine is dynamically regulated in the mouse brain following environmental stress
N6-methyladenine is a covalent epigenetic modification of the genome. Here, Yao and colleagues show that N6-methyladenine level in the mouse brain is dynamic following environmental stress, and the subsequent differential gene expression is correlated with LINE transposon expression.
- Bing Yao
- , Ying Cheng
- & Peng Jin
-
Article
| Open Access DNA methylation and transcriptional trajectories during human development and reprogramming of isogenic pluripotent stem cells
DNA methylation and transcriptional trajectories during human development and reprogramming of isogenic pluripotent stem cells
While DNA methylation and gene expression data are widely available for animal models, comprehensive data from human development is rarer. Here, the authors generated transcriptional and DNA methylation data from 21 organs during human development and 6 isogenic induced pluripotent stem cell lines.
- Matthias S. Roost
- , Roderick C. Slieker
- & Susana M. Chuva de Sousa Lopes
-
Article
| Open Access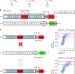 Deciphering TAL effectors for 5-methylcytosine and 5-hydroxymethylcytosine recognition
Deciphering TAL effectors for 5-methylcytosine and 5-hydroxymethylcytosine recognition
Transcription activator-like effector proteins recognise specific DNA sequences via tandem repeats. Here the authors demonstrate TALEs can recognise the methylated bases 5mC and 5hmC, enabling them to detect epigenetic modifications.
- Yuan Zhang
- , Lulu Liu
- & Chengqi Yi
-
Article
| Open Access Non-canonical reader modules of BAZ1A promote recovery from DNA damage
Non-canonical reader modules of BAZ1A promote recovery from DNA damage
ISWI chromatin remodelers regulate DNA accessibility and have been implicated in DNA damage repair. Here, the authors uncover functions, in response to DNA damage, for the bromodomain of the ISWI subunit BAZ1B and for the non-canonical PHD and bromodomain modules of the paralog BAZ1A.
- Mariano Oppikofer
- , Meredith Sagolla
- & Andrea G. Cochran
-
Article
| Open Access LSD1 protects against hippocampal and cortical neurodegeneration
LSD1 protects against hippocampal and cortical neurodegeneration
“LSD1 is a histone demethylase that plays many roles during development. Here, the authors provide evidence that loss of LSD1 in adult mice leads to paralysis and neurodegeneration in the hippocampus and cortex and suggest a potential link between LSD1 and human neurodegenerative disease.
- Michael A. Christopher
- , Dexter A. Myrick
- & David J. Katz
-
Article
| Open Access Genetic and epigenetic features direct differential efficiency of Xist-mediated silencing at X-chromosomal and autosomal locations
Genetic and epigenetic features direct differential efficiency of Xist-mediated silencing at X-chromosomal and autosomal locations
Xist RNA is required for X chromosome inactivation but it is not well understood how Xist silences some regions more efficiently than others. Here, the authors induce ectopic Xist expression from multiple different X-linked and autosomal loci in cells to explore Xist function.
- Agnese Loda
- , Johannes H. Brandsma
- & Joost Gribnau
-
Article
| Open Access ZNF516 suppresses EGFR by targeting the CtBP/LSD1/CoREST complex to chromatin
ZNF516 suppresses EGFR by targeting the CtBP/LSD1/CoREST complex to chromatin
EGFR is a well-known oncogene; however, the mechanisms regulating its expression are still unclear. Here, analysing genome-wide chromatin associations, the authors show that in breast cancer cells ZNF516 represses EGFR transcription through the interaction with the CtBP/LSD1/CoREST complex.
- Lifang Li
- , Xinhua Liu
- & Luyang Sun
-
Article
| Open Access Modular fluorescence complementation sensors for live cell detection of epigenetic signals at endogenous genomic sites
Modular fluorescence complementation sensors for live cell detection of epigenetic signals at endogenous genomic sites
Tools for imaging epigenetic modifications can shed light on the regulation of epigenetic processes. Here, the authors present a fluorescence complementation approach for detection of DNA and histone methylation at endogenous genomic sites allowing following of dynamic changes of these marks by live-cell microscopy.
- Cristiana Lungu
- , Sabine Pinter
- & Albert Jeltsch
-
Article
| Open Access Rapid and reversible epigenome editing by endogenous chromatin regulators
Rapid and reversible epigenome editing by endogenous chromatin regulators
Understanding the link between epigenetic marks and gene regulation requires the development of new tools to directly manipulate chromatin. Here the authors demonstrate a Cas9-based system to recruit chromatin remodelers to loci of interest, allowing rapid, reversible manipulation of epigenetic states.
- Simon M. G. Braun
- , Jacob G. Kirkland
- & Gerald R. Crabtree
-
Article
| Open Access An HDAC3-PROX1 corepressor module acts on HNF4α to control hepatic triglycerides
An HDAC3-PROX1 corepressor module acts on HNF4α to control hepatic triglycerides
HDAC3 is a critical mediator of hepatic lipid metabolism and its loss leads to fatty liver. Here, the authors characterize the liver HDAC3 interactome in vivo, provide evidence that HDAC3 interacts with PROX1, and show that HDAC3 and PROX1 control expression of genes regulating lipid homeostasis.
- Sean M. Armour
- , Jarrett R. Remsberg
- & Mitchell A. Lazar
-
Article
| Open Access Caloric restriction delays age-related methylation drift
Caloric restriction delays age-related methylation drift
Caloric restriction has been shown to increase lifespan in mammals. Here, the authors provide evidence that age-related methylation drift correlates with lifespan and that caloric restriction in mice and rhesus monkeys results in attenuation of age-related methylation drift.
- Shinji Maegawa
- , Yue Lu
- & Jean-Pierre J. Issa
-
Article
| Open Access Copy number rather than epigenetic alterations are the major dictator of imprinted methylation in tumors
Copy number rather than epigenetic alterations are the major dictator of imprinted methylation in tumors
Altered genomic imprinting is frequently reported in cancer. Here, the authors analyze copy number and methylation in cancer cell lines and primary tumors to show that imprinted methylation profiles represent the accumulation of copy number alteration, rather than epigenetic alterations.
- Alex Martin-Trujillo
- , Enrique Vidal
- & David Monk

 Zeb1-Hdac2-eNOS circuitry identifies early cardiovascular precursors in naive mouse embryonic stem cells
Zeb1-Hdac2-eNOS circuitry identifies early cardiovascular precursors in naive mouse embryonic stem cells